Abstract
The COVID-19 pandemic has followed a wave pattern, with an increase in new cases followed by a drop. Several factors influence this pattern, including vaccination efficacy over time, human behavior, infection management measures used, emergence of novel variants of SARS-CoV-2, and the size of the vulnerable population, among others. In this study, we used three statistical approaches to analyze COVID-19 dissemination data collected from 15 November 2021 to 09 January 2022 for the prediction of further spread and to determine the behavior of the pandemic in the top 12 countries by infection incidence at that time, namely Distribution Fitting, Time Series Modeling, and Epidemiological Modeling. We fitted various theoretical distributions to data sets from different countries, yielding the best-fit distribution for the most accurate interpretation and prediction of the disease spread. Several time series models were fitted to the data of the studied countries using the expert modeler to obtain the best fitting models. Finally, we estimated the infection rates (β), recovery rates (γ), and Basic Reproduction Numbers (\(R_{0}\)) for the countries using the compartmental model SIR (Susceptible-Infectious-Recovered). Following more research on this, our findings may be validated and interpreted. Therefore, the most refined information may be used to develop the best policies for breaking the disease's chain of transmission by implementing suppressive measures such as vaccination, which will also aid in the prevention of future waves of infection.
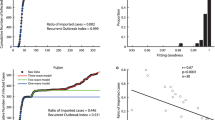
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
So far, the COVID-19 pandemic has followed a wave pattern, with an increase in numbers of new cases followed by a decrease. During the pandemic, several factors have influenced whether new COVID-19 cases are increasing or declining across the countries. The effectiveness of vaccines over time, human behavior, infection control measures, the emergence of new variants, and the size of the population vulnerable due to a lack of immunity, and the size of immunized population, whether through natural infection or by active vaccination programs. For instance, during the winter months of 2020–21 in many countries, a large surge of COVID-19 cases occurred as people traveled and gathered for the holidays [1].Vaccines began arriving in various countries in December 2020, beginning with the USA, and helped to lower the number of new infections in various countries until the spring of 2021. The fast contagious Delta variant emerged and eventually became widespread in July 2021, resulting in a new wave of infections. It was caused due to decline in immunity, as well as weakening of government imposed restrictions and infection-prevention efforts. The first vaccinations in the USA occurred in January 2021. By March, the daily number of new infections had reduced considerably, and the numbers fell even further from April to June. The introduction and spread of the delta variant, however, had generated a new wave of COVID-19 cases by July. As of October 2021, infection rates appear to be decreasing, owing in part to increased vaccination rates. In late 2021, a sudden increase in COVID-19 cases was observed, following the emergence of the Omicron variation, which was first spotted in South Africa and went on to become the most common form worldwide, spreading at least three times faster than the prior variant [2]. The SARS-CoV-2, like all viruses, evolves over time. The majority of changes have little to no effect on its physiological properties. Some changes, however, may affect them and affecting on how easily it spreads the severity of the associated disease, or the performance of vaccines, therapeutic agents, diagnostic tests, or other public health and social measures to be implemented. In collaboration with partners, expert networks, national authorities, institutions, and researchers, the World Health Organization (WHO) has been monitoring and evaluating the evolution of SARS-CoV-2 since January 2020. The emergence of variants posing an increased risk to global public health in late 2020 prompted the identification of specific Variants of Interest (VOIs) and Variants of Concern (VOCs) in order to priorities global monitoring and research, ultimately, informing the ongoing response to the COVID-19 pandemic. The WHO brought together global experts in virology, microbial nomenclature, and communication from various countries and agencies to consider simple, non-stigmatizing nomenclature for VOIs and VOCs. At the moment, this expert group convened by WHO has recommended using Greek Alphabet letters, such as Alpha, Beta, Gamma, Delta, and Omicron, which will be easier and more practical to discuss with non-scientific audiences. The WHO publishes regular updates on SARS-CoV-2 classifications, the geographic distribution of VOCs, and summaries of their phenotypic characteristics (transmissibility, disease severity, risk of re-infection, and impacts on diagnostics and vaccine performance) based on published studies. The Omicron variant of COVID-19 has been called a variant of concern by WHO [3]. Omicron is spreading faster than any previous variant, with a doubling time of 2–3 days. The overall risk related to this variant remains very high. There is no information to suggest that Omicron causes different symptoms from other COVID-19 variants. Omicron does however typically cause less severe disease than previous variants like Delta. The Omicron variant is genetically related to the Gama variant. When compared to the previously emerged variants, the Omicron has more mutations than any other variant ever discovered, with over fifty mutations. This includes 32 different spike protein mutations. Many of these mutations have previously been discovered in other variants, but they have never been discovered together in the same variant. Many of these mutations are found in the spike protein's receptor-binding domain, which is a critical component of the protein that binds to human receptor proteins to allow the coronavirus into the cells, and may thus play a role in antibody recognition by previous infection or vaccinations [4]. Many spike protein mutations have previously been linked to antibody resistance as well as increased transmission. As a result, patients who have developed immunity to previous viral strains may be more susceptible to infection by this variant. While the behavior of the coronavirus cannot be predicted precisely based on evidence generated by individual mutations, such analysis may provide useful insights and pointers for future research because the effect of a combination of mutations is not equal to the sum of individual mutations. More research into the effects of these mutations on transmissibility and vaccine evasion is needed. Some of the spike protein mutations fail primers used in commercially available RT-PCR kits, rendering them undetectable and producing a false-negative result [5, 6].
