Abstract
Several molecular aberrations affect the prognosis of patients with acute myeloid leukemia (AML) and myelodysplastic syndrome (MDS) with excess blasts (EB). This study aimed to determine the incidence and clinical impact of molecular genetic aberrations in Thai patients with AML and MDS-EB, detected by the next-generation sequencing (NGS) technique. This prospective, observational study was conducted between 2018 and 2020 on newly diagnosed Thai AML or MDS-EB patients aged above 15 years. NGS was performed using a custom amplicon-based targeted enrichment assay for 42 genes recurrently mutated in myeloid neoplasms. The molecular results were correlated with baseline patient and disease characteristics as well as outcomes. Forty-nine patients were enrolled in this study. The median age was 56 years (interquartile range [IQR], 44–64), with nearly equal proportions of males and females. The median number of mutations was 3 (IQR, 2–4). The most frequent alterations were FLT3 internal tandem duplications (ITD) (28.6%), DNMT3A (24.5%), and WT1 (22.4%) mutations. FLT3-ITD was more frequent in the de novo AML group than in the MDS/secondary AML group, whereas in the MDS/secondary AML group, ASXL1, ETV6, and SRSF2 mutations were more frequent. Patients aged greater than 65 years and patients with mutated TP53 were more likely to have inferior overall survival from multivariate analysis. FLT3-ITD was the most common mutation among newly diagnosed Thai AML patients. TP53 mutation and advanced age were independent adverse factors for survival outcome. The genetic landscapes of AML patients vary between national populations. Thai Clinical Trials Registry identifier: TCTR20190227003.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Currently, two main standard recommendations for acute myeloid leukemia (AML), published by the European LeukemiaNet (ELN) and the National Comprehensive Cancer Network, are being used to determine patients’ risk stratification, and to select the appropriate therapy [1, 2]. Stratification of risk at the time of diagnosis is primarily based on cytogenetic and molecular genetic findings. Results from the genetic workup are also essential for determining and guiding an appropriate, long-term, treatment strategy. For instance, patients with AML harboring NPM1 mutation and FLT3 wildtype or FLT3-ITD with low allelic ratio, AML with biallelic CEBPA mutation, or with core-binding factor AML are categorized within the favorable risk group and may require only induction chemotherapy followed by consolidation chemotherapy [1, 2]. In contrast, allogeneic stem cell transplantation after induction therapy is generally recommended for patients with adverse genetic profiles, such as complex karyotype or mutated RUNX1, ASXL1, or TP53 [1,2,3,4,5,6].
In addition, personalized treatment is widely integrated with AML treatment strategies, depending on mutational status; for example, the FLT3 inhibitor midostaurin combined with intensive induction and consolidation chemotherapy followed by a 1-year maintenance therapy yielded significantly better survival outcomes of patients with FLT3-mutated AML [7, 8]. The IDH1 inhibitor ivosidenib and IDH2 inhibitor enasidenib show encouraging clinical activity in patients with IDH-mutated AML [9, 10]. Furthermore, myelodysplastic syndrome (MDS) with excess blasts-2 (EB2) shares a similar natural history with AML [11]. Therefore, a comprehensive genetic investigation of MDS-EB patients is required, just as for patients with AML.
Reverse transcriptase-polymerase chain reaction (RT-PCR) is a conventional genetic testing method that can be performed by the majority of institutes. Nevertheless, the main limitation of this technique is that each mutation has to be evaluated separately, which is time-consuming. This is in particular clinically relevant for patients with newly diagnosed AML and MDS-EB. Over the last years, the next-generation sequencing (NGS) technique was introduced for myeloid neoplasms [12, 13]. The technique can be used to evaluate several target genes within a few days. Health-care institutes, especially those in developed countries, are increasingly adopting the NGS method to investigate the mutational status of newly diagnosed AML and MDS-EB patients. However, comprehensive genetic profiles for the Southeast Asian population, including Thais, have not been fully studied.
