Abstract
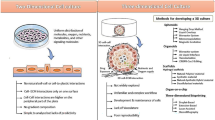
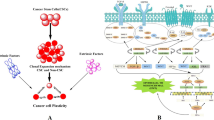
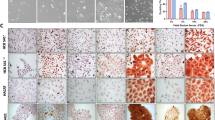
Three-dimensional (3D) melanoma culture is a personalized in vitro model that can be used for high-fidelity pre-clinical testing and validation of novel therapies. However, whether the genomic landscape of 3D cultures faithfully reflects the original primary tumor which remains unknown. The purpose of our study was to compare the genomic landscapes of 3D culture models with those of the original tumors. Patient-derived xenograft (PDX) tumors were established by engrafting fresh melanoma tissue from each patient. Then, a 3D culture model was generated using cryopreserved PDX tumors embedded in pre-gelled porcine skin decellularized extracellular matrix with normal human dermal fibroblasts. Using whole-exome sequencing, the genomic landscapes of 3D cultures, PDX tumors, and the original tumor were compared. We found that 91.4% of single-nucleotide variants in the original tumor were detected in the 3D culture and PDX samples. Putative melanoma driver mutations (BRAF p.V600E, CDKN2A p.R7*, ADAMTS1 p.Q572*) were consistently identified in both the original tumor and 3D culture samples. Genome-wide copy number alteration profiles were almost identical between the original tumor and 3D culture samples, including the driver events of ARID1B loss, BRAF gain, and CCND1 gain. In conclusion, our study revealed that the genomic profiles of the original tumor and our 3D culture model showed high concordance, indicating the reliability of our 3D culture model in reflecting the original characteristics of the tumor.



Similar content being viewed by others
Data availability
Raw sequencing data were deposited in the Sequence Read Archive (SRA) database (Project ID: PRJNA813613).
References
Welch HG, Mazer BL, Adamson AS (2021) The rapid rise in cutaneous melanoma diagnoses. N Engl J Med 384:72–79
Curti BD, Faries MB (2021) Recent advances in the treatment of melanoma. N Engl J Med 384:2229–2240
Switzer B, Puzanov I, Skitzki JJ, Hamad L, Ernstoff MS (2022) Managing metastatic melanoma in 2022: a clinical review. JCO Oncol Practice 18:335–351
Alzeeb G, Metges JP, Corcos L, Le Jossic-Corcos C: Three-Dimensional Culture Systems in Gastric Cancer Research. Cancers (Basel) 2020, 12.
Hidalgo M, Amant F, Biankin AV, Budinská E, Byrne AT, Caldas C, Clarke RB, de Jong S, Jonkers J, Mælandsmo GM et al (2014) Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer Discov 4:998–1013
Herter-Sprie GS, Kung AL, Wong KK (2013) New cast for a new era: preclinical cancer drug development revisited. J Clin Invest 123:3639–3645
LeBlanc VG, Trinh DL, Aslanpour S, Hughes M, Livingstone D, Jin D, Ahn BY, Blough MD, Cairncross JG, Chan JA et al (2022) Single-cell landscapes of primary glioblastomas and matched explants and cell lines show variable retention of inter- and intratumor heterogeneity. Cancer Cell 40:379-392.e379
Powley IR, Patel M, Miles G, Pringle H, Howells L, Thomas A, Kettleborough C, Bryans J, Hammonds T, MacFarlane M, Pritchard C (2020) Patient-derived explants (PDEs) as a powerful preclinical platform for anti-cancer drug and biomarker discovery. Br J Cancer 122:735–744
Templeton AR, Jeffery PL, Thomas PB, Perera MPJ, Ng G, Calabrese AR, Nicholls C, Mackenzie NJ, Wood J, Bray LJ et al (2021) Patient-derived explants as a precision medicine patient-proximal testing platform informing cancer management. Front Oncol 11:767697
Jeong YM, Bang C, Park M, Shin S, Yun S, Kim CM, Jeong G, Chung YJ, Yun WS, Lee JH, Jin S: 3D-Printed Collagen Scaffolds Promote Maintenance of Cryopreserved Patients-Derived Melanoma Explants. Cells 2021, 10.
