Abstract
Recently, several authors described the observation that RNA degradation does not correlate with the postmortem interval (PMI), but rather with other parameters like environmental impact and the circumstances of death. Therefore, the question arose if the analysis of gene expression could be a valuable tool in forensic genetics to contribute to the determination of the cause of death. In our study, six human tissues obtained from six individuals with PMI varying between 15 and 118 h were used for total RNA extraction. Quantification was performed using a GAPDH real-time assay, and the quality of mRNA was checked by amplification of different fragment lengths of the GAPDH transcript. In our set of samples, nearly all tissues in all PMI revealed satisfactory results, while skeletal muscle, followed by brain and heart, gave the best results. No correlation between PMI and RNA degradation could be detected, as very good results were observed for all tissues from the individual with the longest PMI. The highly promising results obtained in this study raise hopes that in the near future several fields of forensic investigation may profit from additional information about gene expression patterns and their correlation with pathological findings.



Similar content being viewed by others
References
Wilusz CJ, Wormington M, Peltz SW (2001) The cap-to-tail guide to mRNA turnover. Nat Rev Mol Cell Biol 2:237–246
Catts VS, Catts SV, Fernandez HR, Taylor JM, Coulson EJ, Lutze-Mann LH (2005) A microarray study of post-mortem mRNA degradation in mouse brain tissue. Brain Res Mol Brain Res 138:164–177
Preece P, Cairns NJ (2003) Quantifying mRNA in postmortem human brain: influence of gender, age at death, postmortem interval, brain pH, agonal state and inter-lobe mRNA variance. Brain Res Mol Brain Res 118:60–71
Fitzpatrick R, Casey OM, Morris D et al (2002) Postmortem stability of RNA isolated from bovine reproductive tissues. Biochim Biophys Acta 1574:10–14
Kobayashi H, Sakimura K, Kuwano R et al (1990) Stability of messenger RNA in postmortem human brains and construction of human brain cDNA libraries. J Mol Neurosci 2:29–34
Reddy PH, McWeeney S, Park BS et al (2004) Gene expression profiles of transcripts in amyloid precursor protein transgenic mice: upregulation of mitochondrial metabolism and apoptotic genes is an early cellular change in Alzheimer’s disease. Hum Mol Genet 13:1225–1240
Preece P, Virley DJ, Costandi M et al (2003) An optimistic view for quantifying mRNA in post-mortem human brain. Brain Res Mol Brain Res 116:7–16
Bauer M, Polzin S, Patzelt D (2003) Quantification of RNA degradation by semi-quantitative duplex and competitive RT-PCR: a possible indicator of the age of bloodstains? Forensic Sci Int 138:94–103
Bauer M, Gramlich I, Polzin S, Patzelt D (2003) Quantification of mRNA degradation as possible indicator of postmortem interval—a pilot study. Leg Med (Tokyo) 5:220–227
Bauer M, Patzelt D (2003) A method for simultaneous RNA and DNA isolation from dried blood and semen stains. Forensic Sci Int 136:76–78
Loddenkötter B, Becker K, Hohoff C, Brinkmann B, Bajanowski T (2005) Real-time quantitative PCR assay for the detection of Helicobacter pylori: no association with sudden infant death syndrome. Int J Legal Med 119:202–206
Bustin SA (2002) Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J Mol Endocrinol 29:23–39
Bustin SA (2000) Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J Mol Endocrinol 25:169–193
Goidin D, Mamessier A, Staquet MJ, Schmitt D, Berthier-Vergnes O (2001) Ribosomal 18S RNA prevails over glyceraldehyde-3-phosphate dehydrogenase and beta-actin genes as internal standard for quantitative comparison of mRNA levels in invasive and noninvasive human melanoma cell subpopulations. Anal Biochem 295:17–21
Aerts JL, Gonzales MI, Topalian SL (2004) Selection of appropriate control genes to assess expression of tumor antigens using real-time RT-PCR. Biotechniques 36:84–86, 88, 90–91
Phang TW, Shi CY, Chia JN, Ong CN (1994) Amplification of cDNA via RT-PCR using RNA extracted from postmortem tissues. J Forensic Sci 39:1275–1279
Mall G, Eckl M, Sinicina I, Peschel O, Hubig M (2005) Temperature-based death time estimation with only partially known environmental conditions. Int J Legal Med 119:185–194
Henssge C, Madea B (2004) Estimation of the time since death in the early post-mortem period. Forensic Sci Int 144:167–175
Takamiya M, Saigusa K, Kumagai R, Nakayashiki N, Aoki Y (2005) Studies on mRNA expression of tissue-type plasminogen activator in bruises for wound age estimation. Int J Legal Med 119:16–21
Hayashi T, Ishida Y, Kimura A, Takayasu T, Eisenmenger W, Kondo T (2004) Forensic application of VEGF expression to skin wound age determination. Int J Legal Med 118:320–325
Suay L, Salvador ML, Abesha E, Klein U (2005) Specific roles of 5′ RNA secondary structures in stabilizing transcripts in chloroplasts. Nucleic Acids Res 33:4754–4761
Ikematsu K, Tsuda R, Nakasono I (2006) Gene response of mouse skin to pressure injury in the neck region. Leg Med (Tokyo) 8:128–131
Liu J, Lewohl JM, Dodd PR, Randall PK, Harris RA, Mayfield RD (2004) Gene expression profiling of individual cases reveals consistent transcriptional changes in alcoholic human brain. J Neurochem 90:1050–1058
Acknowledgement
The authors thank Benedikt Vennemann and Markus Große Perdekamp for help with the sample collection and Manuela Liebers for helpful discussion. This work is part of the bachelor thesis of KM.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Sequences of the forward primers (in 5′-3′ orientation) for the amplification of different lengths of the GAPDH transcript. For all amplicons, the same reverse primer (5′-tgccctgtagaaattcgttg-3′) was used. The different forward primers were chosen to span at least one exon/exon boundary. E/E Exon/Exon boundary according to GenBank NM002046 (mRNA) and J04038 (genomic sequence) (DOC 29 kb)
Supplementary Fig. 1
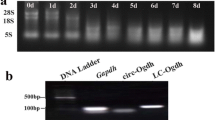
a Example of the real-time PCR results of the GAPDH assay using RNA extracted from muscle tissue of the individuals A–E (according to Table 2). b Example of a denaturing agarose gel electrophoresis using RNA extracted from skeletal muscle of individual A. The band representing 28s ribosomal RNA is stronger than the one representing 18s ribosomal RNA indicating a high quality of the extracted RNA (DOC 96 kb)
Supplementary Fig. 2
Example for the amplification of different lengths of the GAPDH transcripts for individual E. a PCR GAPDH_F1–F3. b PCR GAPDH_F4–F5 (DOC 305 kb)
Rights and permissions
About this article
Cite this article
Heinrich, M., Matt, K., Lutz-Bonengel, S. et al. Successful RNA extraction from various human postmortem tissues. Int J Legal Med 121, 136–142 (2007). https://doi.org/10.1007/s00414-006-0131-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-006-0131-9