Abstract
Fusobacterium nucleatum is supposed to play a critical role in the development of colorectal cancer. The species has also been associated with ulcerative colitis (UC) that can progress into colorectal cancer, however, the involvement of bacteria in this process remains unclear. We analysed 177 colon biopsies obtained from patients during screening, including 20 healthy controls, 56 UC cases and 69 cases at different stages of progression to colitis-associated cancer (CAC); 32 samples of sporadic colorectal carcinoma (sCRC) were also included. The presence of F. nucleatum was detected by quantitative real-time PCR (qPCR). Our data show an association between the presence of the bacteria and the progression of carcinogenesis in UC patients. In 39.5% of CAC samples F. nucleatum was detected, compared to only 1.8% in UC cases. The bacteria were detected in 6.3% of samples with initial neoplastic transformation, so-called low-grade dysplasia (LGD), whereas high-grade dysplasia (HGD) resulted in 33.3% of samples positive for F. nucleatum. The fraction of F. nucleatum-positive samples from sCRC cases was 56.3%, which was not significantly different to the CAC group. We conclude that F. nucleatum is associated with the occurrence and progression of colon carcinogenesis, rather than with UC itself.
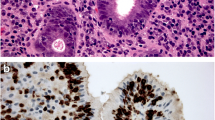
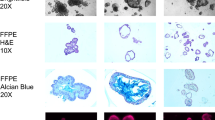
Similar content being viewed by others
Introduction
Ulcerative colitis (UC) is a common form of inflammatory bowel disease (IBD) that is characterized by chronic inflammation of the colon mucosa. The inflammatory process starts from the rectum and spreads continuously to the colon and can even reach the terminal ileum. Clinical characteristics of UC include abdominal pain, often accompanied by bloody diarrhoea [1]. In 2017, the worldwide prevalence of UC was reported as 84.3 per 100,000 people, with by a death rate of 0.51 per 100,000 [2]. The most recent data from Germany are from 2019, when the UC incidence rate was 36 per 100,000, with a prevalence of 529 per 100,000 [3].
Patients suffering from UC have an increased risk of developing colorectal cancer (CRC) [4]. The carcinogenesis of this colitis-associated cancer (CAC) follows a typical sequence of inflammation-dysplasia-carcinoma [5]. The progression starts with the onset of UC-induced inflammation of the colon mucosa, which over time may lead to neoplasia. Precancerous stages of low-grade dysplasia (LGD) and high-grade dysplasia (HGD) can be recognized [6], although occasionally UC can directly progress towards CAC without a precancerous stage [7].
The distinction between CAC that is directly caused by UC and CRC that is independent of UC but initiated by sporadic somatic mutations (sCRC) can be difficult, based on histology only; likewise, the corresponding precancerous lesions are not always easy to diagnose [8]. However, a correct diagnosis is important as it has a significant impact on treatment. If an area in the colon is diagnosed with sporadic lesions, its endoscopic removal is considered curative and in CRC patients without UC a partial colectomy is considered, while CAC or HGD is recommended to be treated by proctocolectomy [9]. So far, the exact molecular mechanisms of pathogenesis for the development of CAC have not been clarified, however, it has been suggested that a genetic predisposition of the host, diet and lifestyle, and the gut microbiome may all play a role.
The human intestine is densely populated with approximately 1012 microorganisms per gram content [10]. Some components of this microbiota can be involved in inflammatory processes or can cause DNA damage leading to cell death, and as such they may have an impact on CRC development [11]. In 2012, two independent research groups described an increased occurrence of Fusobacterium nucleatum in cancerous intestinal tissue [12, 13], and since then, the impact of F. nucleatum in the development or progress of CRC has been substantiated [12,13,14,15,16,17]. However, the underlying mechanisms that lead to the progression of carcinogenesis caused by this bacterial species remain unclear. It has been suggested that F. nucleatum might cause damage to the epithelial barrier of the colon [18], induce DNA damage of intestinal mucosal cells [19] or lead to dysbiosis of the gut microbiota [20].
F. nucleatum is a Gram-negative, obligate anaerobe bacterium typically residing in the oral cavity [21]. It is considered pathogenic when it is involved in dental plaque formation, where it is able to attract other bacterial species [22]. The bacteria have occasionally been detected in various other organs as well, including in the placenta and in foetal tissue [23], in the brain [24] and in the liver [25]. In mice, oral fusobacteria were hematogenously transferred to the placenta and this was associated with stillbirth [26]. When F. nucleatum is present in the colon, this is associated with IBD and this corelated with its invasive potential [27].
The pathogenicity of F. nucleatum is related to its ability to attach to and invade into epithelial cells [27, 28], enabled by virulence factor FadA that is expressed on the bacterial surface [29, 30]. FadA acts as an adhesin and binds to the protein E-cadherin present on host cells at adherens junctions. This activates transcription factor β-catenin signalling, a pathway that is also involved in cell proliferation during carcinogenesis. It has been shown that FadA levels are significantly increased in colon tissue from patients with adenomas and adenocarcinomas, and higher FadA levels in CRC tissue correlate with increased expression of oncogenic and inflammatory genes [14]. Increased expression of cytokines such as IL-6 and IL-1β in patients with an F. nucleatum infection was described that could drive the local inflammatory response [18, 31, 32].
