Abstract
Glucosylglycerol (GG) is a natural compatible solute that can be synthesized by many cyanobacteria and a few heterotrophic bacteria under high salinity conditions. In cyanobacteria, GG is synthesized by GG-phosphate synthase and GG-phosphate phosphatase, and a hydrolase GGHA catalyzes its degradation. In heterotrophic bacteria (such as some Marinobacter species), a fused form of GG-phosphate phosphatase and GG-phosphate synthase is present, but the cyanobacteria-like degradation pathway is not available. Instead, a phosphorylase GGP, of which the coding gene is located adjacent to the gene that encodes the GG-synthesizing enzyme, is supposed to perform the GG degradation function. In the present study, a GGP homolog from the salt-tolerant M. salinexigens ZYF650T was characterized. The recombinant GGP catalyzed GG decomposition via a two-step process of phosphorolysis and hydrolysis in vitro and exhibited high substrate specificity toward GG. The activity of GGP was enhanced by inorganic salts at low concentrations but significantly inhibited by increasing salt concentrations. While the investigation on the physiological role of GGP in M. salinexigens ZYF650T was limited due to the failed induction of GG production, the heterologous expression of ggp in the living cells of the GG-producing cyanobacterium Synechocystis sp. PCC 6803 significantly reduced the salt-induced GG accumulation. Together, these data suggested that GGP may represent a novel pathway of microbial GG catabolism.
Key points
• GGP catalyzes GG degradation by a process of phosphorolysis and hydrolysis
• GGP-catalyzed GG degradation is different from GGHA-based GG degradation
• GGP represents a potential novel pathway of microbial GG catabolism
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Exposure of microorganisms to high salinity conditions usually triggers a fast loss of intracellular water to extracellular environments, which in turn results in a decrease of cell turgor and affects the viability of cells. To counteract this influence, many microorganisms accumulate large amounts of compatible solutes, a functional group of organic compounds generally with low molecular weights, neutral net charge, and good biocompatibility (Brown 1976), in the cell by de novo synthesis and/or uptake from environments (Kempf and Bremer 1998). This strategy ensures basic survival and further growth of microbial cells upon salt stress. When the environmental salinity shifts to standard conditions, the levels of intracellular compatible solutes decline via catabolism or exudation to achieve a new balance of osmotic potential.
Among the compatible solutes (e.g., trehalose, sucrose, proline, 2-O-α-glucosyglycerol [GG], ectoine, and glycine betaine [GB]) identified in microorganisms (Dandapath et al. 2017; Kempf and Bremer 1998; Klähn and Hagemann 2011), GG attracts increasing interest from industry and academia in recent years because of its desirable properties/activities (such as clear-cut sweetness, noncariogenicity, excellent water-holding capacity, protective effects on macromolecules, and antitumor activity) and potential applications in the agronomy, health care, cosmetics, and pharmacy fields (Luo et al. 2022). Nowadays, GG is commercially produced by at least two methods (i.e., enzymatic synthesis and native extraction from cyanobacteria) and has been widely used as an excellent moisturizer in many cosmetic and personal care products (Luo et al. 2022). Concerning the basic research, the genetic/biochemical background and regulatory mechanisms of microbial (especially cyanobacterial) GG anabolism have been elucidated (Hagemann and Erdmann 1994; Mikkat et al. 1996; Novak et al. 2011). GG is synthesized by a two-step process. GG-phosphate synthase (GGPS) first catalyzes the conversion of ADP-glucose and glycerol-3-phosphate to GG-3-phosphate and ADP. A phosphatase, i.e., GG-phosphate phosphatase (GGPP), then hydrolyzes GG-3-phosphate to generate GG. The two enzymes are encoded by two separate genes ggpS and ggpP in nearly all GG-producing cyanobacteria (except Nostoc ellipsosporum NOK) (Fig. 1) (Hagemann et al. 1996, 1997; Marin et al. 1998). In some heterotrophic bacteria such as Stenotrophomonas rhizophila DSM14405T, Pseudomonas mendocina NK-01, Azotobacter vinelandii AEIV, and Marinobacter adhaerens HP15, the GGPS and GGPP candidates are also present, but they are encoded as a fused protein by a single gene (termed ggpPS) with GGPP as its N-terminal domain and GGPS as its C-terminal domain (Hagemann et al. 2008; Klähn and Hagemann 2011). This suggests possible differences of these bacteria from cyanobacteria in GG metabolism.
