Abstract
In vole Microtus levis, cells of preimplantation embryo and extraembryonic tissues undergo imprinted X chromosome inactivation (iXCI) which is triggered by a long non-coding nuclear RNA, Xist. At early stages of iXCI, chromatin of vole inactive X chromosome is enriched with the HP1 heterochromatin-specific protein, trimethylated H3K9 and H4K20 attributable to constitutive heterochromatin. In the study, using vole trophoblast stem (TS) cells as a model of iXCI, we further investigated chromatin of the inactive X chromosome of M. levis and tried to find out the role of Xist RNA. We demonstrated that chromatin of the inactive X chromosome in vole TS cells also contained the SETDB1 histone methyltransferase and KAP1 protein. In addition, we observed that Xist RNA did not contribute significantly to maintenance of X chromosome inactive state during iXCI in vole TS cells. Xist repression affected neither transcriptional silencing caused by iXCI nor maintenance of trimethylated H3K9 and H4K20 as well as HP1, KAP1, and SETDB1 on the inactive X chromosome. Moreover, the unique repertoire of chromatin modifications on the inactive X chromosome in vole TS cells could be disrupted by a chemical compound, DZNep, and then restored even in the absence of Xist RNA. However, Xist transcript was necessary for recruitment of an additional repressive histone modification, trimethylated H3K27, to the inactive X chromosome during vole TS cell differentiation.





Similar content being viewed by others
References
Brockdorff N (2013) Noncoding RNA and polycomb recruitment. RNA 19:429–442. https://doi.org/10.1261/rna.037598.112
Chadwick BP, Willard HF (2004) Multiple spatially distinct types of facultative heterochromatin on the human inactive X chromosome. Proc Natl Acad Sci U S A 101:17450–17455. https://doi.org/10.1073/pnas.0408021101
Chaumeil J, Okamoto I, Heard E (2004) X-chromosome inactivation in mouse embryonic stem cells: analysis of histone modifications and transcriptional activity using immunofluorescence and FISH. Methods in Enzymology:405–419
Coppola G, Pinton A, Joudrey EM et al (2008) Spatial distribution of histone isoforms on the bovine active and inactive X chromosomes. Sex Dev 2:12–23. https://doi.org/10.1159/000117715
Dementyeva EV, Shevchenko AI, Anopriyenko OV et al (2010) Difference between random and imprinted X inactivation in common voles. Chromosoma 119:541–552. https://doi.org/10.1007/s00412-010-0277-6
Dementyeva EV, Shevchenko AI, Zakian SM (2009) X-chromosome upregulation and inactivation: two sides of the dosage compensation mechanism in mammals. BioEssays 31:21–28. https://doi.org/10.1002/bies.080149
Dubois A, Deuve JL, Navarro P et al (2014) Spontaneous reactivation of clusters of X-linked genes is associated with the plasticity of X-inactivation in mouse trophoblast stem cells. Stem Cells 32:377–390. https://doi.org/10.1002/stem.1557
Elisaphenko EA, Nesterova TB, Duthie SM et al (1998) Repetitive DNA sequences in the common vole: cloning, characterization and chromosome localization of two novel complex repeats MS3 and MS4 from the genome of the East European vole Microtus rossiaemeridionalis. Chromosom Res 6:351–360. https://doi.org/10.1023/A:1009284031287
Grigor’eva EV, Shevchenko AI, Mazurok NA et al (2009) FGF4 independent derivation of trophoblast stem cells from the common vole. PLoS One 4:e7161. https://doi.org/10.1371/journal.pone.0007161
Heidenreich M, Zhang F (2016) Applications of CRISPR-Cas systems in neuroscience. Nat Rev Neurosci 17:36–44. https://doi.org/10.1038/nrn.2015.2
Iyengar S, Farnham PJ (2011) KAP1 protein: an enigmatic master regulator of the genome. J Biol Chem 286:26267–26276. https://doi.org/10.1074/jbc.R111.252569
Kalantry S, Purushothaman S, Bowen RB et al (2009) Evidence of Xist RNA-independent initiation of mouse imprinted X-chromosome inactivation. Nature 460:647–651. https://doi.org/10.1038/nature08161