Although a number of studies reported a correlation between the occurrence of F. nucleatum and CRC [33,34,35,36], published data on its involvement in IBD or UC have been contradictory. A Canadian study described the occurrence of F. nucleatum in 50% of IBD patients and suggested that F. nucleatum could serve as a possible biomarker for IBD [27]. The bacteria were also detected in 39% of UC patients from China with a similar detection rate of 37.14% in CRC patients, indicating an association between F. nucleatum and UC as well as CRC [28]. In contrast, a Japanese study found F. nucleatum in only 6.3% of the analysed UC patients [37]. It was reported that the number of colonic F. nucleatum bacteria correlates with a shorter survival in CRC cases [17] and that high numbers aggravate the course of UC by damaging the intestinal barrier and promoting inflammation [18]. Furthermore, F. nucleatum was shown to be associated with resistance to chemotherapy [38, 39].
This conflicting literature concerning the association of F. nucleatum with UC and CRC led to the present study, which aimed to establish the occurrence of qPCR-detected F. nucleatum in German patients with UC, with emphasis on the different stages of CAC carcinogenesis.
Material and Methods
Patient Groups and Sample Collection
In this study, consecutive, retrospective samples of 177 patients that were treated endoscopically or surgically at the Klinikum Bayreuth between 2006 and 2020 were analysed. Biopsies and the surgical resection material were sent to the Institute of Pathology of the Klinikum Bayreuth in Germany for histopathological assessment. UC-associated dysplasia (LGD, HGD) and cancerous lesions (CAC, sCRC) were diagnosed according to common criteria and guidelines [9, 40]. All diagnostic results were confirmed by two independent pathologists with consistent outcomes. The study was approved by the ethics committee of the University Bayreuth (#O 1305/1-GB).
The biopsies (N = 177) were fixed in 4% neutral pH-buffered formalin. Tissue samples were dehydrated and paraffinized using a HistoCore PELORIS 3 (Leica Biosystems, Germany). Formalin-fixed paraffin-embedded (FFPE) blocks were cut into 4-µm-thick slices, followed by hematoxylin–eosin staining carried out on a Tissue-Tek Prisma (Sakura Finetek, Japan) for histopathological diagnosis. According to the diagnosis, the patients were categorized in six groups, with group 1: healthy colon (n = 20), group 2: UC (n = 56), group 3: LGD (n = 16), group 4: HGD (n = 15), group 5: CAC (n = 38) and group 6: sCRC (n = 32).
The FFPE blocks were microdissected to 5 µm and genomic DNA was extracted with the Maxwell LEV Blood DNA Kit (Promega) on a Maxwell 16 instrument. The DNA was quantified with the QuantiFluor dsDNA System Kit (Ref. E4871) in combination with a Quantus Fluorometer (Promega).
As a positive control, a bacterial strain of Fusobacterium nucleatum, subsp. nucleatum (DSM 15643) was obtained from the Leibnitz-Institute, (Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Braunschweig, Germany). The freeze-dried strain was resolved in thioglycollate medium enriched with vitamin K1 and Hemin (Becton Dickinson Ref. 221,788) by colleagues from the Institute for Laboratory Medicine (ILM, Klinikum Bayreuth) and grown under anaerobe conditions using GasPak EZGas (Becton Dickinson, Ref. 260,683) for 48 h at 35 °C. The culture was then incubated on Schaedler Agar with vitamin K1 and 5% sheep blood (Becton Dickinson, Ref. PA-254084). Colonies were transferred to 1 mL 4% formalin and centrifuged for 10 min at 300 RPM. The pelleted bacteria were embedded in paraffin and treated as the FFPE blocks as described above to extract bacterial DNA. All DNA preparations were stored at − 20 °C.
Quantitative Real-Time Polymerase-Chain Reaction (qPCR)
A unique sequence within the F. nucleatum nusG gene was used as the target sequence for qPCR [12]. The used primer sequences were taken from the literature [34], with forward primer 5'-CAACCATTACTTTAACTCTACCATGTTCA and reverse primer 5'-GTTGACTTTACAGAAGGAGATTATGTAAAAATC. The human gene slco2a1 was used as the internal control, with forward primer 5'-ATCCCCAAAGCACCTGGTTT and reverse primer 5'-AGAGGCCAAGATAGTCCTGGTAA. Reactions were performed in 20 µL containing 1 × LightCycler FastStart DNA Master SYBR Green I Kit (Roche, Penzberg), 3 mM MgCl2 and 0.1 µM of each primer in nuclease free, PCR-grade water (Roche, Penzberg). As a template, 0.3–100 ng of FFPE-isolated DNA was used. The positive control contained 20 pg of isolated F. nucleatum DNA and nuclease free, PCR-grade water was used as the negative control. The qPCR reactions were performed with a Cobas LightCycler z480 (Roche, Penzberg) using the following conditions: denaturation for 10 min at 95 °C, 50 cycles of 95 °C for 5 s, 65 °C for 10 s and 72 °C for 6 s, followed by a temperature gradient from 37 to 95 °C with a ramp rate of 0.06 °C/s for melting curve analysis. Melting curves were evaluated with the LightCycler 480 SW Software version 1.5.1.62.
Produced amplicons were checked by agarose gel electrophoresis to confirm their length of 112 bp and 74 bp for nusG and slco2a1, respectively. The correctly sized bands were excised, and the amplified DNA was purified with the QIAquick Nucleotide Removal Kit (Qiagen, Hilden) and sequenced (Eurofins GATC, Köln) using the forward primers of nusG or slco2a1, respectively. The obtained sequences were compared to the corresponding GenBank entries (accession number of nusG from F. nucleatum: AE009951.2; accession number of human control gene slco2a1: NC_000003.12) using BLAST.
Establishment, Precision and Limit of Detection by qPCR
To verify that the nusG and slco2a1 targets are suitable for simultaneous amplification in a multiplex qPCR, the melting temperatures of both products were determined in separate reactions first, for which 20.4 ng isolated F. nucleatum DNA and 24 ng of isolated human DNA, respectively, was used. Reactions were carried out in quadruples to assess intraassay replication. For detection of interassay precision, both products were co-amplified in three independent experiments using the same DNA template samples.
