Abstract
Purpose
Pediatric solid tumors are significantly different from adult tumors. Studies have revealed genomic aberrations in pediatric solid tumors, but these analyses were based on Western populations. Currently, it is not known to what extent the existing genomic findings represent differences in ethnic backgrounds.
Experimental
Design
We retrospectively analyzed the basic clinical characteristics of the patients, including age, cancer type, and sex distribution, and further analyzed the somatic and germline mutations of cancer-related genes in a Chinese pediatric cohort. In addition, we investigated the clinical significance of genomic mutations on therapeutic, prognostic, diagnostic, and preventive actions.
Results
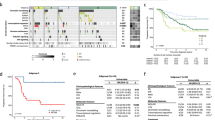
Our study enrolled 318 pediatric patients, including 234 patients with CNS tumors and 84 patients with non-CNS tumors. Somatic mutation analysis showed that there were significant differences in mutation types between CNS tumors and non-CNS tumors. P/LP germline variants were identified in 8.49% of patients. In total, 42.8% patients prompted diagnostic, 37.7% patients prompted prognostic, 58.2% patients prompted therapeutic, and 8.5% patients prompted tumor-predisposing and preventive, and we found that genomic findings might improve clinical management.
Conclusions
Our study is the first large-scale study to analyze the landscape of genetic mutations in pediatric patients with solid tumors in China. Genomic findings in CNS and non-CNS solid pediatric tumors provide evidence for the clinical classification and individualized treatment of pediatric tumors, and they will facilitate improvement of clinical management. Data presented in this study should serve as a reference to guide the future design of clinical trials.




Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the present study are available from the corresponding author on reasonable request.
References
Alaggio R, Hill DA, Jacques TS et al (2021) WHO classification of tumors: pediatric tumors. IARC, Lyon, France
Bagchi S, Yuan R, Engleman EG (2021) Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu Rev Pathol 16:223–249
Berbegall AP, Villamón E, Tadeo I et al (2014) Neuroblastoma after childhood: prognostic relevance of segmental chromosome aberrations, ATRX protein status, and immune cell infiltration. Neoplasia 16(6):471–480
Chan TA, Yarchoan M, Jaffee E et al (2019) Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol 30(1):44–56
Chang W, Brohl AS, Patidar R et al (2016) MultiDimensional clinomics for precision therapy of children and adolescent young adults with relapsed and refractory cancer: a report from the center for cancer research. Clin Cancer Res 22(15):3810–3820
Chen S, Zhou Y, Chen Y et al (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):i884–i890
Doroshow DB, Bhalla S, Beasley MB et al (2021) PD-L1 as a biomarker of response to immune-checkpoint inhibitors. Nat Rev Clin Oncol 18(6):345–362
Ferrari A, Dileo P, Casanova M et al (2003) Rhabdomyosarcoma in adults. A retrospective analysis of 171 patients treated at a single institution. Cancer 98(3):571–580
Fiala EM, Jayakumaran G, Mauguen A et al (2021) Prospective pan-cancer germline testing using MSK-IMPACT informs clinical translation in 751 patients with pediatric solid tumors. Nat Cancer 2:357–365
Filbin M, Monje M (2019) Developmental origins and emerging therapeutic opportunities for childhood cancer. Nat Med 25(3):367–376
GBD 2017 Childhood Cancer Collaborators (2019) The global burden of childhood and adolescent cancer in 2017: an analysis of the global burden of disease study 2017. Lancet Oncol 20(9):1211–1225
George SL, Izquierdo E, Campbell J et al (2019) A tailored molecular profiling programme for children with cancer to identify clinically actionable genetic alterations. Eur J Cancer 121:224–235