In the current study, we have analyzed the data collected in the fresh wave of infections from top 12 countries by incidence of daily new infection incidence. We have used three statistical methods, Distribution Fitting, Time Series Modeling, and the Epidemiological Modeling to analyze the data for obtaining insights on behavior of the pandemic and further forecasting. In epidemiology, statistical modeling and prediction provided a tool for determining the causes of infection transmission as well as treatments for prevention or limitation. We gather, assess, and interpret data when a novel infectious disease emerges or an outbreak of an established infectious disease occurs in order to identify effective strategies to prevent future transmission. Many infectious diseases are unconcerned about country borders, race, creed, caste, or community, and typically affect only one region of the world at a time, before rapidly spreading to other communities and eventually becoming a pandemic [7]. Infectious diseases spread over time, and many factors determine the behavior and kinetics of infection propagation, including the length of the epidemic, the average number of infections per infected person, and the time when patients begin to display symptoms, among others. The infection growth rate, which varies by country due to factors such as health infrastructure and testing processes, determines the total number of illnesses [8]. Some researchers have attempted to fit different theoretical distributions such as the Normal, Negative Binomial, Poisson, Beta, Gamma, Exponential, Lognormal, and many other discrete and continuous parametric and non-parametric distributions to understand the behavior and nature of infectious disease spread. Mayer and Held (2014) analyzed the fitting of power-law distributions of infectious disease spread [9]. Under this category, they fitted power law, student, and Gaussian distributions and tested their sensitivity with the fitting metric AIC (Akaike Information Criteria). They demonstrated that, in contrast to others, the Power law distribution fit well in explaining the spread of infectious diseases. A type of nonparametric distribution called Turnbull and other parametric distributions were fitted by Virlogeux et al. (2015)to estimate the human avian influenza A(H7N9) virus infections incubation durations through the best fitted distribution, similarly, in their another study, Virlogeux et al. (2016) worked on the comparing the incubation period distributions of human infections with MERS-CoV for the countries South Korea along with Saudi Arabia by fitting different parametric and non-parametric theoretical distributions [10, 11].
Based on noisy case reporting data, Li et al. (2018) fitted different discrete and continuous theoretical distributions for forecasting and parameter estimation of infectious diseases. They basically fitted the binomial, negative binomial, beta, gamma, and poisson distributions and forecasted disease infection based on the simulated data based on their best fittings [12]. de Souza et al. (2019) investigated the statistical behavior of admissions in hospitals for respiratory diseases by fitting different distributions [13]. They have fitted the distributions Burr, Inverse Gaussian, Lognormal, Pert, Rayleigh, and Weibull and tested their goodness of fit using the Kolmogorov–Smirnov Test, Anderson–Darling Test, and Chi-square Test to determine the best fitted distribution for the same and demonstrated that different distributions are best fitted for different seasons. Valvo et al., (2020) demonstrated that the Bimodal Lognormal Distribution, as a phenomenological epidemiological model, better predicts COVID-19 deaths [14]. Unglaub and Spendier (2021) fitted different sigmoidal distributions, primarily Gompertz, Richards, Logistic, Stannard, and Schnute, to analyze the infectious diseases dissemination with application to COVID-19 data, and found that the Richards model was fitted most suitable for the COVID-19 data because it had the lowest residual sum of squares[15]. Vazquez et al., (2021) worked on finding the best explaining distribution for the spread of the infectious diseases and for the study the author fitted the exponential, gamma and power-law distributions and shown their short-term and long-term behaviors for the infectious dissemination[16]. Mubarak and Almetwally et al., (2021) suggested a three-parameter exponential distribution for the study of the COVID-19 spread and shown that their distribution is best fitted over the distributions given by other researchers for COVID-19 dissemination on the basis of various fitting measures[17].
Furthermore, time series regression modeling and prediction of infectious diseases are critical for understanding disease behavior and developing better policies to combat the problem. Because the epidemic evolves with the time, it is critical to study its trending behavior to learn many things, such as when it will end, when it will peak, and how many people will be affected by it. Because forecasting time series is critical in disease modeling, it is important to note that proper attention should be given to fitting the suitable model for the given time series. The best time series prediction is, of course, based on the best fitted model. AR (Auto Regressive), MA (Moving Average), ARMA (Auto Regressive Moving Average), ARIMA (Auto Regressive Integrated Moving Average), and SARIMA (Seasonal Auto Regressive Integrated Moving Average) are the most commonly used time series models for predicting infectious diseases. These time series models can help predict the occurrence, dangers, and distribution or dilatation trend of diseases like Dengue, Ebola, Influenza, Malaria, and other infectious diseases [18]. Imai et al. (2015) used time series models, mathematical modeling, wavelet analysis, and ARIMA to assess the short-term relation of air pollution and weather with infectious disease mortality or morbidity and shown that time series regression is superior to other methods [19]. Chaurasia and Pal (2020) used a new method, simple average, moving average, single exponential smoothing, Holt linear trend method, Holt-Winters method, and ARIMA to analyze COVID-19 cases, deaths, and recoveries worldwide [20]. Chyon et al. (2022) used machine learning to perform time series analysis and prediction of COVID-19 affected patients using the ARIMA model [21]. They used the ARIMA model to examine the temporal dynamics of COVID-19's global spread from January 22, 2020 to April 7, 2020. Kibria et al. (2022) fitted five different time series models; AR, MA, ARMA, Rolling Forest Origin, and ARIMA for predicting the spread of the third wave of the COVID-19 pandemic and demonstrated that ARIMA is the best model among them for the purpose under consideration [22].
On the other hand, epidemiological modeling is used to determine the competing risks of infectious disease deaths. Cassels et al. (2008) investigated HIV transmission dynamics mathematical models [23]. Andraud et al. (2012) conducted a systematic review of various structural methods to different epidemiological models for the transmission of dengue [24]. Huppert and Katriel (2013) studied mathematical modeling and forecasting in the field of infectious disease epidemiology [25].The modeling of Ebola virus infection transmission patterns in Liberia was explored and addressed by Xia et al. (2015) [26]. Some more authors have used SIR and modified SIR models for studying the Ebola virus infection dynamics [27,28,29]. Driessche et al., (2017) used the next generation matrix approach to calculate the basic reproduction number,\(R_{0}\), for infectious diseases, as well as other reproduction numbers linked to \(R_{0}\) that are valuable in guiding management measures[30]. Li et al., (2020) developed an infectious disease dynamics model and a time series model to anticipate the trend and short-term prediction of COVID-19 transmission in mainland China [31]. Popov and Nakov (2021) focused on epidemiological models for COVID-19 transmission [32]. For the description and prediction of COVID-19 outbreaks, Moein et al. (2021) used a variety of mathematical tools, including the Susceptible-Infected-Recovered (SIR) model [33]. They simulated a model of the epidemic in Iran's Isfahan area, coupled with three suppressive methods based on the severity of social distancing. Yang et al., (2020) worked on the short-term forecasts and long-term mitigation evaluation on the COVID-19 epidemic in Hubei Province of China [34]. Alvarez et al., (2021) proposed a simple epidemiological model that could be applied in Excel spreadsheets and could accurately simulate the data from COVID-19 pandemic [35]. Wang et al., (2022) worked on the Modified SIR Model to analyze the time dependence of active and hospitalized COVID-19 cases in China [36].