We, therefore, performed this study primarily to determine the incidence of molecular aberrations in Thai patients with AML and MDS-EB, as detected by the NGS technique. The secondary objective was to correlate molecular mutational status with clinical outcomes and to evaluate the genetic landscape differences between Thais and other ethnic groups of AML and MDS patients.
Methods
This prospective observational study was conducted on newly diagnosed Thai AML and MDS-EB patients between January 2018 and March 2020. The inclusion criteria were as follows: (1) patients aged above 15 years; (2) patients with de novo AML, secondary AML, or MDS-EB; and (3) patients requiring treatment and follow-up at Siriraj Hospital, Thailand. Every participant signed a consent form before enrollment. The mononuclear cells from bone marrow specimens were collected and cryopreserved in a biobank. The genomic deoxyribonucleic acid (gDNA) was isolated using the QIAGEN Genomic DNA extraction kit (Qiagen, Hilden, Germany). The qualities and concentrations of gDNA were confirmed by gel electrophoresis and a Qubit 3.0 Fluorometer (life technologies by Thermo Fisher Scientific, Waltham, MA, USA). All of the gDNA products were transferred to the University Hospital of Ulm, Germany, for the performance of the molecular study. The Siriraj Institutional Review Board approved this research, which followed the Declaration of Helsinki guidelines and all subsequent amendments. The study was approved for registration at the Thai Clinical Trial Registry and the identification number is TCTR20190227003. Molecular studies were supported by the Department of Internal Medicine III, University Hospital of Ulm, Ulm, Germany.
NGS library preparation and data interpretation
The NGS covered entire coding regions of 42 genes recurrently mutated in myeloid neoplasms by using a custom amplicon-based targeted enrichment assay (HaloplexHS Target Enrichment System, Agilent, Santa Clara, CA, USA). The library preparation was performed according to the manufactures’ instructions. The library products were sequenced on a MiSeq sequencer using the 300-cycle MiSeq Reagent Kit v2 (Illumina, San Diego, CA, USA). Following demultiplexing, the paired-end sequences were analyzed by an in-house data analysis workflow. In brief, sequences were aligned to the human reference genome GRCh37 (hg19) using BWA-MEM (version 0.7.10) [14]. Based on the molecular barcodes, duplicates were removed and consensus sequences were generated (BamDeduplicateByBarcode, ngs-bits). After local realignment by GATK (version 3.4-16) [15], variants were called using VarScan (v2.3.9) [16] and annotated by ANNOVAR [17]. Semi-automated filtering was applied with intronic variants, synonymous variants, variants with less than one tumor supporting read, and variants with an entry in the Database of Single Nucleotide Polymorphisms (dbSNP, build 138) [18], but not in the Catalogue of Somatic Mutations in Cancer (COSMIC) [19] that were filtered out. The heterozygous variant allele frequency (VAF) detection threshold was 3%. The remaining variants were manually analyzed and curated by the Viewer Integrative Genomics (IGV, California, USA), the UCSC Genome Browser, the COSMIC database, and dbSNP [20, 21]. NPM1 and FLT3-ITD were determined by DNA-based assays, FLT3-ITD as described by Stone et al. [7].
Terminology
This study applied the 2017 ELN risk classification based on the cytogenetic and molecular findings [1]. Secondary AML in this cohort was defined as the patient with a previous history of MDS. CR was defined as < 5% blasts in the bone marrow, an absolute neutrophil count of ≥ 1.0 × 109/l, and a platelet count of ≥ 100 × 109/l, without any of the following: (1) circulating blasts, (2) blasts with Auer rods, and (3) extramedullary disease. The OS was defined as the duration from diagnosis to the time of last follow-up or death, whereas the RFS was defined as the duration from the CR date to the date of relapse or death from any cause.
Statistical analysis
Continuous data were reported as median with interquartile range (IQR) or mean ± standard deviation, as suitable. The Mann–Whitney U test or Student’s t-test was employed to compare continuous data. Categorical data were presented as number and percent, and compared using Fisher’s exact test or a chi-square test. A log-rank test was used to compare the factors correlated with OS and RFS, and it was presented as a Kaplan–Meier survival curve. Cox proportional hazards analysis (enter method) was used to compare the predictors of survival outcome in the univariate and multivariate analyses. The independent variables that have significance in univariate analysis entered the multivariate model. The results were expressed as hazard ratio (HR) and 95% CI. Statistical significance was determined as a p-value of < 0.05. The program IBM SPSS Statistics for Windows, version 20.0 (Armonk, NY: IBM Corp.), was utilized to analyze the data.