The limit of detection (LOD) of the qPCR analysis was determined with serial dilutions of isolated F. nucleatum DNA (range 6.7–0.2 ng) that were complemented with a constant amount (24 ng) of human DNA to create a consistent background for all reactions (Table S1).
Statistics
Fisher’s exact test was used to investigate correlations between the detection of F. nucleatum and the different stages of carcinogenesis. For groups with a sample size > 5, the Pearson’s Correlation Coefficient was used. Statistical analyses were carried out SPSS Statistics Version 23.0.0.0. Power analysis was conducted with G*Power Version 3.1.9.7 (One-sided Fisher’s exact test, power = 0.80, α-error: 0.05) to detect the required sample sizes of all six groups (Table S2).
Results
qPCR Design and Optimization
The presence of F. nucleatum in biopsies from patients with UC at different stages of carcinogenesis was investigated by qPCR. For this, we performed multiplex qPCR to simultaneously amplify the nusG gene as a qualitative detection marker for the presence of F. nucleatum and of slco2a1, a human gene that served as the internal control. First, we established the melting curves of both products in separate reactions. Based on four replicate experiments, the amplicon of nusG had a melting temperature (Tm) of 77.67 ± 0.10 °C, whereas the Tm of the slco2a1 product was 80.29 ± 0.10 °C, indicating a high intraassay precision. The results of the four replicates are summarized in Table S3. The difference of 2.62 °C between the Tm of both products was considered sufficient for a multiplex PCR approach. To determine the interassay precision, nusG and slco2a1 were co-amplified in triplicate, using the same DNA as the template. In this multiplex assay, we observed slightly lower melting temperatures for both products compared to separate amplification: the Tm of the nusG product was now 76.66 ± 0.04 °C and the Tm of the slco2a1 amplicon was 80.17 ± 0.10 °C (see Table S4 for the individual experiments). Both melting temperatures showed high reproducibility, and their difference had even increased, to 3.51 °C on average, which was sufficient for multiplex amplification and detection.
Because our aim was to detect F. nucleatum in samples containing human DNA, a high background of the latter could be expected. Therefore, we determined the limit of detection (LOD) of F. nucleatum DNA by spiking a dilution series of bacterial DNA in the presence of a constant, high background of human DNA (Table S1). Even at the lowest tested concentration, corresponding to a bacterial to human DNA ratio of 0.8%, F. nucleatum DNA was still detected. This corresponded with a limit of detection of 10 pg µl−1 for F. nucleatum.
The PCR products obtained in the single and multiplex assays were validated by gel electrophoresis (Fig. 1). This resulted in amplicons sized 74 bp and 112 bp, for slco2a1 and nusG amplicons, respectively.
Agarose gel electrophoresis of the qPCR products obtained during separate amplification (A) and co-amplification (B) of bacterial nusG and human slco2a1. A M: 100 bp size marker, S1 and S2: F. nucleatum DNA amplified with nusG primers (positive control), S3: human DNA amplified with slco2a1 primers, S4: negative control in the presence of both primer pairs. Lane 5 is empty. B M: 100 bp size marker, S5: amplification of an F. nucleatum-negative human sample with nusG primers only, S6: co-amplification of an F. nucleatum-negative sample with both primer pairs, S7: co-amplification of an F. nucleatum-positive sample with both primer pairs. Arrows indicate the positions of the nusG and slco2a1 amplicons
To confirm that the amplicons were indeed produced from the target genes, the bands of samples S1 and S3 were excised from the agarose gel (Fig. 1) and after purification the DNA was sequenced using the corresponding forward primers utilized for qPCR amplification. When the DNA sequence obtained from sample S1 was compared to a GenBank entry of nusG (Accession number AE009951.2) it aligned with 92% identity. Alignment of the sequence from sample S3 with a GenBank entry of slco2a1 (Accession number NC_000003.12) revealed a sequence identity of 95%. These results proved specificity of the used primers for both genes, producing amplicons of the correct length for both targets.
Melting point analysis was next performed for all 177 human samples by multiplex PCR. Examples of the melting curves obtained with F. nucleatum-positive and F. nucleatum-negative samples, respectively, are shown in Fig. 2. Based on the mean results obtained from all samples, a sample was considered positive for the presence of F. nucleatum if a peak was present at 76.21 °C ± 0.53 in addition to the peak from the internal control of human slco2a1 present at 80.32 °C ± 0.33. Isolated F. nucleatum DNA was included as positive control in each amplification series. This resulted in mean Tm values of 77.71 °C ± 0.17. These data imply consistent and reproducible melting temperatures throughout all samples. A total of 40 samples were found positive for the presence of F. nucleatum, while 137 samples were negative.
Melting point analysis in a multiplex qPCR for detection of F. nucleatum in the presence of human DNA in FFPE samples. The presence of bacteria was indicated by amplification of the F. nucleatum-specific nusG gene with a melting temperature of 76.21 °C ± 0.53 and the absence of PCR inhibition was confirmed with the internal standard of the human slco2a1 gene (melting temperature: 80.32 °C ± 0.33). A shows the melting curve obtained with an F. nucleatum-negative sample and B that of a F. nucleatum-positive sample
Prevalence of F. nucleatum in Colon Biopsies from Patient Groups
The results of the qPCR were analysed per patient group, to investigate the association between the presence of F. nucleatum and either UC, CAC or sCRC (Table 1). The melting curve data are summarized per group in Table S5. The bacteria were not detected in samples from healthy controls and their prevalence strongly differed between groups of patients with different pathology. Whereas F. nucleatum was only detected in 1.8% of patients diagnosed with UC (n = 56), one-third or more of patients with HGD (n = 15) or CAC (n = 38) were found positive (Table 1). Figure 3 summarizes the prevalence of F. nucleatum for each group. Compared to group 2 of UC patients, the higher prevalence of F. nucleatum-positive samples in group 4 (HGD) and 5 (CAC) was highly significant (p = 0.001 and 0.000, respectively). Significant differences were also detected between group 3 (LGD) and either group 4 or 5 (p = 0.021 and 0.000, respectively). The results implicate that patients presenting with HGD already more often had F. nucleatum in their colon than patients with UC or LGD. The prevalence of F. nucleatum in patients with HGD and CAC was comparable (p = 0.761). Interestingly, the bacteria were detected at a similar prevalence in CAC and sCRC samples (p = 0.230).