In this study, we have considered top 12 countries in terms of the number of daily infections for the analysis and prediction of COVID-19 disease infections outbreak having latest variants. For the modeling and prediction of the outbreak of COVID-19, we have considered three methods, distribution fitting, time series modeling and epidemiological modeling as discussed above. We have studied different properties of different methods and on the basis of the analysis, the prediction of infectious disease spread is discussed which may be utilized in making different effective policies for further dissemination of the COVID-19 infectious disease. Ultimately, we have considered three methodologies to model and predict the COVID-19 dissemination and in time series, we used exponential smoothing method along with the ARIMA method. The SIR model has been used as per the availability of the global data after the emergence of the Omicron variant of SARS-CoV-2 and the basic reproduction number has been calculated for the considered countries so that on the basis of the results through the three methods, the most effective and robust policies may be made to prevent the further spread of pandemic in future.
Methodology
Data Description
We obtained the country wise data of infected, recovered, and confirmed cases along with the population data for the all 12 considered countries; Afghanistan, Austria, Belgium, Czech Republic, India, Japan, Mexico, Pakistan, Philippines, South Africa, South Korea, and Thailand using CovsirPhy package (https://github.com/lisphilar/covid19-sir) deployed in Python. The CovsirPhy package retrieves data from ‘COVID-19 Data Hub’ (https://covid19datahub.io/) and ‘COVID-19 dataset in Japan’ (https://github.com/lisphilar/covid19-sir/data/japan) for Japan specific data. For distribution fitting we used the Stats module from Python’s SciPy library (https://scipy.org/) and Python’s Matplotlib (https://matplotlib.org/) for creating visualizations. The time series data analysis using exponential and ARIMA models through expert modeler has been done using SPSS software. For compartmental modeling we deployed deSolve package in R software (https://www.r-project.org/).
Distribution Fitting
The phenomenon which is developing over time may be explained through the best fitted theoretical distribution. In this section for the analysis and forecasting of the COVID-19 infectious disease spread, we have considered 123 theoretical parametric and non-parametric; discrete and continuous distributions. Out of 123 total fitted distributions, the names of 19 discrete distributions are, Bernoulli, Betabinom, Binomial, Boltzmann, Dlaplace, Geometric, Hypergeometric, Logser, Nbinom, Nchypergeom-Fisher, Nchypergeom-Wallenius, Nhypergeom, Planck, Poisson, Randint, Skellam, Yulesimon, Zipf, Zipfian while the 104 continuous distributions are, Alpha, Anglit, Arcsine, Argus, Beta, Betaprime, Bradford, Burr, Burr12, Cauchy, Chi, Chi-Square, Cosine, Crystalball, Dgamma, Dweibull, Erlang, Expon, Exponnorm, Exponpow, Exponweib, F, Fatiguelife, Fisk, Foldcauchy, Foldnorm, Gamma, Gausshyper, Genexpon, Genextreme, Gengamma, Genhalflogistic, Genhyperbolic, Geninvgauss, Genlogistic, Gennorm, Genpareto, Gilbrat, Gompertz, Gumbel_L, Gumbel_R, Halfcauchy, Halfgennorm, Halflogistic, Halfnorm, Hypsecant, Invgamma, Invgauss, Invweibull, Johnsonsb, Johnsonsu, Kappa-3, Kappa-4, Ksone, Kstwo, Kstwobign, Laplace, Laplace_Asymmetric, Levy, Levy_L, Levy_Stable, Loggamma, Logistic, Loglaplace, Lognorm, Loguniform, Lomax, Maxwell, Mielke, Moyal, Nakagami, Ncf, Nct, Ncx-2, Norm, Norminvgauss, Pareto, Pearson-3, Powerlaw, Powerlognorm, Powernorm, Rayleigh, Rdist, Recipinvgauss, Reciprocal, Rice, Semicircular, Skewcauchy, Skewnorm, Studentized-Range, t, Trapezoid, Trapz, Triang, Truncexpon, Truncnorm, Tukeylambda, Uniform, Vonmises, Vonmises-Line, Wald, Weibull-Max, Weibull-Min, Wrapcauchy. The probability mass functions of the discrete distributions and the probability density functions of the continuous distributions may be obtained using the Stats package in Python’s SciPy library.
The all above distributions are fitted to the data set of confirmed COVID-19 cases of each of the considered countries and the best fitted distributions on the basis of the goodness of fit are obtained for each of the considered countries. The graphs of the best fitted distributions along with the values of the location and scale parameters for all the twelve considered distributions are also presented in pictorial form.
Time Series Modeling
Since the COVID-19 infectious disease is developing over time that is why it must be analyzed and predicted through the best fitted time series model. To find the best fitted time series models for the considered all twelve countries, we have used two methods. The first method is the well-known Exponential Smoothing Method (ESM), for which different models namely; Simple Linear Trend, Holt’s Linear Trend, Brown’s Linear Trend, and Damped Linear Trend have been used. The second well established method for time series modeling is the ARIMA method which is a composition of AR Model, Integration (I), and MA Models. We discuss below the ARIMA Model of time series.
ARMA (Autoregressive Moving Average) and ARIMA (Autoregressive Integrated Moving Average) Models
The autoregressive integrated moving average (ARIMA) model, introduced by Box et al. (1994) is an improvement over ARMA model dealing with the non-stationary time series [37]. Non-stationary means that the mean and variance of a time series are not consistent throughout time. The integrated part refers to a differencing starting step that can be used to remove the time series' non-stationarity. Promprou et al. (2006), Liu et al. (2011), and Coutin et al. (2007), all used this method to analyze epidemiological time series [38,39,40]. As a result, we start with the ARMA model and then move on to ARIMA.
ARMA (Autoregressive Moving Average) Model
The autoregressive moving average ARMA of order (\(p,\,q\)) is the combination of the AR(\(p\)) model with order \(\,p\) and the MA(\(\,q\)) model with order \(\,q\). ARMA model, with a few of polynomials, exhibits a weak stationary random process, one for AR and another for MA. The ARMA model forecasts the series' future values for a time series \(Y_{t}\).The AR half of the model regresses the variable for its own lagged values, while the MA part accounts for error terms that occur synchronously at various times in the past. Because of the MA and its behavior, ARMA is suited for unobserved shocks, such as a pandemic. The ARMA (\(p,\,q\)) model suggested by Wold (1938) [41] is as follows,
\(Y_{t} = (\theta_{1} Y_{t - 1} + \theta_{2} Y_{t - 2} + ... + \theta_{p} Y_{t - p} ) + (\phi_{0} \varepsilon_{t} + \phi_{1} \varepsilon_{t - 1} + ... + \phi_{q} \varepsilon_{t - q} ),\quad {\text{or}}\),
where, \(\theta_{1} ,\theta_{2} ,\,...,\,\theta_{p}\) and \(\phi_{1} ,\,\phi_{2} ,\,...,\,\phi_{q}\) along with \(\phi_{0} = 1\) and the error terms \(\varepsilon_{t}\) is white noise. Now, the well-known Box-Jenkins (1976) approach [42], which employs an estimation of the model's parameters or constants and delivers the best diagnostics of these parameters, can be used for better forecasting. The ordinary least square approach is used to estimate the parameters of the ARMA model.