Results
Forty-nine cases were enrolled in this study. The median age was 56 years (IQR, 44–64), with almost equal proportions of males and females. Forty-six (93.9%) patients had AML in which 39 patients (79.6%) were de novo and 7 patients (14.3%) were secondary AML. Two acute promyelocytic leukemia (APL) patients were found in these AML patients. The others were MDS-EB, accounting for 3 (6.1%) patients. The common initial clinical manifestations included anemia symptoms (87.8%), fever (40.8%), bleeding (28.6%), and significant weight loss (24.5%). Table 1 summarizes the baseline patient, disease characteristics, and cytogenetic risks.
Molecular landscape results
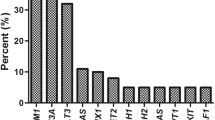
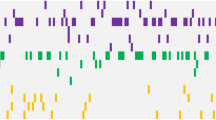
The median number of gene mutations was 3 (IQR, 2–4). The most common mutations were FLT3-ITD (28.6%, as assessed by conventional diagnostic assay), DNMT3A (24.5%), WT1 (22.4%), TET2 (20.4%), RUNX1 (18.4%), NPM1 (16.3%), FLT3-TKD (14.3%), monoallelic CEBPA (10.2%), and biallelic CEBPA (10.2%). The favorable molecular genotypes NPM1 mutation without FLT3-ITD or with FLT3-ITDlow and the biallelic CEBPA mutation represented 6.1% and 10.2% of cases, respectively. The targeted gene sequencing results of the 49 cases, classified by functional gene, [1] are illustrated in Fig. 1.
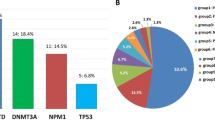
Because the genetic mutations and pathogenesis of MDS and secondary AML are very similar [11], we divided the patients into two groups: a de novo AML group (n=39), and an MDS combined with a secondary AML group (n=10). The common genetic alterations of the de novo group were FLT3-ITD (35.9%), monoallelic or biallelic CEBPA (25.6%), WT1 (25.6%), DNMT3A (23.1%), NPM1 (20.5%), TET2 (20.5%), and FLT3-TKD (15.4%), while the common ones in the latter group were RUNX1 (40.0%), ASXL1 (30.0%), DNMT3A (30.0%), ETV6 (30.0%), NRAS (30.0%), STAG2 (30.0%), SRSF2 (30.0%), BCOR (20.0%), TET2 (20.0%), TP53 (20.0%), and U2AF1 (20.0%). Figure 2 displays the frequencies of the genetic alterations found in the Thai patients categorized by disease type. As to the baseline patient characteristics of the two groups, significantly higher white blood cell counts and higher percentages of peripheral blood blasts and marrow blasts were detected in the de novo AML group than in the MDS/secondary AML group. A comparison of the genetic profiles of the two groups revealed that the FLT3-ITD mutation was more likely to be present in the de novo AML group than in the MDS/secondary AML group (p = 0.044). In contrast, the MDS/secondary AML group more frequently had mutations in ASXL1, ETV6, and SRSF2 genes (p=0.007, 0.023, and 0.007, respectively: Table S1).
Furthermore, we compared the clinical features and mutational status of two age groups, < 65 years (n=38) and ≥ 65 years (n=11). The elderly group had a significantly greater proportion of patients with a poor Eastern Cooperative Oncology Group performance status. The common mutations in the younger group were DNMT3A (26.3%), monoallelic or biallelic CEBPA (23.7%), FLT3-ITD (23.7%), WT1 (23.7%), RUNX1 (15.8%), and TET2 (15.8%). In the elderly group, the common mutations were FLT3-ITD (45.5%), NPM1 (36.4%), TET2 (36.4%), and RUNX1 (27.3%). The proportions of all molecular aberrations found in the two age groups of the Thai patients are illustrated in Fig. 3. There was no significant difference in the proportion of all mutations between the two age groups (Table S2).