Grabovska Y, Mackay A, O’Hare P et al (2020) Pediatric pan-central nervous system tumor analysis of immune-cell infiltration identifies correlates of antitumor immunity. Nat Commun 11:4324
Gröbner SN, Worst BC, Weischenfeldt J et al (2018) The landscape of genomic alterations across childhood cancers. Nature 555(7696):321–327
Gutiérrez-Jimeno M, Alba-Pavón P, Astigarraga I et al (2021) Clinical value of NGS genomic studies for clinical management of pediatric and young adult bone sarcomas. Cancers (basel) 13(21):5436
Harris MH, DuBois SG, Glade Bender JL et al (2016) Multicenter feasibility study of tumor molecular profiling to inform therapeutic decisions in advanced pediatric solid tumors: the individualized cancer therapy (iCat) study. JAMA Oncol 2(5):608–615
Hwang KB, Lee IH, Li H et al (2019) Comparative analysis of whole-genome sequencing pipelines to minimize false negative findings. Sci Rep 9(1):3219
Johnson A, Severson E, Gay L et al (2017) Comprehensive genomic profiling of 282 pediatric low- and high-grade gliomas reveals genomic drivers, tumor mutational burden, and hypermutation signatures. Oncologist 22(12):1478–1490
Jones DTW, Banito A, Grünewald TGP et al (2019) Molecular characteristics and therapeutic vulnerabilities across paediatric solid tumours. Nat Rev Cancer 19(8):420–438
Lai Z, Markovets A, Ahdesmaki M et al (2016) VarDict: a novel and versatile variant caller for next-generation sequencing in cancer research. Nucleic Acids Res 44(11):e108
Li Q, Wang K (2017) InterVar: clinical interpretation of genetic variants by the 2015 ACMG-AMP guidelines. Am J Hum Genet 100(2):267–280
Li MM, Datto M, Duncavage EJ et al (2017) Standards and guidelines for the interpretation and reporting of sequence variants in cancer: a Joint consensus recommendation of the association for molecular pathology, American society of clinical oncology, and college of American pathologists. J Mol Diagn 19(1):4–23
Louis DN, Wesseling P, Paulus W et al (2018) cIMPACT-NOW update 1: not otherwise specified (NOS) and not elsewhere classified (NEC). Acta Neuropathol 135:481–484
Louis DN, Perry A, Wesseling P et al (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol 23:1231–1251
Ma X, Liu Y, Liu Y et al (2018) Pan-cancer genome and transcriptome analyses of 1699 paediatric leukaemias and solid tumours. Nature 555(7696):371–376
Mody RJ, Wu YM, Lonigro RJ et al (2015) Integrative clinical sequencing in the management of refractory or relapsed cancer in youth. JAMA 314(9):913–925
Newman AM, Bratman SV, Stehr H et al (2014) FACTERA: a practical method for the discovery of genomic rearrangements at breakpoint resolution. Bioinformatics 30(23):3390–3393
Newman S, Nakitandwe J, Kesserwan CA et al (2021) Genomes for kids: the scope of pathogenic mutations in pediatric cancer revealed by comprehensive DNA and RNA sequencing. Cancer Discov 11(12):3008–3027
Niu B, Ye K, Zhang Q et al (2014) MSIsensor: microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics 30(7):1015–1016
Oberg JA, Glade Bender JL, Sulis ML et al (2016) Implementation of next generation sequencing into pediatric hematology-oncology practice: moving beyond actionable alterations. Genome Med 8(1):133
Ortiz MV, Kobos R, Walsh M et al (2016) Integrating genomics into clinical pediatric oncology using the molecular tumor board at the memorial sloan kettering cancer center. Pediatr Blood Cancer 63(8):1368–1374
Parsons DW, Roy A, Yang Y et al (2016) Diagnostic yield of clinical tumor and germline whole-exome sequencing for children with solid tumors. JAMA Oncol 2(5):616–624
Pfister SM, Reyes-Múgica M, Chan JKC et al (2022) A summary of the inaugural WHO classification of pediatric tumors: transitioning from the optical into the molecular era. Cancer Discov 12(2):331–355
Richards S, Aziz N, Bale S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American college of medical genetics and genomics and the association for molecular pathology. Genet Med 17(5):405–424