Consider the following scenario for a better understanding Let \(\{ X_{t} \}\) be the sequence of observations that hasn't been observed and \(\{ Y_{t} \}\) be the sequence that was witnessed and we have,
where, \(\varepsilon_{t}\) and \(\eta_{t}\) are independent along with the white noise and further \(\{ X_{t} \}\) is AR (1). Thus we have,
Now it is evident that \(\xi_{t}\) is stationary and \(Cov\,(\xi_{t} ,\,\xi_{t + h} ) = 0\), \(h \ge 2\). Thus \(\xi_{t}\) is possible to model MA (1) model and \(\{ Y_{t} \}\) as ARMA (1, 1) model. The ACF and PACF (Partial ACF) plots are used to determine how much order of \(p\) and \(q\) should be taken for better forecasting of the time series.
ARIMA (Autoregressive Integrated Moving Average) Model
The three components of an ARIMA model are:
-
Auto Regression (AR) model, a model that describes a variable that regresses on its lagged or prior values.
-
Integrated (I), the process of differencing essential observations in order for a time series to become stationary.
-
Moving average (MA) for lag observations, the docility of an observation and a residual from the MA model.
Let \(\{ Y_{t} \}\) be a non-stationary time series. Now we use the differencing approach to make it stationary, and we look at the order of differencing that makes the series stationary. Let the series' first-order difference be,
The differences in the second order are as follows:
\(X_{t} = \nabla^{2} Y_{t} = \nabla (\nabla Y_{t} ) = Y_{t} - 2Y_{t - 1} + Y_{t - 2}\) and so on. We claim the time series is stationary when the differencing process is stationary, and we may use the ARMA model to model this integrated series. Thus \(\{ Y_{t} \}\) is ARIMA model, denoted by \({\text{ARIMA(}}p,\,d,\,q{)}\) if \(X_{t} = \nabla^{d} Y_{t}\) is a \({\text{ARMA(}}p,\,q{)}\) model. Where p is order of AR, d is order of Integration and q is order of MA.
Epidemiological Model SIR
In the current scenario of COVID-19 epidemic, we are having mainly data on Susceptible, Infected, and Recovered individuals for different counties. In this situation SIR model is the most suitable Epidemiological Model for the analysis and prediction of the disease spread. Kermack and McKendrick created the SIR model in 1927 [43], and it has since been applied to a variety of infectious diseases, including airborne pediatric diseases with lifetime immunity following recovery, such as measles, mumps, rubella, and pertussis. S, I, and R in the SIR model show the number of susceptible, infected, and recovered cases, and \(N = S + I + R\) is the total population or \(N(t) = S(t) + I(t) + R(t)\) represents the aggregate population at time \(t\).
SIR Model Dynamics
Vital dynamics may not be evaluated during the infection period with emerging eruption compared to a person's lifetime if the sickness is non-fatal. The deterministic part of the SIR model in the form of ordinary differential equations used in this case is,
where, \(N = S + I + R\) is the whole population or \(N(t) = S(t) + I(t) + R(t)\) is the whole population at the time \(t\), \(\beta\) is the rate of infection and \(\gamma\) is the recovery rate. In a population with births and deaths, an epidemic will eventually fade away since the disease will be carried by the least vulnerable individuals. The ratio \(\beta /\gamma\) is denoted by \(R_{0}\), the Basic Reproduction Number is a potential indicator of infectious disease transmission that measures the average number of secondary infections caused by a typical infected individual in a community comprising all susceptible individuals.
Results and Analyses
Distribution Fitting
From the Fig. 1, it may be observed that different countries are experiencing different best fitted distributions. The best fitted distributions for different countries along with their location parameters (Mean) and the scale parameters (Variance) are presented in Table 1 below.
It may be observed from Table 1 that the confirmed COVID-19 cases follow different distributions for different countries with the respective means and variances. As the best fitted distributions have been identified for each of the considered country, therefore, rest of the properties of the distributions may be obtained and this information may be utilized in making effective policies to check the further spread of the COVID-19 infectious disease.
Time Series
Both the methods exponential smoothing and the ARIMA are fitted to the data sets of confirmed COVID-19 cases for all the considered countries and the best fitted time series models for different countries are obtained on the basis of different fitting measures; Coefficient of determination (\(R^{2}\)), Residual Mean Square Error (RMSE), Mean Squared Percentage Error (MAPE), Mean Absolute Error (MAE) Normalized Bayesian Information Criteria (NBIC) through the expert modeler in SPSS Software. Table 2 is describing the best fitted model for the time series data of confirmed COVID-19 cases for each country along with the fitting measures.
From Table 2 above, it may be observed that there are different best fitted models with their fitting measures for different countries under consideration. The same results are also presented in the graphs of different countries in Fig. 2.
On the basis of the best fitted time series models for different countries, we have forecasted next five days confirmed COVID-19 cases which are presented in Table 3.
The graphs of the best fitted models along with their predicted parts for different countries under consideration are presented in the pictorial form in Fig. 2.
Epidemiological Model
There are various methods available in the literature for estimating \(\beta\) and \(\gamma\) for SIR model. We have considered one of the best methods given in the website statsandr.com, where the codes in R are available for estimating these parameters, which has used ‘ode’ function of ‘deSolve’ R package to solve the differential equations of SIR model. Further after giving the initial values of each of the \(\beta\) and \(\gamma\) as 0.05 and then the ‘optim’ function has been used to get the optimum values of \(\beta\) and \(\gamma\) based on the data sets. For the optimum values of \(\beta\) and \(\gamma\) obtained from the data, we have calculated the values of \(R_{0}\) for all the considered countries. The estimates of \(\beta\), \(\gamma\) and \(R_{0}\) are presented in Table 4.
