Abstract
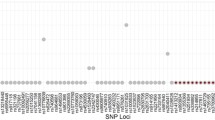
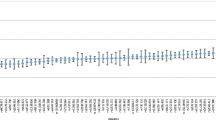
The Ion AmpliSeq™ HID single nucleotide polymorphism (SNP) panel, a primer pool of 103 autosomal SNPs and 33 Y-SNPs, was evaluated using the Ion 314™ Chip on the Ion PGM™ Sequencer with four DNA samples. The study focused on the sequencing of DNA at three different initial target quantities, related interpretation issues, and concordance of results with another sequencing platform, i.e., Genome Analyzer IIx. With 10 ng of template DNA, all genotypes at the 136 SNPs were detected. With 1 ng of DNA, all SNPs were detected and one SNP locus in one sample showed extreme heterozygote imbalance on allele coverage. With 100 pg of DNA, an average of 1.6 SNP loci were not detected, and an average of 4.3 SNPs showed heterozygote imbalance. The average sequence coverage was 945–600× at autosomal SNPs and 465–209× at Y-SNPs for 10 ng–100 pg of DNA. The average heterozygote allele coverage ratio was 89.6–61.8 % for 10 ng–100 pg of DNA. At 10 ng of DNA, all genotypes of the 95 SNPs shared between the two different sequencing platforms were concordant except for one SNP, rs1029047. The error was due to the misalignment of a flanking homopolymer. Overall, the data support that genotyping a large battery of SNPs is feasible with massively parallel sequencing. With barcode systems, better allele balance, and specifically designed alignment software, a more comprehensive rapid genotyping and more cost-effective results may be obtained from multiple samples in one analysis than are possible with current typing and capillary electrophoresis systems.




Similar content being viewed by others
References
Sanchez JJ, Phillips C, Børsting C, Balogh K, Bogus M, Fondevila M, Harrison CD, Musgrave-Brown E, Salas A, Syndercombe-Court D, Schneider PM, Carracedo A, Morling N (2009) A multiplex assay with 52 single nucleotide polymorphisms for human identification. Electrophoresis 27:1713–1724
Pakstis AJ, Speed WC, Fang R, Hyland FC, Furtado MR, Kidd JR, Kidd KK (2010) SNPs for a universal individual identification panel. Hum Genet 127:315–324
Ge J, Eisenberg A, Budowle B (2012) Developing criteria and data to determine best options for expanding the core CODIS loci. Investig Genet 3:1
Tomas C, Axler-DiPerte G, Budimlija ZM, Børsting C, Coble MD, Decker AE, Eisenberg A, Fang R, Fondevila M, Fredslund SF, Gonzalez S, Hansen AJ, Hoff-Olsen P, Haas C, Kohler P, Kriegel AK, Lindblom B, Manohar F, Maroñas O, Mogensen HS, Neureuther K, Nilsson H, Scheible MK, Schneider PM, Sonntag ML, Stangegaard M, Syndercombe-Court D, Thacker CR, Vallone PM, Westen AA, Morling N (2011) Autosomal SNP typing of forensic samples with the GenPlex™ HID System: results of a collaborative study. Forensic Sci Int Genet 5:369–375
Børsting C, Sanchez JJ, Morling N (2005) SNP typing on the NanoChip electronic microarray. Methods Mol Biol 297:155–168
Mengel-Jørgensen J, Sanchez JJ, Børsting C, Kirpekar F, Morling N (2005) Typing of multiple single-nucleotide polymorphisms using ribonuclease cleavage of DNA/RNA chimeric single-base extension primers and detection by MALDI-TOF mass spectrometry. Anal Chem 77:5229–5235
Budowle B, Planz J, Campbell R, Eisenberg A (2004) SNPs and microarray technology in forensic genetics: development and application to mitochondrial DNA. Forens Sci Rev 16:22–36
Budowle B (2004) SNP typing strategies. Forensic Sci Int 146(Suppl):S139–S142
Sobrino B, Brión M, Carracedo A (2005) SNPs in forensic genetics: a review on SNP typing methodologies. Forensic Sci Int 154:181–194
NuGEN (2013) Encore™ 384 Multiplex System. NuGEN. http://www.nugeninc.com/nugen/index.cfm/products/pl/library-preparation/encore-384-multiplex-system/. Accessed 30 January 2013
Wei YL, Li CX, Jia J, Hu L, Liu Y (2012) Forensic identification using a multiplex assay of 47 SNPs. J Forensic Sci 57:1448–1156
Life Technologies (2012) Ion AmpliSeq™ Library Preparation User Guide. Life Technologies, Foster City (CA)
Life Technologies (2011) Ion Library Quantitation Kit User Guide. Life Technologies, Foster City (CA)
Life Technologies (2011) Ion OneTouch™ 200 Template Kit v2 DL. Life Technologies, Foster City (CA)
Life Technologies (2011) Ion PGM™ 200 Sequencing Kit. Life Technologies, Foster City (CA)
Thorvaldsdóttir H, Robinson JT, Mesirov JP (2012) Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. doi:10.1093/bib/bbs017
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26
Davis C, Warshauer, D, Budowle B (2012) DNA profiling of database reference samples using second generation sequencing. 23rd International symposium on human identification, Nashville (TN)
Rothberg JM, Hinz W, Rearick TM, Schultz J, Mileski W, Davey M, Leamon JH, Johnson K, Milgrew MJ, Edwards M, Hoon J, Simons JF, Marran D, Myers JW, Davidson JF, Branting A, Nobile JR, Puc BP, Light D, Clark TA, Huber M, Branciforte JT, Stoner IB, Cawley SE, Lyons M, Fu Y, Homer N, Sedova M, Miao X, Reed B, Sabina J, Feierstein E, Schorn M, Alanjary M, Dimalanta E, Dressman D, Kasinskas R, Sokolsky T, Fidanza JA, Namsaraev E, McKernan KJ, Williams A, Roth GT, Bustillo J (2011) An integrated semiconductor device enabling non-optical genome sequencing. Nature 475:348–352
Voelkerding KV, Dames SA, Durtschi JD (2009) Next-generation sequencing: from basic research to diagnostics. Clin Chem 55:641–658
Keating B, Bansal AT, Walsh S, Millman J, Newman J, Kidd K, Budowle B, Eisenberg A, Donfack J, Gasparini P, Budimlija Z, Henders AK, Chandrupatla H, Duffy DL, Gordon SD, Hysi P, Liu F, Medland SE, Rubin L, Martin NG, Spector TD, Kayser M (2013) First all-in-one inference tool for DNA forensics: parallel genome-wide inference of bio-geographic ancestry, appearance, relatedness and gender with Identitas forensic chip. Int J Leg Med 127:559–572
Acknowledgments
This project was supported in part by Award No. 2012-DN-BX-K033, awarded by the National Institute of Justice, Office of Justice Programs, US Department of Justice. The opinions, findings, and conclusions or recommendations expressed in this publication are those of the author(s) and do not necessarily reflect those of the US Department of Justice. We would like to thank Life Technologies for providing its technical expertise and contributing to the sample preparation and sequencing chemistries used in this study. In addition, we would like to thank Illumina Inc for its contributions for generating the in-house SNP panel.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Seo, S.B., King, J.L., Warshauer, D.H. et al. Single nucleotide polymorphism typing with massively parallel sequencing for human identification. Int J Legal Med 127, 1079–1086 (2013). https://doi.org/10.1007/s00414-013-0879-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-013-0879-7