Illustration of the presence of the key genes participating in microbial GG metabolism in the genomes of the identified or potential GG-producing cyanobacteria and heterotrophic bacteria. ggpS and ggpP compose the GG-synthesizing pathway. gghA represents the GG-degrading pathway. ggp is the putative novel GG-degrading pathway investigated in the present study. The filled and open squares indicate the presence and absence of the target genes, respectively. The number “2” in the squares demonstrates that two target genes are present. The co-location of genes in the genomes is demonstrated by the dotted lines. The locus tags of ggpP/ggpS/gghA are slr0746/sll1566/slr1670 for Syn6803, NIES39_R01210/M02120/M02110 for A. platensis NIES-39, AmaxDRAFT_4256/2350/2351 for L. maxima CS-328, SYNPCC7002_A2841/A2851/A2843 + A2849 for Synechococcus sp. PCC 7002, Lepto7376_0796/0792/0793 + 0794 for Leptolyngbya sp. PCC 7376, and Cce_3291/2391/3236 for C. subtropica ATCC 51142, respectively. The locus tags of ggpP and ggpS in Synechococcus sp. WH8102 are SYNW0860 and SYNW1281, respectively. The locus tags of ggpPS are JMG10_34610 for N. ellipsosporum NOK, DX03_04600 for S. rhizophila DSM 14405T, MDS_3416 for P. mendocina NK-01, AVAEIV_003468 for A. vinelandii AEIV, and HP15_2852 for M. adhaerens HP15, respectively. The locus tag of ggp in M. adhaerens HP15 is HP15_2853
To finely regulate intracellular contents and avoid a net loss of carbon and energy, microbial cells are generally thought to recycle compatible solutes when the stress environment shifts to normal. The molecular basis of microbial GG catabolism was recently unraveled in the model cyanobacterium Synechocystis sp. PCC 6803 (hereafter Syn6803) (Kirsch et al. 2017; Savakis et al. 2016). A glycoside hydrolase GGHA, encoded by slr1670, catalyzed the hydrolysis of GG, generating glycerol and glucose. As predicted, putative gghA genes are widely found in the GG-producing cyanobacteria (Fig. 1). In a few species such as Synechococcus sp. PCC 7002 (hereafter Syn7002) and Leptolyngbya sp. PCC 7376, even two gghA homologs are available. Intriguingly, in most cases, the gghA gene is co-located with the ggpS gene, which supports its functional correlation. This organization pattern may facilitate efficient synergistic regulation of GG anabolism and catabolism. However, the homologs of gghA are not found in the heterotrophic bacteria possessing the GGPPS pathway (Fig. 1), remaining the mechanism of GG catabolism mysterious in these microorganisms.
Recently, we noticed that the genome of the halotolerant bacterium M. adhaerens HP15 harbors a ggpPS gene (HP15_2852) that should enable GG synthesis via a fusion enzyme showing synthase and phosphatase activity as well as a putative hydrolase gene (HP15_2853) that is located downstream of ggpPS (Fig. 1) (Luo et al. 2022). Considering their predicted functions and genomic organization, we supposed that the hydrolase is involved in GG catabolism. Because of the low sequence identity (13%) to GGHA, the HP15_2853 protein may have a different behavior from GGHA. The results of Franceus and colleagues indicated that HP15_2853 codes for a glycoside hydrolase (termed GGP) belonging to the subfamily 18 of glycoside hydrolase family 13 (GH13_18) (Franceus et al. 2018). In the present study, the function of GGP was further determined with a homolog from M. salinexigens ZYF650T by both biochemical characterization and physiological analyses. We found that this GGP catalyzed GG decomposition via a two-step process of phosphorolysis and hydrolysis in vitro. The ggp genes were not only found in M. salinexigens and M. adhaerens, but also present in many members of the Marinobacter genus. It may represent a novel pathway of microbial GG catabolism.
Materials and methods
Strains and cultivation conditions
M. salinexigens strain ZYF650T (MCCC 1K03552T) was provided by Marine Culture Collection of China. Marinobacter cells were grown in 2216E medium or modified marine broth (MB: 5 g/l peptone, 1 g/l yeast extract, 0.04 g/l ferric citrate) at 30 ℃, and different concentrations of NaCl were supplemented as indicated in the text. Syn6803 was from the strain collection of our lab. For standard cultivation, Syn6803 cells were grown in BG11 medium aerated with sterile air at 30 °C under constant white-light illumination of 120 μmol/m2/s. To induce GG production, cyanobacterial cells cultivated under the standard condition were transferred into BG11 medium containing 4% NaCl. Escherichia coli cells were grown in lysogeny broth (LB) at 37 °C. Cell growth was monitored by measuring the optical density at 600 nm (OD600, for Marinobacter and E. coli) or 730 nm (OD730, for Syn6803). Antibiotics such as chloramphenicol (Cm, 5 [in MB] or 20 μg/ml [in 2216E] for Marinobacter, 10 μg/ml for Syn6803) and kanamycin (50 μg/ml for E. coli) were applied when required. For solid medium, 1.6% (w/v) agar was supplemented. All quantitative data were presented as means from three independent replicates.