From Table 4, it is evident that most of the considered countries are having the values of their \(R_{0}\) in the interval {1.012, 1.818} that is on average one infectious person is infecting the respective number of persons in the interval {1.012, 1.818} while India is having its \(R_{0}\) as 2.333 it means that in India an infectious person is infecting on average 2.333 persons in the whole susceptible population. The same results are also presented in the graphs of different countries in Fig. 3.
Graphs of SIR models of fitted and observed cumulative incidence of infected individuals for the 12 countries of infection studied in this work. The color coding of the plots of SIR model is as following: Susceptible = “Black”; Infected = “Red”; Recovered = “Green”; and Cumulative infected cases = “Blue” (Color figure online)
Discussion
In the presented work, we have discussed the problem of modeling for the analysis and forecasting of COVID-19 disease dissemination in the top twelve considered countries in terms of the COVID-19 confirmed cases. We concentrated on the period November 2021 to January 2022, which corresponds to the third wave of infections due to the omicron variant outbreak; a summary of further refinements is as follows. Working on prior and available knowledge and with data sources of varying quality, we have performed our analysis. This article relates the model formulations that were used to understand the dynamics predict cases and advise policies during the first and second waves of COVID-19 infections. As the outbreak progressed, a comparison of model predictions and data revealed an evident decrease in errors, implying that our models and methods used have explained and predicted the phenomenon very well. We have addressed in some depth the choice of data sets used to infer model parameters and the impact of this decision on the major emergent features of the latent parameters and the derived parameters from the considered models. Many of the decisions concerning which data sets to use are value judgments based on an epidemiological understanding of the links between disease dynamics and observed outcomes, as we point out. None of the data sets accessible to epidemiological modelers are flawless; they all contain biases and delays; we believe that by applying various best-fit models to the data set under consideration, we can find models that achieve a natural compromise.
We tried to model the problem in three different methods namely, Distribution Fitting, Time Series Regression modeling, and the Epidemiological Modeling. We fitted 123 different discrete and continuous theoretical distributions to the data sets of all considered countries and we find the best fitted distributions of COVID-19 confirmed cases for all the countries on the basis of goodness of fit. We also fitted different time series regression models for the COVID-19 confirmed cases of all the countries and through the expert modeler, we have found the best fitted time series models for different countries under consideration. Through these best fitted time series models, we have also predicted COVID-19 confirmed cases for next five days for each of the countries. Since COVID-19 is an infectious disease, therefore, we have also fitted the well-established SIR model for all the considered countries. We estimated the transmission rates and the recovery rates for all the countries and calculated the Basic Reproduction numbers for all the countries.
From the Table 1, we can observe the best fitted theoretical distributions for all the countries under consideration. The Location and the Scale parameters for the best distributions are also presented in Table 1. The graphs of the best fitted distributions for all the countries are also presented in Fig. 1. Out of 123 distributions for the twelve considered countries, Gausshyper, Halfcauchy, Laplace-Asymmetric, Genhalflogistic, Moyal, Genextreme, Nakagami, DGamma, Beta, Triangular are the ten best fitted distribution for the considered countries. Many researchers have fitted different theoretical discrete and continuous distributions and find the best fitted distributions to predict the phenomenon [12,13,14,15,16,17]. Similarly, we have also fitted various distributions and found the best fitted distributions for different countries to study the behavior of the confirmed cases of COVID-19 infectious disease. From the graphs in Fig. 1, we observe that different country is having different best fitted theoretical distributions for the COVID-19 dissemination and on the basis of the fitted distributions; we infer the average number of infections during the considered period along with the variations in the cases. On the basis of these inferences the better polices for the further dissemination of the infections and the work on the requirement of the further health infrastructure may be carried out.
From Table 2, it is observed that Holt is the best fitted time series model for Afghanistan only and Brown is the best fitted model for Austria, India, Pakistan, Philippines, and South Africa while ARIMA is best fitted model for the rest of the countries Belgium, Czech Republic, Japan, Mexico, South Korea, and Thailand. The best fitted times series models along with their predicted graphs are also presented in Fig. 2. The time series models are capable for further predictions of the disease spread. As far as the time series model fitting is concerned for the COVID-19 cases, Imai et al. (2015), Chaurasia and Pal (2020), Chyon et al. (2022), and Kibria et al. (2022) have fitted the ARIMA model for the better explanation and prediction of COVID-19 spread [19,20,21,22]. These studies were performed only for single countries in contrast to our study, which is being done on top 12 countries by infection incidence across the globe. In our analysis, we have used ARIMA along with the exponential smoothing method and obtained the best fitted time series models for different countries under consideration using expert modeler. In our study, Holt, Brown and ARIMA have been found as the best fitted time series models for different countries. From the best fitted time series regression models, we get the insight to make the best policies for further spread of the infectious disease and we may plan to get it down. Further as the time series regression model has capability to predict the future values of the phenomenon under consideration, thus on the basis of the predicted values of the infectious cases we may prepare ourselves to fight against it with some robust policies.
It’s critical to explain uncertainty in the parameters regulating transmission dynamics, as well as its impact on the expected outcomes. Without it, decision-makers will be missing important information and may mistakenly believe they have a high level of precision. The structure of infectious disease transmission models, the inference technique, and the usage of data streams to underpin these models must all evolve as we get a better understanding of the SARS-CoV-2 virus and the COVID-19 disease it causes. There is a large amount of literature describing mathematical models for various infectious outbreaks and the inference of associated parameters from epidemiological data. However, in most cases, these models are fitted retrospectively, using all of the data collected during an outbreak. In many cases, fitting models with such hindsight is significantly more accurate than making predictions in real time. There are additional issues connected with the quick flow of precise and accurate data when models are applied during active epidemics. Even if reliable models and procedures existed at the outset of an outbreak, there are still considerable delays in receiving, processing, and inferring parameters from incoming data. In conclusion, if epidemiological models are to be utilized in scientific discussions about how to contain a disease epidemic, it is critical that they reflect current biological knowledge and are constantly matched to all available data in real-time. Our research on COVID-19 dissemination, which is presented here, demonstrates some of the insights in anticipating a novel outbreak in a fast changing environment. From Table 3, it may be inferred that countries Afghanistan, Austria, Czech Republic, Japan, Mexico, Pakistan, Philippines, South Africa, South Korea, and Thailand are having \(R_{0}\) in the interval {1.0, 1.2} while Belgium has 1.8 and India has the highest as 2.3. The graphs of SIR model for all countries are also presented in Fig. 3. The results obtained in Tables 1, 2, and 3 may be utilized in making the best and effective policies for breaking the further dissemination of the COVID-19 infectious disease. Researchers have fitted the SIR epidemiological compartmental model for the analysis of COVID-19 infectious disease as it seems the most suitable compartmental model in case of COVID-19 infectious disease [33, 35, 36]. We have also fitted the SIR model for the analysis of COVID-19 so that the best measures and policies may be made to handle the spread of the pandemic.