Treatment response and clinical outcomes
Sixty-one percent of patients received intensive induction therapy (7+3 regimen, n = 29; 5+2 regimen, n = 1), 10.2% hypomethylating agents (HMAs), and 4.1% APL induction. The CR rate of the patients who received intensive induction therapy reached 60%, while the induction mortality rate was 11.4%. A quarter (24.5%) of the patients received supportive treatment or low-intensive therapy, comprised of hydroxyurea, oral low-dose chemotherapy, and subcutaneous cytarabine. An allogeneic stem cell transplant after CR using a matched sibling donor was performed in only one patient because of the cost-effectiveness-based transplantation policy in Thailand, under which only matched-sibling donor allogeneic stem cell transplants are approved. There was no difference in the CR rates of the de novo AML and the MDS/secondary AML groups (63.3% and 66.7%, respectively).
The median OS time of all patients was 274 days (95% confidence interval [CI]: 199.83–348.17), with a 1-year OS rate of 33.8%; the median RFS time was 235 days (95% CI: 173.59–296.41), with a 1-year RFS rate of only 24.9%. The Kaplan–Meier survival curves for the OS and RFS outcomes of the entire cohort are illustrated in Fig. 4. A subgroup analysis of several factors impacting on survival outcome did not identify any difference in the OS outcomes of the de novo AML group and the MDS/secondary AML group (the median OS times were 303.6 days and 273.4 days, respectively; hazard ratio [HR]: 1.19; 95% CI: 0.50–2.87; p = 0.687). The patients who were aged 65 years or less had a significantly better OS than those aged over 65, with a median OS time of 355.9 days versus 135.0 days, respectively (HR: 3.51; 95% CI: 1.55–7.94; p = 0.003). The former group received intensive chemotherapy accounting for 89.5% compared to only 9.1% in the latter group. As to the mutational status analysis, the TP53 was the only mutation that showed a statistically significant difference in survival outcomes. Patients with wild-type TP53 had a median OS time of 300.4 days, whereas those with mutated TP53 had a median OS time of 105.0 days (HR: 4.45; 95% CI: 1.59–12.46; p = 0.004). Figure 5 presents the Kaplan–Meier survival curves of the patients, classified by disease type, age group, and genetic mutation. A multivariate analysis was performed to assess all possible factors that were significantly correlated to the survival time of the patients. Patients aged greater than 65 years and patients with the mutated TP53 were more likely to have an inferior OS, with the HRs of 3.22 (p = 0.006) and 4.38 (p = 0.006), respectively (Table 2).
Discussion
This is the first prospective study to comprehensively investigate the genetic profiles of Thai patients with AML and MDS-EB using the NGS technique. FLT3-ITD (assessed by conventional PCR technique) was the most common genetic mutation in this cohort. Other mutations that were commonly found in our study were mutations in DNA methylation (DNMT3A and TET2), the myeloid transcription factor (CEBPA and RUNX1), and the NPM1 gene, known to be frequently mutated in AML. However, the high proportion of WT1 mutation in our results was markedly different from the frequencies reported by other studies [22,23,24,25].
In de novo AML patients, the most common genetic mutations were FLT3-ITD, NPM1, and CEBPA, which is in line with another report from an Asian cohort [26]. DNMT3A mutation was commonly found in a high percentage especially in younger AML patients, which compares well with a previous report [27].
As to the MDS/secondary AML patients, mutations in the myeloid transcription factor (RUNX1 and ETV6), chromatin-modifying gene (ASXL1 and BCOR), spliceosome complex (SRSF2 and U2AF1), cohesion complex gene (STAG2), and tumor suppressor gene (TP53) were demonstrated in high proportions compared with the frequencies in the de novo AML patients; this finding was similar to the results of previous reports. As previously shown, these mutational groups appear to be a signature of this patient subgroup [1, 12, 28, 29].