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Keniry A, Gearing LJ, Jansz N et al (2016) Setdb1-mediated H3K9 methylation is enriched on the inactive X and plays a role in its epigenetic silencing. Epigenetics Chromatin 9:16. https://doi.org/10.1186/s13072-016-0064-6
Kumaki Y, Oda M, Okano M (2008) QUMA: quantification tool for methylation analysis. Nucleic Acids Res 36:W170–W175. https://doi.org/10.1093/nar/gkn294
Lee J-K, Kim K-C (2013) DZNep, inhibitor of S-adenosylhomocysteine hydrolase, down-regulates expression of SETDB1 H3K9me3 HMTase in human lung cancer cells. Biochem Biophys Res Commun 438:647–652. https://doi.org/10.1016/j.bbrc.2013.07.128
Minkovsky A, Sahakyan A, Rankin-Gee E et al (2014) The Mbd1-Atf7ip-Setdb1 pathway contributes to the maintenance of X chromosome inactivation. Epigenetics Chromatin 7:12. https://doi.org/10.1186/1756-8935-7-12
Namekawa SH, Payer B, Huynh KD et al (2010) Two-step imprinted X inactivation: repeat versus genic silencing in the mouse. Mol Cell Biol 30:3187–3205. https://doi.org/10.1128/MCB.00227-10
Nesterova TB, Duthie SM, Mazurok NA et al (1998) Comparative mapping of X chromosomes in vole species of the genus Microtus. Chromosom Res 6:41–48. https://doi.org/10.3897/CompCytogen.v5i3.1429
Nesterova TB, Slobodyanyuk SY, Elisaphenko EA et al (2001) Characterization of the genomic Xist locus in rodents reveals conservation of overall gene structure and tandem repeats but rapid evolution of unique sequence. Genome Res 11:833–849. https://doi.org/10.1101/gr.174901
Okamoto I, Arnaud D, Le Baccon P et al (2005) Evidence for de novo imprinted X-chromosome inactivation independent of meiotic inactivation in mice. Nature 438:369–373. https://doi.org/10.1038/nature04155
Okamoto I, Heard E (2006) The dynamics of imprinted X inactivation during preimplantation development in mice. Cytogenet Genome Res 113:318–324. https://doi.org/10.1159/000090848
Okamoto I, Patrat C, Thépot D et al (2011) Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature 472:370–374. https://doi.org/10.1038/nature09872
Orishchenko KE, Pavlova SV, Elisaphenko EA et al (2012) A regulatory potential of the Xist gene promoter in vole M. rossiaemeridionalis. PLoS One 7:e33994. https://doi.org/10.1371/journal.pone.0033994
Patrat C, Okamoto I, Diabangouaya P et al (2009) Dynamic changes in paternal X-chromosome activity during imprinted X-chromosome inactivation in mice. Proc Natl Acad Sci 106:5198–5203. https://doi.org/10.1073/pnas.0810683106
Ran FA, Hsu PD, Wright J et al (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308. https://doi.org/10.1038/nprot.2013.143
Rubtsov NB, Rubtsova NV, Anopriyenko OV et al (2002) Reorganization of the X chromosome in voles of the genus Microtus. Cytogenet Genome Res 99:323–329. https://doi.org/10.1159/000071611
Sado T, Sakaguchi T (2013) Species-specific differences in X chromosome inactivation in mammals. Reproduction 146:R131–R139. https://doi.org/10.1530/REP-13-0173
Schultz DC, Ayyanathan K, Negorev D et al (2002) SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16:919–932. https://doi.org/10.1101/gad.973302
Sherstyuk VV, Shevchenko AI, Zakian SM (2015) Mapping of replication origins in the X inactivation center of vole Microtus levis reveals extended replication initiation zone. PLoS One 10:e0128497. https://doi.org/10.1371/journal.pone.0128497
Shevchenko AI, Demina VV, Mazurok NA et al (2008) Extraembryonic endoderm stem cell lines from common voles of the genus Microtus. Russ J Genet 44:1280–1289. https://doi.org/10.1134/S1022795408110057
Shevchenko AI, Malakhova AA, Elisaphenko EA et al (2011) Variability of sequence surrounding the Xist gene in rodents suggests taxon-specific regulation of X chromosome inactivation. PLoS One 6:e22771. https://doi.org/10.1371/journal.pone.0022771
Shevchenko AI, Pavlova SV, Dementyeva EV, Zakian SM (2009) Mosaic heterochromatin of the inactive X chromosome in vole Microtus rossiaemeridionalis. Mamm Genome 20:644–653. https://doi.org/10.1007/s00335-009-9201-x