The results for each group were analysed for patient’s sex and age (Table 1). Whereas 28% of female patients were found positive, only 18% of male patients were positive (Table 1). Thus, females were about 1.5 times more likely to be positive for the bacteria than males, but this difference was not significant (p = 0.206). The results for sex within the patient groups are summarized in Fig. 4. Since no positive samples were obtained from the healthy controls of group 1 and only one of the 56 patients in group 2 was positive for F. nucleatum, these two groups were omitted here. Concentrating on groups 3 to 6, a total of 45% female and 55% male patients were found positive for F. nucleatum. In 28% of samples obtained from females and in 19% of samples from males, F. nucleatum was detected. The most remarkable differences between male and female patients were seen in group 4 (HGD with 50% prevalence in females versus 22% in males) and group 6 (sCRC, 65% versus 42%, Fig. 4A), but this difference was not statistically significant (HGD: p = 0.329; sCRC: p = 0.277). Interestingly, about half of the analysed patients for both sexes in group 5 presenting with CAC (male: 40%, 10/25; female: 39%, 5/13) were infected with F. nucleatum (p = 1.000), which differs from the results obtained for groups 4 and 6. A comparison of patients with regard to the sex in groups 5 and 6 revealed no statistical relevance (female: p = 0.169, male: p = 1.000, respectively). Together, these results underline that the infection is not correlating with the sex nor with the type of cancer in the investigated patients.
Since patients with UC are usually quite young, with an initial manifestation of UC between the age of 20 and 30 [41], we further analysed the data according to the patients’ age. For this, the patients were divided into five different subgroups of < 21 years (y), 21–40 years, 41–60 years, 61–80 years and > 80 years. The highest prevalence of F. nucleatum was found in samples from patients 41–60 years (21%) and 61–80 years (27%), and as many as 5 of 8 samples obtained from > 80-year-old patients were positive (Table 1). Ignoring the clinical presentation, a statistically significant difference was found in F. nucleatum prevalence between patients up to 40 y and those older (p = 0.011).
Figure 4B summarizes the data for subgroups 41–60 years and 61–80 years only per clinical manifestation; the other age subgroups had too few samples. A slight increase in prevalence was observed, in both age groups, going from LGD to HGD, CAC and then sCRC. While none of the patients with LGD were positive for F. nucleatum, that fraction increased to 25% (1/4) and 42% (8/19) in HGD and CAC samples, respectively (p = 0.444, 0.130). A similar increase was found in the age group of 61–80 years, where the fraction of positive patients was the lowest in group 3 with LGD (10%, 1/10) and the highest (40%, 6/15) in group 5 with CAC, which was not statistically different (p = 0.303, 0.107, respectively).
These findings all suggest that F. nucleatum prevalence increases the closer the patient is to the diagnosis of CAC. We point out that, of all clinical groups, samples from sCRC cases resulted in the highest prevalence of F. nucleatum (Fig. 3), for both sexes (Fig. 4A) and in three age subgroups: sCRC samples resulted in a prevalence of 50% (4/8) at 21–40 years, 47% (9/19) at 41–60 years and all four samples from > 80 years patients were positive. The frequencies of positive samples were similar between the CAC and sCRC groups, not resulting in a significant difference (Fig. 3), and the distribution of the positive samples in the age groups 41–60 years and 61–80 years within the groups 3–5 was similar.
Discussion
F. nucleatum can be involved in opportunistic infections and a role in CRC has been speculated, however, the species is a common member of the oral microbiota and can also have benign relationships with the host [21]. A recent study reported the association of F. nucleatum DNA in stool samples from IBD patients [19]. In mouse models, F. nucleatum had an impact on the intestinal inflammation for dextran sodium sulphate (DSS)-induced colitis, achieved by causing damage of the epithelial barrier [18, 42]. In our study, colon biopsies of 177 German patients presenting with various clinical pathologies were examined for the presence of F. nucleatum. Our data show that the incidence of F. nucleatum is not only increased in CAC cases compared to controls, but also in the precancerous stages LGD and in particular HGD. The prevalence of the bacteria was significantly higher in HGD and CAC compared to tissue samples from cases of LGD or UC. From this, we conclude that F. nucleatum infection does not represent an early event in CAC carcinogenesis. This implies that F. nucleatum may not be useful as an early diagnostic marker for CRC development, however, its presence may indicate a progression in malignant transformation. A metagenomics study analysing human faeces indicated that F. nucleatum possibly acts as a key factor for the initiation of disorders, implying that this bacterium might be a useful early biomarker for CRC development [43].