Pati F, Jang J, Ha DH, Won Kim S, Rhie JW, Shim JH, Kim DH, Cho DW (2014) Printing three-dimensional tissue analogues with decellularized extracellular matrix bioink. Nat Commun 5:3935
Ahn G, Min KH, Kim C, Lee JS, Kang D, Won JY, Cho DW, Kim JY, Jin S, Yun WS, Shim JH (2017) Precise stacking of decellularized extracellular matrix based 3D cell-laden constructs by a 3D cell printing system equipped with heating modules. Sci Rep 7:8624
Kim BS, Kwon YW, Kong JS, Park GT, Gao G, Han W, Kim MB, Lee H, Kim JH, Cho DW (2018) 3D cell printing of in vitro stabilized skin model and in vivo pre-vascularized skin patch using tissue-specific extracellular matrix bioink: A step towards advanced skin tissue engineering. Biomaterials 168:38–53
Won JY, Lee MH, Kim MJ, Min KH, Ahn G, Han JS, Jin S, Yun WS, Shim JH (2019) A potential dermal substitute using decellularized dermis extracellular matrix derived bio-ink. Artif Cells Nanomed Biotechnol 47:644–649
Park W, Gao G, Cho DW: Tissue-Specific Decellularized Extracellular Matrix Bioinks for Musculoskeletal Tissue Regeneration and Modeling Using 3D Bioprinting Technology. Int J Mol Sci 2021, 22.
Kim YS, Shin S, Jung SH, Park YM, Park GS, Lee SH, Chung YJ (2022) Genomic progression of precancerous actinic keratosis to squamous cell carcinoma. J Invest Dermatol 142:528-538.e528
Van der Auwera GA OCB: Genomics in the Cloud: Using Docker, GATK, and WDL in Terra (1st Edition). O'Reilly Media; 2020.
Okonechnikov K, Conesa A, García-Alcalde F (2015) Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32:292–294
Shen R, Seshan VE (2016) FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res 44:e131
Jo SY, Kim E, Kim S (2019) Impact of mouse contamination in genomic profiling of patient-derived models and best practice for robust analysis. Genome Biol 20:231
Yang H, Wang K (2015) Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc 10:1556–1566
Brash DE (2015) UV signature mutations. Photochem Photobiol 91:15–26
Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO, Whitwham A, Keane T, McCarthy SA, Davies RM, Li H: Twelve years of SAMtools and BCFtools. Gigascience 2021, 10.
Hodis E, Watson IR, Kryukov GV, Arold ST, Imielinski M, Theurillat JP, Nickerson E, Auclair D, Li L, Place C et al (2012) A landscape of driver mutations in melanoma. Cell 150:251–263
Hayward NK, Wilmott JS, Waddell N, Johansson PA, Field MA, Nones K, Patch AM, Kakavand H, Alexandrov LB, Burke H et al (2017) Whole-genome landscapes of major melanoma subtypes. Nature 545:175–180
Zack TI, Schumacher SE, Carter SL, Cherniack AD, Saksena G, Tabak B, Lawrence MS, Zhsng CZ, Wala J, Mermel CH et al (2013) Pan-cancer patterns of somatic copy number alteration. Nat Genet 45:1134–1140
Ben-David U, Ha G, Tseng Y-Y, Greenwald NF, Oh C, Shih J, McFarland JM, Wong B, Boehm JS, Beroukhim R, Golub TR (2017) Patient-derived xenografts undergo mouse-specific tumor evolution. Nat Genet 49:1567–1575
Acknowledgements
This study was supported by grants from National Research Foundation of Korea (2020R1C1C11011296, RS-2022-00165497, and 2019R1A5A2027588). This research was supported by a grant of the MD-Phd/Medical Scientist Training Program through the Korea Health Industry Development Institue (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea. The biospecimens for this study were provided by The Biobank of Seoul St. Mary's Hospital, the Catholic University of Korea.
Author information
Authors and Affiliations
Contributions
Conception and design: Y.-J. Chung, C.-H. Bang Development of methodology: S. Jin, Y.-J. Chung, C.-H. Bang Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): All authors Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): All authors Writing, review, and/or revision of the manuscript: Y.-S. Kim, Y.-J. Chung, C.-H. Bang Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): Y.-J. Chung, C.-H. Bang Study supervision: Y.-J. Chung, C.-H. Bang
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, YS., Park, M., Jin, S. et al. Genomic comparison between an in vitro three-dimensional culture model of melanoma and the original primary tumor. Arch Dermatol Res 315, 1225–1231 (2023). https://doi.org/10.1007/s00403-022-02502-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00403-022-02502-4