Abstract
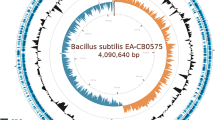
Bacillus sp. TYF-LIM-B05, which is isolated from spoilage vinegar, is resistant to high temperature, high concentrated alcohol, acid, and salt, and can produce ethanol from mono-, di-, polysaccharide, and complex biomass as the sole carbon source. Thus, this strain is a potential candidate for consolidated bioprocessing (CBP) of fermenting lignocellulose to ethanol in a single step. To provide insight into the key enzymes involved in lignocellulose degradation and ethanol production, a draft genome of TYF-LIM-B05 was developed in this study. The results indicated that 348 genes are related to carbohydrate transport and metabolism according to the clusters of orthologous groups of proteins and annotated 187 CAZy domains from a total of 61 different families. The presence of genes encoding laccases, quinone oxidoreductases/reductases, and aryl-alcohol dehydrogenases further implies that TYF-LIM-B05 has the potential to degrade lignin. Remarkably, this strain has the ability to catalyze the oxidative decarboxylation of pyruvate to acetyl-CoA by pyruvate dehydrogenase complexes. The genomic information provided in this study will help researchers to better understand the mechanisms of the lignocellulose degradation and ethanol production pathway in thermophiles.






Similar content being viewed by others
References
Almarsdottir AR, Sigurbjornsdottir MA, Orlygsson J (2012) Effect of various factors on ethanol yields from lignocellulosic biomass by Thermoanaerobacterium AK17. Biotechnol Bioeng 109(3):686–694. https://doi.org/10.1002/bit.24346
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Andlar M, Rezic T, Mardetko N, Kracher D, Ludwig R, Santek B (2018) Lignocellulose degradation: an overview of fungi and fungal enzymes involved in lignocellulose degradation. Eng Life Sci 18(11):768–778. https://doi.org/10.1002/elsc.201800039
Arora R, Behera S, Kumar S (2015) Bioprospecting thermophilic/thermotolerant microbes for production of lignocellulosic ethanol: a future perspective. Renew Sustain Energy Rev 51:699–717. https://doi.org/10.1016/j.rser.2015.06.050
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29. https://doi.org/10.1038/75556
Bhattacharya AS, Bhattacharya A, Pletschke BI (2015) Synergism of fungal and bacterial cellulases and hemicellulases: a novel perspective for enhanced bio-ethanol production. Biotechnol Lett 37(6):1117–1129. https://doi.org/10.1007/s10529-015-1779-3
Boch J, Nau-Wagner G, Kneip S, Bremer E (1997) Glycine betaine aldehyde dehydrogenase from Bacillus subtilis: characterization of an enzyme required for the synthesis of the osmoprotectant glycine betaine. Arch Microbiol 168(4):282–289
Currie DH, Raman B, Gowen CM, Tschaplinski TJ, Land ML, Brown SD, Covalla SF, Klinge-man DM, Yang ZK, Engle NL, Johnson CM, Rodriguez M, Shaw AJ, Kenealy WR, Lynd LR, Fong SS, Mielenz JR, Davison BH, Hogsett DA, Herring CD (2015) Genome-scale resources for Thermoanaerobacterium saccharolyticum. BMC Syst Biol 9(1):30. https://doi.org/10.1186/s12918-015-0159-x
De Vrije T, Bakker RR, Budde MAW, Lai MH, Mars AE, Claassen PAM (2009) Efficient hydrogen production from the lignocellulosic energy crop Miscanthus by the extreme thermophilic bacteria Caldicellulosiruptor saccharolyticus and Thermotoga neapolitana. Biotechnol Biofuels 2(1):1–15. https://doi.org/10.1186/1754-6834-2-12
Eram MS, Ma K (2013) Decarboxylation of pyruvate to acetaldehyde for ethanol production by hyperthermophiles. Biomolecules 3(3):578–596. https://doi.org/10.3390/biom3030578
Ezeilo UR, Zakaria II, Huyop F, Wahab RA (2017) Enzymatic breakdown of lignocellulosic biomass: the role of glycosyl hydrolases and lytic polysaccharide monooxygenases. Biotechnol Biotechnol Equip 31(4):647–662. https://doi.org/10.1080/13102818.2017.1330124
Gardner PP, Daub J, Tate JG, Nawrocki EP, Kolbe DL, Lindgreen S, Wilkinson AC, Finn RD, Griffiths-Jones S, Eddy SR, Bateman A (2009) Rfam: updates to the RNA families database. Nucleic Acids Res 37:D136–D140. https://doi.org/10.1093/nar/gkn766
Grissa I, Vergnaud G, Pourcel C (2007) CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res 35:W52–W57. https://doi.org/10.1093/nar/gkm360
Hsiao W, Wan I, Jones SJ, Brinkman FSL (2003) IslandPath: aiding detection of genomic islands in prokaryotes. Bioinformatics 19(3):418–420. https://doi.org/10.1093/bioinformatics/btg004
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280. https://doi.org/10.1093/nar/gkh063
Kuhad RC, Gupta R, Khasa YP, Singh A, Zhang YHP (2011) Bioethanol production from pentose sugars: current status and future prospects. Renew Sustain Energy Rev 15(9):4950–4962. https://doi.org/10.1016/j.rser.2011.07.058