Our results from the studied German cohort of patients corroborated data from international studies. It has been demonstrated in patients from North America [12, 34], South America [44], Asia [35, 36, 45] and Europe [33, 46] that F. nucleatum is overabundant in tumorous colon samples from tissue or stool, compared to adequately matched healthy controls. Still, there are differences within literature reports in the frequency of F. nucleatum detection. Whereas a Chinese study identified 87.13% of CRC samples positive for the bacteria [36], a Japanese study [47] and two studies from the USA reported positivity for CRCs in only 8.7% and 13%, respectively [17, 34]. Compared to those findings, our data included 70 cases of CRC (CAC and sCRC combined) of which 47.1% (33/70) were found positive, which places our German cohort somewhere in the middle of the reported values from other studies. Differences between studies may be due to the method of detection and the nature of the samples. A recent study demonstrated the impact of the sample type on F. nucleatum detection by qPCR [48]: while F. nucleatum was detected in 23% of CRCs from fresh-frozen tissues, only 5.8% of CRC-derived FFPE tissues were positive. In this respect, our reported prevalence, detected in FFPE samples, is eight times higher, possibly related with the high sensitivity and low limit of detection that was achieved.
A Chinese study reported a prevalence of F. nucleatum in only 6.3% (4/64) of examined UC samples [36]. Our data confirm that the bacteria are not common in the colonic mucosa of patients with UC, as we detected it in less than 2% (1/56) of the UC samples. The single positive sample was obtained from a patient suffering from UC at a highly active state, which could mean that F. nucleatum had been attracted by the active inflammation of this area, or F. nucleatum had induced the active inflammation.
Considering the initial stage of neoplasia (LGD), we observed a slightly higher prevalence of the bacteria (6.3%, 1/16) than in the UC group, but the difference was not significant. There was, however, a significant increase in the number of positive samples in HGD cases, which represent the precancerous stage to CAC (Fig. 3). We conclude from this that the presence of F. nucleatum is more likely associated with the progression of carcinogenesis, rather than the inflammation of UC itself. It is still possible that this species directly or indirectly triggers the development of carcinoma, perhaps at a later stage than LGD. This is supported by the results we obtained from sCRC samples, which produced the highest prevalence (56.3%) of F. nucleatum. Because this type of carcinoma does not involve chronic inflammation of the colon, the high prevalence of the bacteria here indicates that UC is not required for bacterial colonization by F. nucleatum in a cancerous colon. Rather, the fact that F. nucleatum can be detected in increasing fractions during progression to CRC suggests that F. nucleatum is involved in carcinogenesis. However, it remains unclear, to what part the bacterium contributes to the pathogenic cascade leading to cancerogenesis, and to reveal this, future studies are required.
The observed increase in the fraction of F. nucleatum-positive samples within the cascade of progression towards CAC implicates that the pathogen may represent a central factor within the pathways leading to carcinogenesis. A previous European study analysing CRC in general and colorectal adenoma (CRA) detected F. nucleatum with results comparable to our data [33]. While the presence of F. nucleatum was significantly higher in CRC and HGD samples compared to normal tissue, this was not found in CRA samples [33]. Similar results were also obtained in an independent study that involved a smaller sample size, but again reported a higher prevalence of F. nucleatum in CRCs compared to matching healthy individuals [49]. That study focussed on CRA as the precancerous stage of CRC, and the authors postulated that patients with high abundance of F. nucleatum are more likely to develop CRA, indicating an association of these bacteria with development of CRC [49].
A limitation of our study is the relatively small number of patients with LGD or HGD in our cohort, and further analyses should be conducted with expanded sample sizes for each group. This would increase the statistical power of analysis, thereby possibly fortifying our conclusion that F. nucleatum is probably not a suitable early diagnostic marker for cancer development.
Methods based on qPCR are commonly used for pathogen detection, as it can reach high sensitivity and specificity. The usage of intercalating dyes such as SYBR Green results in a cost-effective method, but it can lead to unspecific binding to dsDNA. In addition, unspecific amplification of by-products can contribute to the fluorescent signal. To avoid erroneous results, we applied strict measures to avoid contamination, included an internal standard to detect PCR inhibition and verified the produced amplicons to ensure the target sequences were correctly detected. When such measures are taken, qPCR is a reliable detection method. Nevertheless, the results strongly depend on the type of material used. DNA isolated from fresh-frozen tissue is more suitable for qPCR analysis than DNA from FFPE material [48]. Unfortunately, in pathology FFPE represents the standard tissue type, as it clearly is advantageous with easy sample handling and suitability for long-time storage. Although our qPCR method for FFPE samples was highly sensitive, with a detection limit of 10 pg µl−1 of F. nucleatum DNA, alternative methods should be considered, such as droplet digital PCR (ddPCR) for identification of low-abundant targets. That method was recently shown to have an even higher sensitivity than to qPCR for the detection of F. nucleatum and it can be applied FFPE tissue [50]. Metagenomic sequencing can also be used for the detection of the bacteria [38] and this has the advantage to determine the complete microbiome of a sample, but the costs, time requirement and downstream bioinformatic analysis still hamper routine application. For the time being, that leaves qPCR as a suitable method of detection of F. nucleatum in FFPE samples.
Conclusions
F. nucleatum, a Gram-negative bacterium that typically resides in the oral cavity, has been implied to contribute to the development of CRC. While the underlying mechanisms have still not been resolved, it is undisputed that F. nucleatum is overabundant in inflammatory intestinal lesions. We investigated the presence of these bacteria by qPCR in inflammatory colon biopsies from UC cases and at different stages of carcinogenesis. Our data confirm an increased prevalence of F. nucleatum in lesions with sCRC, CACs and colitis-associated HGD compared to tissues from a non-inflamed colon and from cases presenting with UC only. From this, we conclude that a higher prevalence of F. nucleatum is associated with advancing cancerous development, but not with UC itself.
Availability of Data and Material
Not applicable.
Code availability
Not applicable.