Sequence analyses
The protein datasets of 250 Marinobacter isolates (supplementary information Table S1) were obtained from the “Genome” database of NCBI (https://www.ncbi.nlm.nih.gov/) using the integrated program Batch Entrez. Blastp analyses were conducted in a local manner using BLAST + (version ncbi-blast-2.13.0 +), and the threshold of outputs was set at the sequence identity of 30%. The software Origin was used to generate heatmaps. The phylogenetic relation of GGP proteins was analyzed with the software MEGA using the neighbor-joining method.
Analyses of intracellular compatible solutes
The extraction and determination of intracellular compatible solutes (mainly GG) of Syn6803 were performed as previously described (Qiao et al. 2019). The same procedure was applied to Marinobacter.
Protein expression and purification
The coding sequence (FWJ25_14990) of M. salinexigens ZYF650T GGP was synthesized by BGI (Beijing, China) after codon optimization (supplementary information), and cloned into pET-28b vector with an N-terminal His-tag. To overexpress ggp, E. coli BL21(DE3) cells harboring the expression plasmid were induced by 0.2 mM isopropyl-β-D-thiogalactopyranoside for 20 h at 20 ℃. Centrifugation was conducted at 8228 × g for 20 min to harvest cells. Cell pallets were resuspended and disrupted by sonication in prechilled binding buffer (20 mM KH2PO4-K2HPO4, pH 7.0, 300 mM NaCl). The lysates were centrifuged at 8228 × g for 60 min. After filtration through 0.22-μm polyethersulfone membrane, the supernatants were applied to Ni–NTA affinity columns for protein purification. Non-specific proteins were removed with wash buffer (binding buffer containing 50–60 mM imidazole), and the target protein was obtained at an imidazole concentration of 200 mM. The removal of imidazole was conducted by ultrafiltration of 30-kDa, and the resulting GGP protein was stored in the binding buffer for enzyme assays. The purity of the target protein was examined by SDS-PAGE. The protein content was determined according to the Bradford method (Bradford 1976). For the phosphorolytic and hydrolytic assays, GGP was purified using 20 mM Tris–HCl (pH7.0, containing 300 mM NaCl) as the basic buffer, and the purification procedure was the same as above.
Enzyme assays
All enzyme assays were conducted in 200-μl reaction mixtures. Initially the enzymatic degradation of GG by GGP was evaluated by incubating 20 mM GG and a proper amount of GGP in 20 mM PB buffer (KH2PO4-K2HPO4, pH7.0) at 30 ℃. Unless noted otherwise, further assays were conducted in the mixtures of 20 mM PB or MES buffer (as indicated in the text), 20 mM GG, and 0.05 mg/ml GGP at 45 ℃ for 0.5 h to determine enzyme activity. To determine the hydrolytic activity, 20 mM MES solution (pH7.0) was used as the reaction buffer, and 20 mM α-glucose-1-phosphate (αG1P) was used as the substrate. All reactions were terminated by heating at 90 ℃ for 10 min. The amounts of GG, glucose, and glycerol in reactions were determined using an ICS5000/5000+ ion-exchange chromatography system (ThermoFisher, Waltham, MA, USA) as previously described (Qiao et al. 2018). The content of αG1P was determined using a αG1P assay kit (Grace Biotechnology, Suzhou, China). All quantitative data were presented as means from three independent replicates.
Construction of the ggp-deficient mutant of Marinobacter
All plasmids and primers used in the present study are listed in Table 1. To inactivate ggp in M. salinexigen ZYF650T, the Cm-resistant marker Cmr was amplified from plasmid pQL199 employing primers Msa-Cm-F/Msa-Cm-R, and the flanking regions of ggp were amplified using M. salinexigen ZYF650T genomic DNA and primers MsaGGPko-up-F/MsaGGPko-up-R and MsaGGPko-dn-F/MsaGGPko-dn-R. The three fragments were linked and cloned into the pCE-Zero vector using the ClonExpress® Ultra One-Step-Cloning Kit (Vazyme Biotech, Nanjing, China), yielding plasmid pCL8. M. salinexigens cells, cultivated in 2216E medium at 30 °C for 24 h, were harvested at 4 °C. The cells were washed twice with prechilled sucrose solution (300 mM) and resuspended in the same solution to a final cell density of OD600≈2.0. pCL8 was introduced into M. salinexigens cells by electroporation. Briefly, 100 μl of cells was mixed with 100 ng plasmid DNA and electro-pulse treated for 5 ms using a micropulser (Bio-Rad, CA, USA) with the following settings: 12.5 kV/cm, 25 μF, and 200 Ω. Immediately after pulse, the cells were supplemented with 900 μl of 2216E medium and regenerated at 30 °C for 20 h by shaking. The cells were plated on 2216E agar plates containing 20 μg/ml Cm. Resistant transformants were obtained after 24 h of selection. The genotypes of the transformants were verified by PCR.