Laurent CVFP, Breslmayr E, Tunega D, Ludwig R, Oostenbrink C (2019) Interaction between cellobiose dehydrogenase and lytic polysaccharide monooxygenase. Biochemistry 58(9):1226–1235. https://doi.org/10.1021/acs.biochem.8b01178
Li R, Li Y, Kristiansen K, Wang J (2008) SOAP: short oligonucleotide alignment program. Bioinformatics 24(5):713–714. https://doi.org/10.1093/bioinformatics/btn025
Li R, Zhu H, Ruan J, Qian W, Fang X, Shi Z, Li Y, Li S, Shan G, Kristiansen K, Li S, Yang H, Wang J, Wang J (2010) De novo assembly of human genomes with massively parallel short read sequencing. Genome Res 20(2):265–272. https://doi.org/10.1101/gr.097261.109
Lim HJ, Lee EH, Yoon Y, Chua B, Son A (2016) Portable lysis apparatus for rapid single-step DNA extraction of Bacillus subtilis. J Appl Microbiol 120(2):379–387. https://doi.org/10.1111/jam.13011
Ma K, Loessner H, Heider J, Johnson MK, Adams MW (1995) Effects of elemental sulfuron the metabolism of the deep-sea hyperthermophilic archaeon Thermococcus strain ES-1: characterization of a sulfur-regulated, non-heme iron alcohol dehydrogenase. J Bacteriol 177(16):4748–4756. https://doi.org/10.1128/jb.177.16.4748-4756.1995
Margeot A, Hahn-Hagerdal B, Edlund M, Slade R, Monot F (2009) New improvements for lignocellulosic ethanol. Curr Opin Biotechnol 20(3):372–380. https://doi.org/10.1016/j.copbio.2009.05.009
Mistry FR, Cooney CL (1989) Production of ethanol by Clostridium thermosaccharolyticum: I. Effect of cell recycle and environmental parameters. Biotechnol Bioeng 34(10):1295–1304. https://doi.org/10.1002/bit.260341008
Nawrocki EP, Kolbe DL, Eddy SR (2009) Infernal 1.0: inference of RNA alignments. Bioinformatics 25(13):1713–1713. https://doi.org/10.1093/bioinformatics/btp326
Rastogi M, Shrivastava S (2017) Recent advances in second generation bioethanol production: an insight to pretreatment, saccharification and fermentation processes. Renew Sustain Energy Rev 80:330–340. https://doi.org/10.1016/j.rser.2017.05.225
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evolut 4(4):406–442. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Scully SM, Orlygsson J (2015) Recent advances in second generation ethanol production by Thermophilic Bacteria. Energies 8(1):1–30. https://doi.org/10.3390/en8010001
Singh N, Mathur AS, Tuli DK, Gupta RP, Barrow CJ, Puri M (2017) Cellulosic ethanol productionvia consolidated bioprocessing by a novel thermophilic anaerobic bacterium isolated from a Hima-layan hot spring. Biotechnol Biofuels 10(1):73. https://doi.org/10.1186/s13068-017-0756-6
Tamaru Y, Miyake H, Kuroda K, Ueda M, Doi RH (2010) Comparative genomics of the m-esophilic cellulosome-producing Clostridium cellulovorans and its application to biofuel product-ion via consolidated bioprocessing. Environ Technol 31(8–9):889–903. https://doi.org/10.1080/09593330.2010.490856
Taylor MP, Eley KL, Martin S, Tuffin MI, Burton SG, Cowan DA (2009) Thermophilic ethanologenesis: future prospects for second-generation bioethanol production. Trends Biotechnol 27(7):398–405. https://doi.org/10.1016/j.tibtech.2009.03.006
Velasco-Garcia R, Villalobos M, Ramirez-Romero M, Mujica-Jimenez C, Iturriaga G, Munoz-Clares R (2006) Betaine aldehyde dehydrogenase from Pseudomonas aeruginosa: cloning, over-expression in Escherichia coli, and regulation by choline and salt. Arch Microbiol 185(1):14–22. https://doi.org/10.1007/s00203-005-0054-8
Wiegel J, Ljungdahl LG (1981) Thermoanaerobacter ethanolicus gen. nov., spec. nov., a new, extreme thermophilic, anaerobic bacterium. Arch Microbiol 128(4):343–348. https://doi.org/10.1007/bf00405910
Witzmann S, Bisswanger H (1998) The pyruvate dehydrogenase complex from thermophilic organisms: thermal stability and re-association from the enzyme components. Biochim Biophys Acta 1385(2):341–352. https://doi.org/10.1016/s0167-4838(98)00078-8
Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS (2011) PHAST: a fast phage search tool. Nucleic Acids Res 39:W347–W352. https://doi.org/10.1093/nar/gkr485
Acknowledgements
We thank the Taiyuan Ninghuafu Yiyuanqing Vinegar Industry for providing spoilage vinegar samples. This study was funded by the Foundation of Shanxi Province (Grant Nos. 20180008, 201703D121044).
Nucleotide Sequence Accession Number
The whole genome sequencing has been deposited at GenBank under the accession no.QXII00000000. The version described in this paper is QXII00000000.1.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fan, L., Li, M., Li, Y. et al. Draft Genome Sequence of Thermophilic Bacillus sp. TYF-LIM-B05 Directly Producing Ethanol from Various Carbon Sources Including Lignocellulose. Curr Microbiol 77, 491–499 (2020). https://doi.org/10.1007/s00284-019-01833-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01833-2