References
Ordás I, Eckmann L, Talamini M, Baumgart DC, Sandborn WJ (2012) Ulcerative colitis. Lancet 380(9853):1606–1619. https://doi.org/10.1016/S0140-6736(12)60150-0
Alatab S, Sepanlou SG, Ikuta K, Vahedi H, Bisignano C, Safiri S et al (2020) The global, regional, and national burden of inflammatory bowel disease in 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol Hepatol 5(1):17–30. https://doi.org/10.1016/S2468-1253(19)30333-4
Mueller S, Khalid M, Patel H, Wilke T, Dittmar A (2021) P662 A retrospective claims analysis on the prevalence and incidence of ulcerative colitis in Germany and the frequency of advanced therapy use. J Crohns Colitis 15(Supplement_1):S587–S588. https://doi.org/10.1093/ecco-jcc/jjab076.782
Eaden JA, Abrams KR, Mayberry JF (2001) The risk of colorectal cancer in ulcerative colitis: a meta-analysis. Gut 48(4):526–535. https://doi.org/10.1136/gut.48.4.526
Ullman TA, Itzkowitz SH (2011) Intestinal inflammation and cancer. Gastroenterology 140(6):1807–1816. https://doi.org/10.1053/j.gastro.2011.01.057
Rogler G (2014) Chronic ulcerative colitis and colorectal cancer. Cancer Lett 345(2):235–241. https://doi.org/10.1016/j.canlet.2013.07.032
Woolrich AJ, DaSilva MD, Korelitz BI (1992) Surveillance in the routine management of ulcerative colitis: The predictive value of low-grade dysplasia. Gastroenterology 103(2):431–438. https://doi.org/10.1016/0016-5085(92)90831-I
Lang-Schwarz C, Adler W, Geppert M, Seitz G, Sterlacci W, Falkeis-Veits C, Veits L, Drgac J, Melcher B, Lang-Schwarz K, Nikolaev S, Dregelies T, Krugmann J, Vieth M (2020) Sporadic adenoma or ulcerative colitis associated neoplasia? The endoscopist’s information has an impact on diagnosis and patient management. Pathol Res Pract 216(11):153162. https://doi.org/10.1016/j.prp.2020.153162
Kucharzik T, Dignass AU, Atreya R, Bokemeyer B, Esters P, Herrlinger K, Kannengießer K, Kienle P, Langhorst J, Lügering A, Schreiber S, Stallmach A, Stein J, Sturm A, Teich N, Siegmund B (2020) Aktualisierte S3-leitlinie Colitis ulcerosa—living guideline. Z Gastroenterol 58(12):e241–e326. https://doi.org/10.1055/a-1296-3444
Hooper LV, Gordon JI (2001) Commensal host-bacterial relationships in the gut. Science 292(5519):1115–1118. https://doi.org/10.1126/science.1058709
Janney A, Powrie F, Mann EH (2020) Host-microbiota maladaptation in colorectal cancer. Nature 585:509–517. https://doi.org/10.1038/s41586-020-2729-3
Castellarin M, Warren RL, Freeman JD, Dreolini L, Krzywinski M, Strauss J, Barnes R, Watson P, Allen-Vercoe E, Moore RA, Holt RA (2012) Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res 22(2):299–306. https://doi.org/10.1101/gr.126516.111
Kostic AD, Gevers D, Pedamallu CS, Michaud M, Duke F, Earl AM, Ojesina AI, Jung J, Bass AJ, Tabernero J, Baselga J, Liu C, Shivdasani RA, Ogino S, Birren BW, Huttenhower C, Garrett WS, Meyerson M (2012) Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res 22(2):292–298. https://doi.org/10.1101/gr.126573.111
Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW (2013) Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe 14(2):195–206. https://doi.org/10.1016/j.chom.2013.07.012
Rubinstein MR, Baik JE, Lagana SM, Han RP, Raab WJ, Sahoo D, Dalerba P, Wang TC, Han YW (2019) Fusobacterium nucleatum promotes colorectal cancer by inducing Wnt/β-catenin modulator Annexin A1. EMBO Rep. https://doi.org/10.15252/embr.201847638
Kostic AD, Chun E, Robertson L, Glickman JN, Gallini CA, Michaud M, Clancy TE, Chung DC, Lochhead P, Hold GL, El-Omar EM, Brenner D, Fuchs CS, Meyerson M, Garrett WS (2013) Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14(2):207–215. https://doi.org/10.1016/j.chom.2013.07.007
Mima K, Nishihara R, Qian ZR, Cao Y, Sukawa Y, Nowak JA, Yang J, Dou R, Masugi Y, Song M, Kostic AD, Giannakis M, Bullman S, Milner DA, Baba H, Giovannucci EL, Garraway LA, Freeman GJ, Dranoff G, Garrett WS, Huttenhower C, Meyerson M, Meyerhardt JA, Chan AT, Fuchs CS, Ogino S (2016) Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut 65(12):1973–1980. https://doi.org/10.1136/gutjnl-2015-310101
Liu H, Hong XL, Sun TT, Huang XW, Wang JL, Xiong H (2020) Fusobacterium nucleatum exacerbates colitis by damaging epithelial barriers and inducing aberrant inflammation. J Dig Dis 21(7):385–398. https://doi.org/10.1111/1751-2980.12909
Li R, Shen J, Xu Y (2022) Fusobacterium nucleatum and colorectal cancer. Infect Drug Resist 15:1115–1120. https://doi.org/10.2147/IDR.S357922
Ahmad Kendong SM, Raja Ali RA, Nawawi KNM, Ahmad HF, Mokhtar NM (2021) Gut dysbiosis and intestinal barrier dysfunction: potential explanation for early-onset colorectal cancer. Front Cell Infect Microbiol 11:744606. https://doi.org/10.3389/fcimb.2021.744606