Construction of recombinant Syn6803 strains
To overexpress ggp in the neutral site slr0168 of Syn6803, an expression platform, which harbors Cmr, the strong cyanobacterial promoter Pcpc560 (Zhou et al. 2014) and the flanking regions of slr0168, was amplified from plasmid pJS40 employing primers Trbcl-ggp-F/Pcpc560-ggp-R. The open reading frame (ORF) of M. salinexigens ZYF650T ggp was amplified with primers ggp-Pcpc560-F/ggp-Trbcl-R. The two fragments were linked by One-Step-Cloning (see above) to generate plasmid pCL12. To overexpress ggp at a gghA-deficient background of Syn6803, a Cmr-Pcpc560 fragment was amplified from pJS40 with primers Cm-gghA-F/Pcpc560-ggp-R. The ggp gene was amplified as above using primers ggp-Pcpc560-F/ggp-gghA-R, and the flanking regions of gghA were amplified from Syn6803 using gghA-up-F/gghA-Cm-up-R and gghA-ggp-F/gghA-dn-R. The four fragments were linked and cloned into pCE-Zero, yielding plasmid pCL10. In both pCL10 and pCL12, the ggp gene was under the control of Pcpc560. To construct a gghA-deficient control, the Cmr fragment (amplified by Cm-gghA-F/Cm-gghA-R) and the gghA flanking regions (amplified by gghA-up-F/gghA-Cm-up-R and gghA-Cm-dn-F/gghA-dn-R) were combined and cloned into pCE-Zero, yielding plasmid pCL11. For the inducible expression of ggp, the promoter Pcpc560 of pCL10 was replaced by a theophyllin-regulated riboswitch ENYC4 (Qiao et al. 2018). The ENYC4 and vector fragments were amplified by Cm-ENYC4-F/ENYC4-ggp-R and gghA-ENYC4-F/gghA-ENYC4-R, respectively. After one-step-cloning, pCL14 was generated.
The final constructs, pCL10, pCL11, pCL12, and pCL14 were transferred to the wild-type strain of Syn6803 by natural transformation. The Cmr transformants were obtained after at least 7 days of selection on BG11 agar plates. The genotypes of the transformants were verified by PCR. Recombinant strains QL513, QL514, QL515, and QL518 were obtained.
Results
Potential GG metabolism in Marinobacter by sequence analyses
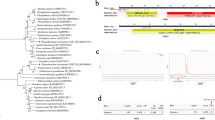
To determine the occurrence of potential GG metabolism in the genus Marinobacter, local blastp analyses of GGP (HP15_2853) and GGPPS (HP15_2852) of M. adhaerens HP15 were performed against 250 annotated Marinobacter genomes (Table S1). The GGPPS homologs exhibiting ≥ 30% sequence identity were identified in 53 isolates, among which the GGP homologs were detected in 49 cases (Fig. 2). Despite annotated as sucrose phosphorylases that belong to the GH13_18 family, these GGP homologs were classified into a different clade from the well-characterized sucrose phosphorylase of Leuconostoc mesenteroides (Goedl et al. 2007), in the phylogenic analysis (Fig. 2). Excluding five isolates (Marinobacter sp. CBIW17, M. gelidimuriae BF04_CF-4, M. adhaerens UBA9380, M. adhaerens UBA8974, and M. adhaerens SW_4_16), in which the putative ggpPS or ggp genes occur at the ends of genomic contigs, the organization of the two genes in the genome is similar to that in M. adhaerens HP15, i.e., the ggpPS gene is located immediately upstream (17–76 bp) of ggp in the same direction. The search for the reported GG-degrading pathway, GGHA, was also performed. No homologs showing ≥ 30% sequence identity were identified, indicating that Marinobacter may have a different GG catabolic pathway from that of cyanobacteria.