The presented study was carried out amid COVID-19 pandemic on the basis of the availability of the global data on the most relevant factors responsible for the disease outbreak on country scale to make some robust policies against the current pandemic. Some of the factors which affect the disease dissemination are not considered in the study due to the unavailability of such specific data for these countries. The asymptomatic persons may be responsible for the high rate of transmission of the disease. Amid the COVID-19 pandemic, it is difficult to identify the asymptomatic persons. This identification is only possible by contact tracing and mandatory testing of every exposed person. However, some authors have considered the asymptomatic cases as well but for smaller geographical areas and/or for small scale data [44,45,46]. This study was carried out using the available official data released by the top incidence countries. There may be other relevant data, which may be important to analyze, e.g., the possible underestimation of the deaths due to pandemic. Few of the authors have attempted to calculate the possible underestimation of the deaths which may affect the model parameters and hence the outputs as well. However, such studies are dependent upon projected total deaths parameter which includes deaths caused by other causes and diseases as well, which were further subtracted from the total deaths to estimate the genuine number of deaths caused by the COVID-19 exclusively. In such cases the studies report that the projected total deaths by other causes and diseases were found to be lower than the reported data, thereby making the COVID-19 deaths underestimated [47,48,49,50]. Although, such data may not be available for all countries, therefore, it is difficult to apply it to all countries.
A COVID-19 diagnostic test may produce a false-negative result. This means that the virus was not detected by the test, even though the person was infected with it. If one person has symptoms and her/his test is false-negative, he/she may unknowingly spread the virus to others if proper precautions are not taken, such as wearing a face mask when appropriate. If the instructions are not followed carefully, a COVID-19 rapid antigen test may produce false-positive results. False-positive results indicate that the test results show an infection when there is none. The possibility of false-negative or false-positive test results is determined by the type and sensitivity of the COVID-19 diagnostic test, the thoroughness of sample collection, and the accuracy of the lab analysis. One should be careful about any offers for at-home COVID-19 tests that have not been approved by the Food & Drug Administration, USA. Such tests frequently produce inaccurate results. Thus false-negative diagnosis may affect the estimate of the model parameters. In many of the countries with large population such as India and China, 100% testing of the entire population is not possible after community spread due to lack of health infrastructure and trained human resource. However, again, it is difficult to estimate the extent of false-negative/positive cases in such large countries [51,52,53,54].
The statistical estimates of infection could be affected by the confounding factors such as pollution, temperature, relative humidity, population dynamics, age groups, co-morbidities, restriction policies, and health systems. Many of the other studies where the data is being collected for specific small regions have been studied in these contexts [55,56,57,58]. However, our study is an attempt to estimate the infection on global scale after the appearance of Omicron variant. Further the estimates of \(R_{0}\), β, and γ may vary if we do not take into account the confounding factors, but in order to make quick policy decisions in the middle of the pandemic using the overall available data and impose restrictions so that the pandemic can be contained, we have conducted this study using the most influencing factors. It will be interesting to conduct such studies in future, where we may incorporate the confounding factors in our model. If specific data is available for smaller geographical units and populations, these factors may play crucial roles in determining the dynamics of the infection spread [59,60,61,62,63].
Conclusion
In this work, from the distribution fitting, we have calculated daily average along with the variance confirmed cases in the respective countries. After knowing the best fitted distribution, we can calculate rest of the important parameters such as mode, median, coefficient of skewness, and coefficient of kurtosis, which may help in simulating the future incidence of infection and dimension of spread of the pandemic. We have also predicted the confirmed cases using time series analysis based best fitted time series models for the different countries. This gave indication of the nature of the spread. In the final part of the work, we have used epidemiological compartmental model, SIR. As this model is showing its very good fitting for all the considered countries, therefore, the estimates of infection rates (β), recovery rates (γ) and the Basic Reproduction Numbers (\(R_{0}\)) are very close to reality for these countries. Since, the value for \(R_{0}\) for all the countries are greater than one, therefore, after further validation of our findings by more research, it may be suggestive that some suppressive measures should have been applied at this point of time, when the data was collected and these measures should be applicable till the values of \(R_{0}\) would not become less than one. These modeling approaches may be used to design policies for restriction of future waves of the ongoing pandemic and any future pandemics.