Brennan CA, Garrett WS (2019) Fusobacterium nucleatum – symbiont, opportunist and oncobacterium. Nat Rev Microbiol 17(3):156–166. https://doi.org/10.1038/s41579-018-0129-6
Bradshaw DJ, Marsh PD, Watson GK, Allison C (1998) Role of Fusobacterium nucleatum and coaggregation in anaerobe survival in planktonic and biofilm oral microbial communities during aeration. Infect Immunol 66(10):4729–4732. https://doi.org/10.1128/IAI.66.10.4729-4732.1998
Han YW, Fardini Y, Chen C, Iacampo KG, Peraino VA, Shamonki JM, Redline RW (2010) Term stillbirth caused by oral Fusobacterium nucleatum. Obstet Gynecol 115(2 Pt 2):442–445. https://doi.org/10.1097/AOG.0b013e3181cb9955
Kai A, Cooke F, Antoun N, Siddharthan C, Sule O (2008) A rare presentation of ventriculitis and brain abscess caused by Fusobacterium nucleatum. J Med Microbiol 57(Pt 5):668–671. https://doi.org/10.1099/jmm.0.47710-0
Yoneda M, Kato S, Mawatari H, Kirikoshi H, Imajo K, Fujita K, Endo H, Takahashi H, Inamori M, Kobayashi N, Kubota K, Saito S, Tohnai I, Watanuki K, Wada K, Maeda S, Nakajima A (2011) Liver abscess caused by periodontal bacterial infection with Fusobacterium necrophorum. Hepatol Res 41(2):194–196. https://doi.org/10.1111/j.1872-034X.2010.00748.x
Han YW, Redline RW, Li M, Yin L, Hill GB, McCormick TS (2004) Fusobacterium nucleatum induces premature and term stillbirths in pregnant mice: implication of oral bacteria in preterm birth. Infect Immunol 72(4):2272–2279. https://doi.org/10.1128/IAI.72.4.2272-2279.2004
Strauss J, Kaplan GG, Beck PL, Rioux K, Panaccione R, Devinney R, Lynch T, Allen-Vercoe E (2011) Invasive potential of gut mucosa-derived Fusobacterium nucleatum positively correlates with IBD status of the host. Inflamm Bowel Dis 17(9):1971–1978. https://doi.org/10.1002/ibd.21606
Li D-H, Li Z-P, Yan Z, Zhou G-Z, Ren R-R, Zhao H-J, Zhang N-N, Li J-F, Peng L-H, Yang Y-S (2021) Fecal Fusobacterium nucleatum harbored virulence gene fadA are associated with ulcerative colitis and clinical outcomes. Microb Pathog 157:104964. https://doi.org/10.1016/j.micpath.2021.104964
Xu M, Yamada M, Li M, Liu H, Chen SG, Han YW (2007) FadA from Fusobacterium nucleatum utilizes both secreted and nonsecreted forms for functional oligomerization for attachment and invasion of host cells. J Biol Chem 282(34):25000–25009. https://doi.org/10.1074/jbc.M611567200
Fardini Y, Wang X, Témoin S, Nithianantham S, Lee D, Shoham M, Han YW (2011) Fusobacterium nucleatum adhesin FadA binds vascular endothelial cadherin and alters endothelial integrity. Mol Microbiol 82(6):1468–1480. https://doi.org/10.1111/j.1365-2958.2011.07905.x
Ma C-T, Luo H-S, Gao F, Tang Q-C, Chen W (2018) Fusobacterium nucleatum promotes the progression of colorectal cancer by interacting with E-cadherin. Oncol Lett 16(2):2606–2612. https://doi.org/10.3892/ol.2018.8947
Liu Le, Liang L, Liang H, Wang M, Lu B, Xue M, Deng J, Chen Y (2019) Fusobacterium nucleatum aggravates the progression of colitis by regulating M1 macrophage polarization via AKT2 pathway. Front Immunol 10:1324. https://doi.org/10.3389/fimmu.2019.01324
Flanagan L, Schmid J, Ebert M, Soucek P, Kunicka T, Liska V, Bruha J, Neary P, Dezeeuw N, Tommasino M, Jenab M, Prehn JHM, Hughes DJ (2014) Fusobacterium nucleatum associates with stages of colorectal neoplasia development, colorectal cancer and disease outcome. Eur J Clin Microbiol Infect Dis 33(8):1381–1390. https://doi.org/10.1007/s10096-014-2081-3
Mima K, Sukawa Y, Nishihara R, Qian ZR, Yamauchi M, Inamura K, Kim SA, Masuda A, Nowak JA, Nosho K, Kostic AD, Giannakis M, Watanabe H, Bullman S, Milner DA, Harris CC, Giovannucci E, Garraway LA, Freeman GJ, Dranoff G, Chan AT, Garrett WS, Huttenhower C, Fuchs CS, Ogino S (2015) Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol 1(5):653–661. https://doi.org/10.1001/jamaoncol.2015.1377
Yan X, Liu L, Li H, Qin H, Sun Z (2017) Clinical significance of Fusobacterium nucleatum, epithelial-mesenchymal transition, and cancer stem cell markers in stage III/IV colorectal cancer patients. Onco Targets Ther 10:5031–5046. https://doi.org/10.2147/OTT.S145949
Li Y-Y, Ge Q-X, Cao J, Zhou Y-J, Du Y-L, Shen B, Wan Y-JY, Nie Y-Q (2016) Association of Fusobacterium nucleatum infection with colorectal cancer in Chinese patients. World J Gastroenterol 22(11):3227–3233. https://doi.org/10.3748/wjg.v22.i11.3227
Tahara T, Shibata T, Kawamura T, Okubo M, Ichikawa Y, Sumi K, Miyata M, Ishizuka T, Nakamura M, Nagasaka M, Nakagawa Y, Ohmiya N, Arisawa T, Hirata I (2015) Fusobacterium detected in colonic biopsy and clinicopathological features of ulcerative colitis in Japan. Dig Dis Sci 60(1):205–210. https://doi.org/10.1007/s10620-014-3316-y