A heatmap of the putative proteins involved in microbial GG (pink area), ectoine (pale green area), GB (pale purple area), GGA (gray area), and sucrose (pale blue area) metabolisms in 49 Marinobacter isolates. The query sequences used for the local blastp analyses are given on the top. GGP (HP15_2853) and GGPPS (HP15_2852) are from M. adhaerens HP15. GGHA (Slr1670), SPS (Sll0045), SPP (Slr0953), and invertase (Sll0626) are from Synechocystis sp. PCC 6803. EctB (HALO_2589), EctA (HALO_2588), EctC (HALO_2590), DoeA (HALO_3665), DoeB (HALO_3664), and DoeD (HALO_3661) are from H. elongate DSM 2581T. BetA (CAA37093.1) and BetB (CAA37092.1) are from E. coli. GSMT (AAF87202.1) and SDMT (AAF87203.1) are from Ectothiorhodospira halochloris. GPGS (SYNPCC7002_A2021), GPGP (SYNPCC7002_A2023), and GGAP (SYNPCC7002_A2022) are from Synechococcus sp. PCC 7002. The frames and connecting lines on GGPs and GGPPSs indicate their adjacent gene organization. The phylogenic relation of 44 GGPs, which have complete ORFs, is presented on the left and L. mesenteroides sucrose phosphorylase (BAA14344.1) was used as a reference
The metabolic pathways of ectoine, GB, glucosylglycerate (GGA), and sucrose, the other four well-identified compatible solutes in bacteria (Costa et al. 2007; da Costa et al. 1998; Kirsch et al. 2019; Klähn et al. 2010), were additionally surveyed in the above 250 genomes (Table S1 and Fig. 2). The homologs (≥ 30% identity) of diaminobutyrate-2-oxoglutarate transaminase (EctB), diaminobutyrate acetyltransferase (EctA), and ectoine synthase (EctC), which compose the ectoine-synthesizing pathway of Halomonas elongata (Cánovas et al. 1998), were simultaneously identified in 228 Marinobacter isolates. No homologs of ectoine hydrolase (DoeA), the key enzyme involved in the well-identified ectoine-degrading pathway DoeABD (Schwibbert et al. 2011), were found in these isolates. In the case of GB synthesis, the E. coli-like BetAB pathway, catalyzing GB synthesis via choline oxidation by choline dehydrogenase (BetA) and betaine dehydrogenase (BetB) (Lamark et al. 1991), were detected in most of the Marinobacter isolates (i.e., 229 out of 250), whereas the de novo pathway consisting of glycine-sarcosine methyltransferase (GSMT) and sarcosine-dimethylglycine methyltransferase (SDMT) seemed not present in Marinobacter (Nyyssola et al. 2000). Regarding GGA, the homologs (≥ 30% sequence identity) of glucosyl-3-phosphoglycerate synthase (GPGS) and glucosyl-3-phosphoglycerate phosphatase (GPGP), which are responsible for GGA synthesis (Klähn et al. 2010), and the possible GGA-degrading enzyme GGAP (GGA phosphorylase) (Franceus et al. 2017), were simultaneously identified in 198 isolates. These results suggested that ectoine, GB, and GGA may be also employed as compatible solutes by Marinobacter for salt acclimation. On the other hand, only a few homologs of the sucrose-synthesizing enzymes, SPS and/or SPP, were found, and no homologs of the sucrose-degrading enzyme invertase were identified. Thus, the genus Marinobacter seems not to have the complete pathway of sucrose metabolism.
Preliminary evidence of GG degradation by GGP
To explore the GG-degrading activity, the putative ggp gene (FWJ25_14990) of M. salinexigens ZYF650T, a deep sea isolate from the Mariana Trench (Ahmad et al. 2020), was heterologously expressed in E. coli with an N-terminal His-tag. After Ni affinity chromatography, GGP was purified to apparent homogeneity. The SDS-PAGE as well as native PAGE analyses showed a single protein band with an estimated molecular mass of 57 kDa (Fig. 3a and Fig. S1), in agreement with its theoretical value (57.0 kD) deduced from the amino acid sequence.
Analyses of the GG-degrading activity and other basic properties of GGP. a Examination of GGP purification by SDS-PAGE. M, protein marker; S, the supernatant fraction of cell extracts; E1–E5, different elution fractions at 200 mM imidazole. The solid triangle indicates the position of GGP in the gel. b The qualitative analysis of GG decomposition catalyzed by purified GGP. The assays were performed in PB buffer at 30 °C for 12 h with GG as the substrate. Gly, glycerol; glu, glucose. c Evaluation of the residual GG content in the presence ( +) or absence ( −) of GGP after different reaction times. The measured amounts were normalized by comparing them with the starting values (at 0 h). d, e The quantitative analysis of GG decomposition catalyzed by GGP (0.05 mg/ml). f, g The effects of reaction temperature and pH on the activity of GGP
The degrading activity of GGP was examined using GG as the substrate (Fig. 3b). After 12-h incubation, two products, glycerol and glucose, were detected. The amounts of the products exhibited an elevated pattern with the increase of GGP amount, whereas the GG level gradually decreased. To exclude possible spontaneous decomposition, the stability of GG was analyzed in a time-course manner (Fig. 3c). In the absence of GGP, GG stayed on a relatively stable level within 24 h. However, the GG level declined rapidly within the first 4 h in the presence of GGP and was 77.8% of the initial level at 24 h. Further quantitative analyses showed that the amount of GG degradation matched well to that of glycerol generation but was clearly more than that of glucose generation (Fig. 3d and e), suggesting a two-step process of GG decomposition by GGP. Considering that GGP is predicted to be a phosphorylase of GH13_18 family, we supposed that it first phosphorolyzes GG to glycerol and αG1P and the latter compound is further hydrolyzed to produce glucose (Fig. 4a). Before we further verified this hypothesis, basic features of the enzyme were determined. GGP exhibited relatively high activity at temperatures of 30–50 °C and the highest activity was detected at 45 °C. At temperatures of above 50 °C, the activity rapidly declined (Fig. 3f). The optimal pH was pH6.6 (Fig. 3g). The thermostability of the enzyme was also determined by measuring the residual activity after heat treatment. The semi-inactivation temperature (T5010) was 53 °C and the half-life time at this temperature (t1/2) was around 8 h (Fig. S2).