References
IHME COVID-19 Forecasting Team (2021) Modeling COVID-19 scenarios for the United States. Nat Med 2021 27(1):94–105 https://doi.org/10.1038/s41591-020-1132-9
Kandeel M, Mohamed MEM, Abd El-Lateef HM, Venugopala KN, El-Beltagi HS (2021) Omicron variant genome evolution and phylogenetics. J Med Virol. https://doi.org/10.1002/jmv.27515
WHO. (2022). Tracking sars-COV-2 variants. World Health Organization. Retrieved June 16, 2022, from https://www.who.int/activities/tracking-SARS-CoV-2-variants
Araf Y, Akter F, Tang YD, Fatemi R, Parvez MSA, Zheng C, Hossain MG (2022) Omicron variant of SARS-CoV-2: Genomics, transmissibility, and responses to current COVID-19 vaccines. J Med Virol 94(5):1825–1832. https://doi.org/10.1002/jmv.27588
Kannan SR, Spratt AN, Sharma K, Chand HS, Byrareddy SN, Singh K (2022) Omicron SARS-CoV-2 variant: Unique features and their impact on pre-existing antibodies. J Autoimmun 126:102779. https://doi.org/10.1016/j.jaut.2021.102779
Akhter, Y. (2022). How are we going to defeat Omicron? Science Reporter. Retrieved June 16, 2022, from http://sciencereporter.niscair.res.in/home/article/619?fbclid=IwAR1D9Y-4dbEKSzzJyE5qks4eNV3CTCxuRnPVg-PY6A9A3Zon2DpVda1GYwg
CDC, US Department of Health and Human Services (2006) Principles of epidemiology in public health practice: an introduction to applied epidemiology and biostatistics, Atlanta, GA: U.S. Department of Health and Human Services, Centers for Disease Control and Prevention
Liang K (2020) Mathematical model of infection kinetics and its analysis for COVID-19, SARS and MERS Infection, genetics and evolution. J Mole Epid Evolu Gen Infe Dis 82:104306. https://doi.org/10.1016/j.meegid.2020.104306
Meyer SH (2014) Power-Law models for infectious disease spread. The Annals of Applied Statistics. 8(3):1612–1639
Virlogeux VL, Tsang M, Feng TK, Fang L, Jiang VJ, H, et al (2015) Estimating the distribution of the incubation periods of human avian influenza A(H7N9) virus infections. Am J Epidemiol 182(8):723–729
Virlogeux VF, Park VJ, Wu M, Cowling JT (2016) Comparison of incubation period distribution of human infections with MERS-CoV in South Korea and Saudi Arabia. Sci Rep 6:35839. https://doi.org/10.1038/srep35839
Li MD, Bolker J, BM, (2018) Fitting mechanistic epidemic models to data: A comparison of simple Markov chain Monte Carlo approaches. Stat Meth Med Res 27(7):1956–1967
de Souza A, Aristone FF, Olaofe WA, Abreu Z, De-Oliveira-Júnio MC et al (2019) Statistical behavior of hospital admissions for respiratory diseases by probability distribution functions. J Infectious Dis Epidemiol 5(6):098. https://doi.org/10.23937/2474-3658/1510098
Valvo PS (2020) A bimodal lognormal distribution model for the prediction of COVID-19 deaths. Appl Sci 10:8500. https://doi.org/10.3390/app10238500
Unglaub RAG, Spendier K (2021) A Model for the spread of infectious diseases with application to COVID-19. Challenges 12:3. https://doi.org/10.3390/challe12010003
Vazquez A (2021) Exact solution of infection dynamics with gamma distribution of generation intervals. Phys Rev E 103(4):042306
Mubarak AES, Almetwally EM (2021) A new extension exponential distribution with applications of COVID-19 Data. J Financial Business Res 22(1):444–460
Zhang XL, Yang Y, Zhang M, Young T, Li AA, X, (2013) Comparative Study of four time series methods in forecasting typhoid fever incidence in China. PLoS ONE 8(5):e63116. https://doi.org/10.1371/journal.pone.0063116
Imai C, Armstrong B, Chalabi Z, Mangtani P, Hashizume M (2015) Time series regression model for infectious disease and weather. Environ Res 142:319–327
Chaurasia VP (2020) Application of machine learning time series analysis for prediction COVID-19 pandemic. Res Biomed Eng. https://doi.org/10.1007/s42600-020-00105-4
Chyon FA, Suman MNH, Fahim MRI, Ahmmed MS (2022) Time series analysis and predicting COVID-19 affected patients by ARIMA model using machine learning. J Virol Methods 301:114433
Kibria HB, Jyoti O, Matin A (2022) Forecasting the spread of the third wave of COVID-19 pandemic using time series analysis in Bangladesh. Informatics in Medicine Unlocked 28:100815
Cassels S, Clark SJ, Morris M (2008) Mathematical models for HIV transmission dynamics: tools for social and behavioral science research. J Acq Imm Def Synd 1(47):S34–S39. https://doi.org/10.1097/QAI.0b013e3181605da3
Andraud M, Hens N, Marais C, Beutels P (2012) Dynamic epidemiological models for dengue transmission: a systematic review of structural approaches. PLoS ONE 7(11):e49085. https://doi.org/10.1371/journal.pone.0049085
Huppert AK (2013) Mathematical modelling and prediction in infectious disease epidemiology. Clin Microbiol Infect 19:999–1005
Xia Z-Q, Wang S-F, Li S-L, Huang L-Y, Zhang W-Y, Sun G-Q, Gai Z-T, Jin Z (2015) Modeling the transmission dynamics of Ebola virus disease in Liberia. Sci. Reports. 5:13857. https://doi.org/10.1038/srep13857
Drake JM, Bakach I, Just MR, O’Regan SM, Gambhir M, Fung ICH (2015) Transmission models of historical Ebola outbreaks. Emerg Infect Dis 21(8):1447
Atinuke, B., & Bagbe, A. S. (2019) Statistical analysis of ebola virus disease outbreak in some west africa countries using sir model
Hayman DT, Sam John R, Rohani P (2022) Transmission models indicate Ebola virus persistence in non-human primate populations is unlikely. J R Soc Interface 19(187):20210638
Driessche PVD (2017) Reproduction numbers of infectious disease models. Infectious Disease Modelling. 2:288–303
Li YW, Peng B, Zhou R, Zhan C, Liu Y, Z, et al (2020) Mathematical modeling and epidemic prediction of COVID-19 and its significance to epidemic prevention and control measures. Annals Infectious Disease Epidemiol 5(1):1052
Popov GN, O. (2021) An epidemic model of COVID-19 disease with variable spreading. Appl Math Eng Economics AIP Conf Proc 10:0041742
Moein S, Nickaeen N, Roointan A, Borhani NH, Javanmard SH et al (2021) Inefficiency of SIR models in forecasting COVID-19 epidemic: a case study of Isfahan. Sci Rep 11:4725
Yang Q, Yi C, Vajdi A, Cohnstaedt LW, Wu H, Guo X, Scoglio CM (2020) Short-term forecasts and long-term mitigation evaluations for the COVID-19 epidemic in Hubei Province. China, Infectious Disease Modelling 5:563–574