Notably, just over 10% of the patients aged 65 years or less had mutated NPM1, whereas the rate in a previous report was as high as 30% [1]. Besides, high percentages of unfavorable mutations were found in elderly patients, including FLT3-ITD and RUNX1 mutations. These genetic profiles are possibly important adverse factors for Thai elderly patients with AML, and the profiles appear to support the published results of a study of Thai elderly patients with AML who had dismal survival outcomes, with a median OS time of only 4 months [30]. However, the percentages of mutations in the elderly group need to be interpreted with caution due to the low number of patients. The frequencies of the genetic mutations detected by the NGS technique in the AML patients in the present study and previous research are tabulated in Table 3 [4, 22,23,24, 31].
Furthermore, TP53 mutation was an independent adverse factor for outcome in our study, which is comparable with other published results [5, 28, 32]. In a previous report, decitabine has been suggested as an effective treatment for AML patients with mutated TP53 [5]. However, this observation is not supported by data from controlled clinical trials [4, 33]. New agents, such as the p53 reactivator APR-246 or the monoclonal anti-CD47 antibody magrolimab that have shown promising activity in this high-risk subset of AML, are currently in clinical development [34, 35]. Apart from the limited transplant access, the poor OS and RFS rates of our patients were possible because age > 65 years and unfavorable genetic mutations (especially the TP53 mutation) were adverse factors. The reason why the old age in this study confers grave prognosis could be explained by less availability of HMAs utilization.
Overall, there are different genetic abnormalities in AML patients in each race. A comprehensive genetic investigation of AML and MDS patients could categorize patients’ risks and prognoses. Moreover, personalized treatment based on each molecular mutation in individual patients could improve their treatment responses and long-term survival outcomes.
The main limitation of this study was the low number of enrolled patients. Consequently, several genetic mutations could not reach a level of statistical difference for outcome measures between the wild-type and the mutated gene. Establishing the exact prevalence and clinical outcome of each mutation in Thai AML patients would be interesting but would require the collection and analysis of more cases.
Conclusions
FLT3-ITD was the most common mutation in newly diagnosed Thai AML patients. TP53 mutation and advanced age were independent, poor prognostic factors for patients’ survival. The genetic landscape of AML patients for each disease type, each age group, and each nation differ; hence, a comprehensive genetic investigation should guide the most suitable treatment to improve individual patients’ outcomes.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
References
Dohner H, Estey E, Grimwade D, Amadori S, Appelbaum FR, Buchner T et al (2017) Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood 129(4):424–447. https://doi.org/10.1182/blood-2016-08-733196
National Comprehensive Cancer Network (NCCN), Acute Myeloid Leukemia (version 3.2021), Available at: https://www.nccn.org/professionals/physician_gls/pdf/aml.pdf. Accessed 18 March 2021
Hamilton BK, Rybicki L, Hirsch C, Przychodzen B, Nazha A, Gerds AT, Hanna R, Kalaycio M, Sekeres MA, Sobecks R, de Lima M, Majhail NS, Maciejewski J (2019) Mutation clonal burden and allogeneic hematopoietic cell transplantation outcomes in acute myeloid leukemia and myelodysplastic syndromes. Bone Marrow Transplant 54(8):1281–1286. https://doi.org/10.1038/s41409-019-0444-1