The proposed process of GG degradation by GGP (a) and biochemical verification of the phosphorolytic and hydrolytic steps during GG degradation (b–e). In b and c, the phosphorolytic assays were performed in 20 mM MES buffer (pH 6.6) with GG as the substrate. In d and e, the hydrolytic assays were performed in 20 mM MES buffer (pH 7.0) with αG1P as the substrate. A proper volume of PB buffer (pH6.6 or pH7.0) was supplemented to achieve the indicated Pi concentrations. Gly, glycerol; αG1P, α-glucose-1-phosphate; Glu, glucose
A two-step degradation of GG by GGP in vitro
To identify GG phosphorolysis, the PB buffer used in the above experiments was changed to MES buffer and a Pi gradient was supplemented (Fig. 4b). Elevated Pi concentrations caused improved generation of glycerol and αG1P (Fig. 4b and c), indicating a proportional increase of GG degradation with the elevated Pi level. To verify the subsequent hydrolytic reaction, αG1P was used as the substrate and the glucose product was determined. As predicted, the amount of glucose was elevated with the increase of GGP content (Fig. 4d) and reduced with the increase of Pi concentration (Fig. 4e). Taken together, these results confirmed that GG degradation catalyzed by GGP in vitro is a successive process of phosphorolysis and hydrolysis.
Salt-mediated regulation of GGP activity and its substrate specificity
GG is known to function as a compatible solute for microbial salt acclimation. Its metabolism can be regulated on transcriptional, translational, and post-translational levels in cyanobacteria, and the salt-mediated modulation of the activities of the key enzymes (e.g., GGPS and GGHA) plays an important role for the fast response during acclimation (Kirsch et al. 2017; Marin et al. 2002; Novak et al. 2011). We supposed that GGP activity might be affected by salt concentrations. To this end, the impact of increasing inorganic salt concentrations on GGP activity was determined. Under the salt-free condition, GGP exhibited a noticeable activity of 1.17 μmol/min/mg (Fig. 5a). Although slightly activated at low concentrations (0–0.2 M), the enzyme activity was inhibited by the addition of NaCl (0.2–2.0 M). The activity declined to 22% (0.36 μmol/min/mg) of the highest level (1.63 μmol/min/mg, at 0.2 M) at 0.8 M NaCl and to almost non-detectable level at NaCl concentrations of 1.2 M or higher. The decreased activity seemed not due to the disruption of the protein structure. An impact of high salinity (up to 2 M NaCl) on protein status was not observed in the native PAGE analysis (Fig. S1). To explore whether such regulation exhibits salt preference, the impacts of additional three inorganic salts (i.e., KCl, NaNO3, and NH4Cl) were investigated in parallel with NaCl (Fig. 5b). Similarly, a strong inhibition, starting with a gentle activation at low salt concentrations (0–0.2 M for KCl, 0–0.05 M for NaNO3 and NH4Cl), by increasing salt concentrations was observed for all these salts. Thus, GGP exhibits a salt-dependent regulation of enzyme activity, similar to that of the previously identified GG- and sucrose-catabolizing enzymes, GGHA and invertase (Kirsch et al. 2018, 2017).
Effects of inorganic salts on the GG-degrading activity of GGP (a, b) and examination of the substrate specificity of GGP (c). In a and b, the assays were performed in 20 mM PB buffer (pH6.6) for 0.5 h with GG as the substrate. Different concentrations of NaCl, KCl, NaNO3, and NH4Cl were included in the reaction mixtures as indicated. The reaction without salt addition was used as the control. In c, the assays were performed for 12 h in the presence ( +) or absence ( −) of GGP with GG, sucrose, cellobiose, trehalose, melibiose, maltose, lactose, and kojibiose as the substrates, respectively. The obtained values were normalized by comparing them with the starting values (at 0 h). The asterisk indicates significant difference between the tested samples (Student’s t test, P < 0.01)
The substrate specificity of GGP was analyzed with seven disaccharides (sucrose, cellobiose, trehalose, melibiose, maltose, lactose, and kojibiose) in parallel with GG (Fig. 5c). After 12-h reaction, the residual amount of GG in the presence of GGP was apparently lower than that in the absence of the enzyme. Under the same condition, the residual amounts of other disaccharides were similar to the controls. These observations indicated a strict specificity of GGP toward GG.