Yu T, Guo F, Yu Y, Sun T, Ma D, Han J, Qian Y, Kryczek I, Sun D, Nagarsheth N, Chen Y, Chen H, Hong J, Zou W, Fang J-Y (2017) Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 170(3):548-563.e16. https://doi.org/10.1016/j.cell.2017.07.008
Zhang S, Yang Y, Weng W, Guo B, Cai G, Ma Y, Cai S (2019) Fusobacterium nucleatum promotes chemoresistance to 5-fluorouracil by upregulation of BIRC3 expression in colorectal cancer. J Exp Clin Cancer Res 38(1):14. https://doi.org/10.1186/s13046-018-0985-y
(2019) Digestive system tumours, 5th edition. World Health Organization classification of tumours series, 5th ed, Vol. 1. International Agency for Research on Cancer, Lyon
Gajendran M, Loganathan P, Jimenez G, Catinella AP, Ng N, Umapathy C, Ziade N, Hashash JG (2019) A comprehensive review and update on ulcerative colitis. Dis Mon 65(12):100851. https://doi.org/10.1016/j.disamonth.2019.02.004
Su W, Chen Y, Cao P, Chen Y, Guo Y, Wang S, Dong W (2020) Fusobacterium nucleatum promotes the development of ulcerative colitis by inducing the autophagic cell death of intestinal epithelial. Front Cell Infect Microbiol 10:594806. https://doi.org/10.3389/fcimb.2020.594806
Huh J-W, Roh T-Y (2020) Opportunistic detection of Fusobacterium nucleatum as a marker for the early gut microbial dysbiosis. BMC Microbiol 20(1):208. https://doi.org/10.1186/s12866-020-01887-4
Proença MA, Biselli JM, Succi M, Severino FE, Berardinelli GN, Caetano A, Reis RM, Hughes DJ, Silva AE (2018) Relationship between Fusobacterium nucleatum, inflammatory mediators and microRNAs in colorectal carcinogenesis. World J Gastroenterol 24:5351–5365. https://doi.org/10.3748/wjg.v24.i47.5351
Suehiro Y, Sakai K, Nishioka M, Hashimoto S, Takami T, Higaki S, Shindo Y, Hazama S, Oka M, Nagano H, Sakaida I, Yamasaki T (2017) Highly sensitive stool DNA testing of Fusobacterium nucleatum as a marker for detection of colorectal tumours in a Japanese population. Ann Clin Biochem 54:86–91. https://doi.org/10.1177/0004563216643970
Eklöf V, Löfgren-Burström A, Zingmark C, Edin S, Larsson P, Karling P, Alexeyev O, Rutegård J, Wikberg ML, Palmqvist R (2017) Cancer-associated fecal microbial markers in colorectal cancer detection. Int J Cancer 141:2528–2536. https://doi.org/10.1002/ijc.31011
Nosho K, Sukawa Y, Adachi Y, Ito M, Mitsuhashi K, Kurihara H, Kanno S, Ymamamoto I, Ishigami K, Igarashi H, Maruyama R, Imai K, Yamamoto H, Shinomura Y (2016) Association of Fusobacterium nucleatum with immunity and molecular alterations in colorectal cancer. World J Gastroenterol 22:557–566. https://doi.org/10.3748/wjg.v22.i2.557
de Carvalho AC, de Mattos Pereira L, Datorre JG, Dos Santos W, Berardinelli GN, Matsushita MdM, Oliveira MA, Durães RO, Guimarães DP, Reis RM (2019) Microbiota profile and impact of Fusobacterium nucleatum in colorectal cancer patients of barretos cancer hospital. Front Oncol 9:813. https://doi.org/10.3389/fonc.2019.00813
McCoy AN, Araújo-Pérez F, Azcárate-Peril A, Yeh JJ, Sandler RS, Keku TO (2013) Fusobacterium is associated with colorectal adenomas. PLoS One 8(1):e53653. doi:https://doi.org/10.1371/journal.pone.0053653
Datorre JG, Carvalho AC de, Dos Reis MB, Dos Reis M, Matsushita M, Santos F, Guimarães DP, Reis RM (2022) Accuracy and Clinical Relevance of Intra-Tumoral Fusobacterium nucleatum Detection in Formalin-Fixed Paraffin-Embedded (FFPE) Tissue by Droplet Digital PCR (ddPCR) in Colorectal Cancer. Diagnostics (Basel) 12. https://doi.org/10.3390/diagnostics12010114
Acknowledgements
We thank the Forschungskommission of the Klinikum Bayreuth for the financial support to F. Haumaier and T. Dregelies.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was financially supported by the Forschungskommission of the Klinikum Bayreuth/Germany to F. Haumaier and T. Dregelies.
Author information
Authors and Affiliations
Contributions
FH designed the study. TD performed the analysis, evaluated and compiled the data and wrote the manuscript. MV, SB and SW supervised the studies, reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics Approval
The study was approved by the ethics committee of the University Bayreuth (#O 1305/1-GB).
Consent to Participate
Informed consent forms were signed by all patients of the Klinikum Bayreuth at the timepoint of hospitalization.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dregelies, T., Haumaier, F., Sterlacci, W. et al. Detection of Fusobacterium nucleatum in Patients with Colitis-Associated Colorectal Cancer. Curr Microbiol 80, 293 (2023). https://doi.org/10.1007/s00284-023-03398-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-023-03398-7