Roux A, Pallud J, Saffroy R et al (2020) High-grade gliomas in adolescents and young adults highlight histomolecular differences from their adult and pediatric counterparts. Neuro Oncol 22(8):1190–1202
Steliarova-Foucher E, Colombet M, Ries LAG et al (2017) International incidence of childhood cancer, 2001–10: a population-based registry study. Lancet Oncol 18(6):719–731
Suh E, Stratton KL, Leisenring WM et al (2020) Late mortality and chronic health conditions in long-term survivors of early-adolescent and young adult cancers: a retrospective cohort analysis from the childhood cancer survivor study. Lancet Oncol 21(3):421–435
Surrey LF, MacFarland SP, Chang F et al (2019) Clinical utility of custom-designed NGS panel testing in pediatric tumors. Genome Med 11(1):32
Sweet-Cordero EA, Biegel JA (2019) The genomic landscape of pediatric cancers: implications for diagnosis and treatment. Science 363(6432):1170–1175
Talevich E, Shain AH, Botton T et al (2016) CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLoS Comput Biol 12(4):e1004873
Thomas RK, Baker AC, Debiasi RM et al (2007) High-throughput oncogene mutation profiling in human cancer. Nat Genet 39(3):347–351
Vellichirammal NN, Chaturvedi NK, Joshi SS et al (2021) Fusion genes as biomarkers in pediatric cancers: a review of the current state and applicability in diagnostics and personalized therapy. Cancer Lett 499:24–38
Walterhouse D, Watson A (2007) Optimal management strategies for rhabdomyosarcoma in children. Paediatr Drugs 9(6):391–400
Wienke J, Dierselhuis MP, Tytgat GAM et al (2021) The immune landscape of neuroblastoma: challenges and opportunities for novel therapeutic strategies in pediatric oncology. Eur J Cancer 144:123–150
Wong M, Mayoh C, Lau LMS et al (2020) Whole genome, transcriptome and methylome profiling enhances actionable target discovery in high-risk pediatric cancer. Nat Med 26(11):1742–1753
Zhang J, Walsh MF, Wu G et al (2015) Germline mutations in predisposition genes in pediatric cancer. N Engl J Med 373(24):2336–2346
Acknowledgements
The authors would like to thank all patients and their families who participated in this study. The authors would also like to thank Mr. Chuang Qi, Mr. Wanglong Deng, Mr. Chao Song, and Mr. Fanfeng Bu from Simceredx for the kindly assistance.
Funding
This work was supported by the National Natural Science Foundation of China [Grant Nos. 81874083; 82072776], Taishan Pandeng Scholar Program of Shandong Province [Grant No. tspd20210322], Natural Science Foundation of Shandong Province of China [Grant No. ZR2021LSW025], Key Clinical Research Project of Clinical Research Center of Shandong University [Grant No. 2020SDUCRCA011], and Jinan Science and Technology Bureau of Shandong Province [Grant No. 2021GXRC029].
Author information
Authors and Affiliations
Contributions
JG, BH, and GL planed the study, analyzed data and interpreted the results. JG, LD, CW, and NL wrote the manuscript. NL, TH, XZ, and ML participated in data analysis. TS, CQ, RD, BH, and GL contributed the manuscript revision and editing. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare there are no competing interests.
Ethical approval and consent to participate
The study was approved by the Ethics Committee of Qilu Hospital of Shandong University.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2023_4756_MOESM2_ESM.tif
Supplementary file2 Schematic representation of the protein domain structure of the most common BRAF fusion in CNS tumors and the most common EWSR1 fusion in non-CNS solid tumors. The number of fusion events is displayed on the far right (TIF 1487 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gong, J., Dong, L., Wang, C. et al. Molecular genomic landscape of pediatric solid tumors in Chinese patients: implications for clinical significance. J Cancer Res Clin Oncol 149, 8791–8802 (2023). https://doi.org/10.1007/s00432-023-04756-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04756-5