Effects of ggp inactivation in M. salinexigens
Before the genetic manipulations in M. salinexigens ZYF650T, the salt tolerance and antibiotic sensitivity of the strain were determined. M. salinexigens ZYF650T showed growth in the medium containing 3–12% NaCl (Fig. S3a), similar to that reported by Ahamad et al. (Ahmad et al. 2020). Under the NaCl-free and 15% NaCl conditions, no growth was observed. Whereas M. salinexigens ZYF650T showed apparent resistances to kanamycin, apramycin, and neomycin (minimum inhibitory concentration > 100 μg/ml), high sensitivities to ampicillin and chloramphenicol were seen (Fig. S3b). To explore the GG-catabolizing activity of GGP in vivo, the ggp gene was inactivated (Δggp) by inserting a Cmr cassette and the possible GG production was tracked (Fig. 6a). Marinobacter cells were cultivated in MB medium containing 3% NaCl and then treated by elevated NaCl concentrations (3–12%). Contrary to our expectation, GG was not detected in the wild-type cells at all NaCl concentrations during the whole testing period (0–24 h). No detectable GG production was observed in the Δggp cells as well under the same condition. Similar experiments were also performed in minimal salt medium with glycerol as the sole carbon and energy source. As seen above, no GG was accumulated by the wild-type and Δggp cells under elevated NaCl concentrations (Fig. S4a). The capability of M. salinexigens ZYF650T to assimilate exogenous GG was also analyzed (Fig. 6b). Cells were grown in minimal salt medium with GG as the sole carbon and energy source. Glycerol was used as the positive control. At GG concentrations of 20 and 100 mM, the wild-type strain did not grow as the negative control (without carbon and energy source). A weak growth was observed at a relatively high GG concentration of 200 mM within the first 2 days. Under the same condition, the Δggp mutant did not grow at all.
Analyses of the possible GG accumulation of the M. salinexigens ZYF650T wild-type (WT) and Δggp strains under different NaCl concentrations (a) and the growth of the M. salinexigens ZYF650T wild-type and Δggp strains with GG as the sole carbon and energy source (b). In a, the cells grown in MB medium containing 3% NaCl were transferred into the same medium with 3–12% NaCl. In b, cells were grown in minimal salt medium (MSM: 5.3 g/l MgCl2-6H2O, 0.75 g/l KCl, 0.1 g/l MgSO4-7H2O, 50 mg/l K2HPO4, 1 g/l NH4Cl, 0.74 g/l CaCl2-2H2O, 0.42 g/l NaHCO3, and 20 g/l NaCl) supplemented with 20 mM glycerol or 20–200 mM GG
Effects of ggp expression in Syn6803
The physiological function of ggp was further investigated in the GG-producing cyanobacterium Syn6803. The mutant strain QL513, expressing the ggp gene of M. salinexigens ZYF650T under the control of a strong cyanobacterial promoter Pcpc560 at the neutral site slr0168, was constructed (Fig. 7a). Syn6803 cells were exposed to 4% NaCl for GG production (Fig. 7b). As predicted, the wild-type Syn6803 exhibited a fast accumulation of GG within 24 h upon salt stress. Under the same condition, GG production of QL513 was greatly depressed. The GG content of the mutant was only 1.6 mg/l/OD730 in 24 h, which was much lower than the wild-type level (24.7 mg/l/OD730). Similar genetic construction was performed in the gghA-deficient background (QL514), generating QL515. Due to the inactivation of the native GG-catabolizing pathway, QL514 showed an enhanced GG accumulation compared with the wild-type. Like that in QL513, the production of GG was dramatically depressed by the expression of ggp.