Dohner H, Dolnik A, Tang L, Seymour JF, Minden MD, Stone RM et al (2018) Cytogenetics and gene mutations influence survival in older patients with acute myeloid leukemia treated with azacitidine or conventional care. Leukemia 32(12):2546–2557. https://doi.org/10.1038/s41375-018-0257-z
Welch JS (2018) Patterns of mutations in TP53 mutated AML. Best Pract Res Clin Haematol 31(4):379–383. https://doi.org/10.1016/j.beha.2018.09.010
Gaidzik VI, Teleanu V, Papaemmanuil E, Weber D, Paschka P, Hahn J et al (2016) RUNX1 mutations in acute myeloid leukemia are associated with distinct clinico-pathologic and genetic features. Leukemia 30(11):2160–2168. https://doi.org/10.1038/leu.2016.126
Stone RM, Mandrekar SJ, Sanford BL, Laumann K, Geyer S, Bloomfield CD, Thiede C, Prior TW, Döhner K, Marcucci G, Lo-Coco F, Klisovic RB, Wei A, Sierra J, Sanz MA, Brandwein JM, de Witte T, Niederwieser D, Appelbaum FR, Medeiros BC, Tallman MS, Krauter J, Schlenk RF, Ganser A, Serve H, Ehninger G, Amadori S, Larson RA, Döhner H (2017) Midostaurin plus chemotherapy for acute myeloid leukemia with a FLT3 mutation. N Engl J Med 377(5):454–464. https://doi.org/10.1056/NEJMoa1614359
Schlenk RF, Weber D, Fiedler W, Salih HR, Wulf G, Salwender H, Schroeder T, Kindler T, Lübbert M, Wolf D, Westermann J, Kraemer D, Götze KS, Horst HA, Krauter J, Girschikofsky M, Ringhoffer M, Südhoff T, Held G, Derigs HG, Schroers R, Greil R, Grießhammer M, Lange E, Burchardt A, Martens U, Hertenstein B, Marretta L, Heuser M, Thol F, Gaidzik VI, Herr W, Krzykalla J, Benner A, Döhner K, Ganser A, Paschka P, Döhner H, German-Austrian AML Study Group (2019) Midostaurin added to chemotherapy and continued single-agent maintenance therapy in acute myeloid leukemia with FLT3-ITD. Blood 133(8):840–851. https://doi.org/10.1182/blood-2018-08-869453
DiNardo CD, Stein EM, de Botton S, Roboz GJ, Altman JK, Mims AS et al (2018) Durable remissions with ivosidenib in IDH1-mutated relapsed or refractory AML. N Engl J Med 378(25):2386–2398. https://doi.org/10.1056/NEJMoa1716984
Stein EM, DiNardo CD, Pollyea DA, Fathi AT, Roboz GJ, Altman JK et al (2017) Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood 130(6):722–731. https://doi.org/10.1182/blood-2017-04-779405
Bejar R (2018) What biologic factors predict for transformation to AML? Best Pract Res Clin Haematol 31(4):341–345. https://doi.org/10.1016/j.beha.2018.10.002
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, Potter NE, Heuser M, Thol F, Bolli N, Gundem G, van Loo P, Martincorena I, Ganly P, Mudie L, McLaren S, O’Meara S, Raine K, Jones DR, Teague JW, Butler AP, Greaves MF, Ganser A, Döhner K, Schlenk RF, Döhner H, Campbell PJ (2016) Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med 374(23):2209–2221. https://doi.org/10.1056/NEJMoa1516192
Bacher U, Shumilov E, Flach J, Porret N, Joncourt R, Wiedemann G, Fiedler M, Novak U, Amstutz U, Pabst T (2018) Challenges in the introduction of next-generation sequencing (NGS) for diagnostics of myeloid malignancies into clinical routine use. Blood Cancer J 8(11):113
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L et al (2012) VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 22(3):568–576. https://doi.org/10.1101/gr.129684.111
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164. https://doi.org/10.1093/nar/gkq603
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29(1):308–311. https://doi.org/10.1093/nar/29.1.308
Tate JG, Bamford S, Jubb HC, Sondka Z, Beare DM, Bindal N, Boutselakis H, Cole CG, Creatore C, Dawson E, Fish P, Harsha B, Hathaway C, Jupe SC, Kok CY, Noble K, Ponting L, Ramshaw CC, Rye CE, Speedy HE, Stefancsik R, Thompson SL, Wang S, Ward S, Campbell PJ, Forbes SA (2019) COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res 47(D1):D941–D947. https://doi.org/10.1093/nar/gky1015