Takagi N, Sasaki M (1975) Preferential inactivation of the paternally derived X chromosome in the extraembryonic membranes of the mouse. Nature 256:640–642. https://doi.org/10.1038/256640a0
Vallot C, Ouimette J-F, Makhlouf M et al (2015) Erosion of X chromosome inactivation in human pluripotent cells initiates with XACT coating and depends on a specific heterochromatin landscape. Cell Stem Cell 16:533–546. https://doi.org/10.1016/j.stem.2015.03.016
Vaskova EA, Dementyeva EV, Shevchenko AI et al (2014) Dynamics of the two heterochromatin types during imprinted X chromosome inactivation in vole Microtus levis. PLoS One 9:e88256. https://doi.org/10.1371/journal.pone.0088256
Vaskova EA, Medvedev SP, Sorokina AE et al (2015) Transcriptome characteristics and X-chromosome inactivation status in cultured rat pluripotent stem cells. Stem Cells Dev 24:2912–2924. https://doi.org/10.1089/scd.2015.0204
West JD, Frels WI, Chapman VM, Papaioannou VE (1977) Preferential expression of the maternally derived X chromosome in the mouse yolk sac. Cell 12:873–882
Wutz A (2011) Gene silencing in X-chromosome inactivation: advances in understanding facultative heterochromatin formation. Nat Rev Genet 12:542–553. https://doi.org/10.1038/nrg3035
Zakharova IS, Shevchenko AI, Shilov AG et al (2011) Histone H3 trimethylation at lysine 9 marks the inactive metaphase X chromosome in the marsupial Monodelphis domestica. Chromosoma 120:177–183. https://doi.org/10.1007/s00412-010-0300-y
Acknowledgments
We thank S.I. Bayborodin for the technical assistance.
Funding
The work was funded by the Russian Academy of Sciences (program II.2P/VI.60-1 No 0324-2016-0032 and the Russian Foundation for Basic Research (grant No. 15-04-03947a).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Fig. S1
A) SETDB1 distribution on metaphase spreads of vole XY TS cells. X and Y chromosomes are shown by arrows. B) HP1-gamma (green) and H4K20me3 (red) in nuclei of vole trophoblast cells after two weeks of differentiation. Metaphase chromosomes and nuclei were counterstained with DAPI (blue). MS4 is a DNA repeat specific for a giant telomeric block of constitutive heterochromatin on X chromosome of M. levis. (GIF 116 kb)
Fig. S2
Chromatin modifications on Xi in DZNep treated and untreated vole TS cells. MS4 repeat marks presence of two X chromosomes in nuclei of DZNep treated TS cells. (GIF 185 kb)
Fig. S3
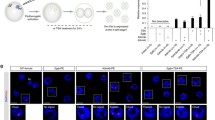
DZNep treatment does not affect the level of Xist expression in vole TS cells. Relative level of Xist expression was examined by semi-quantitative RT-PCR. Blue bars represent untreated TS cell line, orange bars do the TS cell treated with DZNep. Xa – / Xi +, the original TS cell line, R2, and TS cell clones without the deletion; Xa Δ / Xi +, TS cell clones with the deletion in the Xist promoter and normal Xist expression; Xa – / Xi Δ, TS cell clones with the deletion in the Xist promoter and depleted Xist expression. R1, XO TS cell line. (GIF 71 kb)
Fig. S4
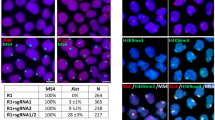
A) RNA FISH detecting monoallelic Hprt1 expression from Xa in XX TS cell nuclei with presence and absence of Xist RNA on Xi. Xi is enriched with H3K9me3. Xa – / Xi +, Xist is properly expressed; Xa – / Xi Δ, Xist is repressed. Scale bar is 10 μm. В) RNA FISH signals of monoallelically expressed Pgk1 (red) and Hprt1 (green) were revealed in a close proximity within nucleus. IF, immunofluorescent staining. (GIF 512 kb)
Table S1
Antibodies used in the study (DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Shevchenko, A.I., Grigor’eva, E.V., Medvedev, S.P. et al. Impact of Xist RNA on chromatin modifications and transcriptional silencing maintenance at different stages of imprinted X chromosome inactivation in vole Microtus levis . Chromosoma 127, 129–139 (2018). https://doi.org/10.1007/s00412-017-0650-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-017-0650-9