Alvarez MM, González-González ET, Santiago G (2021) Modeling COVID-19 epidemics in an Excel spreadsheet to enable first-hand accurate predictions of the pandemic evolution in urban areas. Scientific Rep 11:4327
Wang J, Liu Y, Liu X, Shen K (2022) A Modified SIR model for the COVID-19 epidemic in China. J Phys: Conf Ser 2148:012002
Box GJ, Reinsel G, G. (1994) Time Series Analysis: Forecasting and Control, 3rd edn. Prentice Hall, Canada
Promprou SJ, Jaroensutasinee M (2006) Forecasting dengue haemorrhagic fever cases in southern Thailand using ARIMA models. Dengue Bull 30:99–106
Liu QL, Jiang X, Yang B (2011) Forecasting incidence of hemorrhagic fever with renal syndrome in china using ARIMA model. BMC Infe Dis 11(1):218
Coutín, MG. Use of ARIMA models for communicable disease surveillance. Cub Jour Pub Health (2007) 33(2)
Wold, H. (1938) A study in the analysis of stationary time series. Stockholm: Almqvist and Wiksell
Box GJ, G. (1976) Time Series Analysis Forecasting and Control. Holden Day, San Francisco, USA
Kermack WOM (1927) A contribution to the mathematical theory of epidemics. Proc R Soc Lond 115:722. https://doi.org/10.1098/rspa.1927.0118
Sah P, Fitzpatrick MC, Zimmer CF, Abdollahi E, Juden-Kelly L, Moghadas SM et al (2021) Asymptomatic SARS-CoV-2 infection: A systematic review and meta-analysis. Proceedings of National Academy of Science, USA 118(34):e2109229118. https://doi.org/10.1073/pnas.2109229118
Ooi EE, Low JG (2020) Asymptomatic SARS-CoV-2 infection. The Lancet. https://doi.org/10.1016/S1473-3099(20)30460-6
Almadhi MA, Abdulrahman A, Sharaf SA, AlSaad D, Stevenson NJ, Atkin SL et al (2021) The high prevalence of asymptomatic SARS-CoV-2 infection reveals the silent spread of COVID-19. Int J Infect Dis 105:656–661
Kung S, Doppen M, Black M, Braithwaite I, Kearns C, Weatherall M et al (2021) Underestimation of COVID-19 mortality during the pandemic. ERJ Open Research 7:00766–02020. https://doi.org/10.1183/23120541.00766-2020
Adam D (2022) COVID’s true death toll: much higher than ocial records. Nature 603:562. https://doi.org/10.1038/d41586-022-00708-0
Jha P, Deshmukh Y, Tumbe C, Suraweera A, Bhowmick A, Sharma S et al (2021) COVID mortality in India: National survey data and health facility deaths. Science 375:667–671
Guilmoto CZ (2022) An alternative estimation of the death toll of the Covid-19 pandemic in India. PLoS ONE 17(2):e0263187. https://doi.org/10.1371/journal.pone.0263187
Itamura K, Wu A, Illing E, Ting J, Higgins T (2021) YouTube Videos Demonstrating the Nasopharyngeal Swab Technique for SARS-CoV-2 Specimen Collection: Content Analysis. JMIR Public Health Surveill 7(1):e24220. https://doi.org/10.2196/24220
Bhattacharyya R, Kundu R, Bhaduri R, Ray D, Beesley LJ, Salvatore M et al (2021) Incorporating false negative tests in epidemiological models for SARS-CoV-2 transmission and reconciling with seroprevalence estimates. Sci Rep. https://doi.org/10.1038/s41598-021-89127-1
Mouliou DS, Gourgoulianis KI (2021) False-positive and false-negative COVID-19 cases: respiratory prevention and management strategies, vaccination, and further perspectives. Expert Rev Respir Med. https://doi.org/10.1080/17476348.2021.1917389
Bhaduri R, Kundu R, Purkayastha S, Kleinsasser M, Beesley LJ, Mukherjee B et al (2022) Extending the susceptible-exposed-infected-removed (SEIR) model to handle the false negative rate and symptom-based administration of COVID-19 diagnostic tests: SEIR-fansy. Stat Med 41:2317–2337
Pluchino A, Biondo AE, Giurida N, Inturri G, Latora V, Le MR, Rapisarda A et al (2021) Author correction: a novel methodology for epidemic risk assessment of COVID-19 outbreak. Sci Rep 11(1):15719. https://doi.org/10.1038/s41598-021-94234-0.Erratumfor:ScientificReports,11(1),5304
Fontal A, Bouma MJ, San-José A et al (2021) Climatic signatures in the dierent COVID-19 pandemic waves across both hemispheres. Nat-Comput Sci 1:655–665. https://doi.org/10.1038/s43588-021-00136-6
Kifer D, Bugada D, Villar-Garcia J, Gudelj I, Menni C, Sudre C et al (2021) Effects of environmental factors on severity and mortality of COVID-19. Front Med 7:607786. https://doi.org/10.3389/fmed.2020.607786
Maiti A, Zhang Q, Sannigrahi S, Pramanik S, Chakraborti S, Cerda A et al (2021) Exploring spatiotemporal effects of the driving factors on COVID-19 incidences in the contiguous United States. Sustain Cities Soc 68:102784. https://doi.org/10.1016/j.scs.2021.102784
Flynn D, Moloney E, Bhattarai N, Scott J, Breckons M, Avery L et al (2020) COVID-19 pandemic in the United Kingdom. Health Policy Technology 9(4):673–691. https://doi.org/10.1016/j.hlpt.2020.08.003
Rovetta A, Bhagavathula AS, Castaldo L (2020) Modeling the epidemiological trend and behavior of COVID-19 in Italy. Cureus 12(8):e9884. https://doi.org/10.7759/cureus.9884.Erratum.In:Cureus,12(9),c37
Naik PA, Zu J, Ghori MB, Naik M (2021) Modeling the effects of the contaminated environments on COVID-19 transmission in India. Results in Physics 29:104774. https://doi.org/10.1016/j.rinp.2021.104774
Kong JD, Tekwa EW, Gignoux-Wolfsohn SA (2021) Social, economic, and environmental factors influencing the basic reproduction number of COVID-19 across countries. PLoS ONE 16(6):e0252373. https://doi.org/10.1371/journal.pone.0252373
Nakada LYK, Urban RC (2021) COVID-19 pandemic: environmental and social factors influencing the spread of SARS-CoV-2 in São Paulo Brazil. Environ Sci Pollution Res 28:40322–40328
Acknowledgements
We are thankful to Babasaheb Bhimrao Ambedkar University for providing us the basic research infrastructure, which helped us in carrying out this work.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or nonprofit sectors.
Author information
Authors and Affiliations
Contributions
SKY and YA have conceived the idea. VK has retrieved the data. SKY and VK have performed the analysis. SKY wrote the first draft. SKY and YA edited the final version and coordinated the overall study.
Corresponding authors
Ethics declarations
Conflict of interest
We do not have any conflicts of interests to report.
Ethical Approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yadav, S.K., Kumar, V. & Akhter, Y. Modeling Global COVID-19 Dissemination Data After the Emergence of Omicron Variant Using Multipronged Approaches. Curr Microbiol 79, 286 (2022). https://doi.org/10.1007/s00284-022-02985-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-02985-4