Robinson JT, Thorvaldsdóttir H, Wenger AM, Zehir A, Mesirov JP (2017) Variant review with the integrative genomics viewer. Cancer Res 77(21):e31–e34. https://doi.org/10.1158/0008-5472.CAN-17-0337
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12(6):996–1006. https://doi.org/10.1101/gr.229102
Lin PH, Li HY, Fan SC, Yuan TH, Chen M, Hsu YH, Yang YH, Li LY, Yeh SP, Bai LY, Liao YM, Lin CY, Hsieh CY, Lin CC, Lin CH, Lien MY, Chen TT, Ni YH, Chiu CF (2017) A targeted next-generation sequencing in the molecular risk stratification of adult acute myeloid leukemia: implications for clinical practice. Cancer Med 6(2):349–360. https://doi.org/10.1002/cam4.969
Hussaini MO, Mirza AS, Komrokji R, Lancet J, Padron E, Song J (2018) Genetic Landscape of Acute Myeloid Leukemia Interrogated by next-generation sequencing: a large cancer center experience. Cancer Genomics Proteomics 15(2):121–126. https://doi.org/10.21873/cgp.20070
Cao XX, Cai H, Mao YY, Wu Q, Zhang L, Zhou DB, Li J (2018) Next-generation sequencing-based genetic landscape and its clinical implications for Chinese acute myeloid leukemia patients. Cancer Cell Int 18:215. https://doi.org/10.1186/s12935-018-0716-7
Nagel G, Weber D, Fromm E, Erhardt S, Lubbert M, Fiedler W et al (2017) Epidemiological, genetic, and clinical characterization by age of newly diagnosed acute myeloid leukemia based on an academic population-based registry study (AMLSG BiO). Ann Hematol 96(12):1993–2003. https://doi.org/10.1007/s00277-017-3150-3
Zhang Y, Wang F, Chen X, Liu W, Fang J, Wang M, Teng W, Cao P, Liu H (2019) Mutation profiling of 16 candidate genes in de novo acute myeloid leukemia patients. Front Med 13(2):229–237. https://doi.org/10.1007/s11684-018-0616-1
Gaidzik VI, Schlenk RF, Paschka P, Stölzle A, Späth D, Kuendgen A, von Lilienfeld-Toal M, Brugger W, Derigs HG, Kremers S, Greil R, Raghavachar A, Ringhoffer M, Salih HR, Wattad M, Kirchen HG, Runde V, Heil G, Petzer AL, Girschikofsky M, Heuser M, Kayser S, Goehring G, Teleanu MV, Schlegelberger B, Ganser A, Krauter J, Bullinger L, Döhner H, Döhner K (2013) Clinical impact of DNMT3A mutations in younger adult patients with acute myeloid leukemia: results of the AML Study Group (AMLSG). Blood 121(23):4769–4777. https://doi.org/10.1182/blood-2012-10-461624
Taskesen E, Havermans M, van Lom K, Sanders MA, van Norden Y, Bindels E, Hoogenboezem R, Reinders MJT, Figueroa ME, Valk PJM, Löwenberg B, Melnick A, Delwel R (2014) Two splice-factor mutant leukemia subgroups uncovered at the boundaries of MDS and AML using combined gene expression and DNA-methylation profiling. Blood 123(21):3327–3335. https://doi.org/10.1182/blood-2013-07-512855
McGraw KL, Nguyen J, Komrokji RS, Sallman D, Al Ali NH, Padron E et al (2016) Immunohistochemical pattern of p53 is a measure of TP53 mutation burden and adverse clinical outcome in myelodysplastic syndromes and secondary acute myeloid leukemia. Haematologica 101(8):e320–e323. https://doi.org/10.3324/haematol.2016.143214
Owattanapanich W, Utchariyaprasit E, Tantiworawit A, Rattarittamrong E, Niparuck P, Puavilai T, Julamanee J, Saelue P, Chanswangphuwana C, Polprasert C, Limvorapitak W, Kanitsap N, Wanitpongpun C, Nakhakes C, Sriswasdi C, Prayongratana K (2018) Improved survival of elderly-fit patients with acute myeloid leukemia requiring intensive therapy: 3-year multicenter analysis from TALWG. Clin Lymphoma Myeloma Leuk 18(12):e509–e514. https://doi.org/10.1016/j.clml.2018.08.002
Eisfeld AK, Kohlschmidt J, Mrozek K, Blachly JS, Walker CJ, Nicolet D et al (2018) Mutation patterns identify adult patients with de novo acute myeloid leukemia aged 60 years or older who respond favorably to standard chemotherapy: an analysis of Alliance studies. Leukemia 32(6):1338–1348. https://doi.org/10.1038/s41375-018-0068-2
Rücker FG, Schlenk RF, Bullinger L, Kayser S, Teleanu V, Kett H, Habdank M, Kugler CM, Holzmann K, Gaidzik VI, Paschka P, Held G, von Lilienfeld-Toal M, Lübbert M, Fröhling S, Zenz T, Krauter J, Schlegelberger B, Ganser A, Lichter P, Döhner K, Döhner H (2012) TP53 alterations in acute myeloid leukemia with complex karyotype correlate with specific copy number alterations, monosomal karyotype, and dismal outcome. Blood 119(9):2114–2121. https://doi.org/10.1182/blood-2011-08-375758
DiNardo CD, Jonas BA, Pullarkat V, Thirman MJ, Garcia JS, Wei AH et al (2020) Azacitidine and venetoclax in previously untreated acute myeloid leukemia. N Engl J Med 383(7):617–629. https://doi.org/10.1056/NEJMoa2012971
Sallman DA, DeZern AE, Garcia-Manero G, Steensma DP, Roboz GJ, Sekeres MA et al (2021) Eprenetapopt (APR-246) and azacitidine in TP53-mutant myelodysplastic syndromes. J Clin Oncol:JCO2002341. https://doi.org/10.1200/JCO.20.02341
Sallman DA, Asch AS, Al Malki MM, Lee DJ, Donnellan WB, Marcucci G et al (2019) The first-in-class anti-CD47 antibody magrolimab (5F9) in combination with azacitidine is effective in MDS and AML patients: ongoing phase 1b results. Blood 134(Supplement_1):569. https://doi.org/10.1182/blood-2019-126271
Acknowledgements
The authors are grateful to the Department of Internal Medicine III, University Hospital of Ulm, Germany, and Faculty of Medicine Siriraj Hospital, Mahidol University, Thailand, for grant support. We thank Ms. Pattaraporn Tunsing and Ms. Khemajira Karaketklang for their assistance with the data recording and the statistical analyses. We thank Mr. Wannachai Srisaard and Ms. Yaowalak U-Pratya for assistance with the mononuclear cell and gDNA preparation, and Ms. Silvia Schuhhardt and Ms. Adriane Deisboeck for their assistance with the NGS library preparation.
Funding
Open Access funding enabled and organized by Projekt DEAL. The study was funded by the Department of Internal Medicine III, University Hospital of Ulm, Ulm, Germany and Faculty of Medicine Siriraj Hospital, Mahidol University, Thailand.
Author information
Authors and Affiliations
Contributions
All authors designed the study. WO and TR collected the data. WO performed the NGS library preparation, analyzed the NGS results, conducted the statistical analyses, and drafted and revised the manuscript. KD, JH, NJ, and EP supervised the NGS library preparation and analysis of the NGS results. HD, KD, JH, and SI supervised the project and made critical revisions to the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was approved by the Ethics Committee for Research in Human Subjects at the Siriraj Institutional Review Board. All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (Siriraj Institutional Review Board, Faculty of Medicine Siriraj Hospital, Mahidol University, Thailand) and with the Helsinki Declaration of 1975, as revised in 2008.
Consent to participate
Informed consent was obtained from all patients for being included in the study.
Consent for publication
This manuscript has been approved by all authors. A copy of the consent document is available for review from the Editor-in-Chief of Annals of Hematology.
Conflicts of interest
The authors declare no competing interests..
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 21.5 kb).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Owattanapanich, W., Herzig, J., Jahn, N. et al. Genetic alterations in Thai adult patients with acute myeloid leukemia and myelodysplastic syndrome—excess blasts detected by next-generation sequencing technique. Ann Hematol 100, 1983–1993 (2021). https://doi.org/10.1007/s00277-021-04513-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00277-021-04513-z