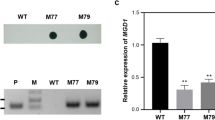
Analyses of the salt-induced GG accumulation of the wild-type (WT) and mutant strains of Syn6803. In a, the genotypes of the WT and mutant strains are illustrated. “up” and “dn” represent the upstream and downstream flanking regions of target genes, respectively. In b, the WT and mutant (QL513, QL514, QL515, and QL518) cells grown in the standard BG11 medium were stressed by 4% NaCl to induce GG production. In c, the cells of QL514 and QL518 were cultivated in BG11 medium containing 4% NaCl for 48 h and then supplemented with different concentrations of theophylline to induce ggp expression. T0, T500, T750, and T1000 represent 0, 500, 750, and 1000 μM theophylline, respectively. The asterisks indicate significant differences from the starting levels (0 h) (Student’s t test, P < 0.01)
In additional to the perspective of GG accumulation, GGP function was also analyzed from the perspective of GG degradation. Mutant QL518, in which the ggp gene was placed under the control of the theophylline-dependent riboswitch ENYC4 (Nakahira et al. 2013), was constructed (Fig. 7a). Cells were first cultivated under the salt stress condition (4% NaCl) for 48 h to accumulate GG. Then theophylline was supplemented to induce ggp expression, and the content of GG was monitored (Fig. 7c). Apparent GG degradation induced by theophylline was observed, in particular at high theophylline concentrations. At 750 and 1000 mM of theophylline, the intracellular GG of QL518 was significantly decreased in 48 h, with a reduction of 27.1% and 39.1%, respectively. With regard to the control QL514, it maintained a stable level of GG throughout the testing period in the presence of 1000 mM theophylline. Together, these data demonstrated a clear function of GGP in the degradation of GG in the living cells of cyanobacteria.
Discussion
Marinobacter species are often isolated from marine and other saline environments. They are generally known for the good tolerance to high salinity (up to 20%) (Nie et al. 2021; Yoo et al. 2020; Zhang et al. 2008, 2020a). In agreement with this property, a few pathways for compatible solute syntheses were detected in many isolates of Marinobacter (Table S1 and Fig. 2). We found that 53 out of 250 annotated Marinobacter genomes contained the putative ggpPS gene (Table S1), indicating the potential of GG synthesis under special conditions (e.g., stresses). More interestingly, a ggp homolog was always present in associated with ggpPS (excluding four cases with partial ggpPS at the termini of genomic contigs) and located adjacent to the latter gene. This phenomenon strongly suggested the involvement of ggp in GG metabolism. To date, only one pathway, namely GGHA, for microbial GG catabolism was physiologically verified (Kirsch et al. 2017; Savakis et al. 2016). This was done in cyanobacteria. The genetic and biochemical investigations of Syn6803 showed that GG is catabolized by GGHA under low salinity conditions via a hydrolytic process, generating glucose and glycerol. In most of the GG-producing cyanobacteria, the putative gghA gene is closely linked with the key GG-synthesizing gene ggpS, echoing their metabolic correlation (Fig. 1). However, in the genomes of Marinobacter, no gghA homologs were detected, indicating the absence of the GGHA pathway. Hence, it becomes logic to suppose a GG-catabolizing role of ggp in Marinobacter.
Here, detailed investigations on the function of GGP of M. salinexigens ZYF650T, an isolate from the hadal seawater with a salt tolerance of 0–14% NaCl (Ahmad et al. 2020), were performed. The purified GGP catalyzed apparent decomposition of GG with the products glycerol and glucose, supposing the hydrolysis of GG. However, our quantitative analyses of the products did not support a pure hydrolysis. The amount of glycerol product was significantly higher than that of glucose (Fig. 3d), raising a question that “where is the other part of glucose.” In 2018, Franceus and colleagues provided the first evidence for the function of GGP (Franceus et al. 2018). The M. adhaerens HP15 GGP (HP15_2853), which had ever been thought to be a sucrose phosphorylase, was identified as a GG phosphorylase, showing a preferred specificity on GG instead of sucrose. The recombinant GGP catalyzed reversible phosphorolysis of GG with αG1P and glycerol as the products or substrates in vitro. In consideration of the high sequence identity between the M. salinexigens ZYF650T and M. adhaerens HP15 GGPs, the decomposition of GG in the present study was thought to be a two-step process composed of phosphorolysis and hydrolysis. In line with this hypothesis, an increased generation of αG1P using GG as the substrate under Pi elevation conditions and a production of glucose using αG1P as the substrate under Pi free conditions were detected (Fig. 4c and e).
Although the GG-decomposing activity of Marinobacter GGPs has been solidly verified by the previous study as well as the present one (Franceus et al. 2018), the physiological roles of these enzymes are still unclear. Physiological investigations are thereby expected. In the cell, the Pi content is expected to be higher than that of αG1P, because the latter compound can be easily captured by glycolysis. From this perspective, GGP is thought to perform a catabolic function via phosphorolysis in vivo. A phenomenon that may be related with this hypothesis is that, a weak increase of cell density was seen for the wild-type M. salinexigens ZYF650T grown in minimal salt medium with GG (0.2 M) as the sole carbon and energy source, while the Δggp mutant did not show any growth at all (Fig. 6b). To give more information for the function of GGP, physiological analyses were further performed both natively (in M. salinexigens ZYF650T) and heterologously (in Syn6803) under salinity shift conditions. No matter in the wild-type Syn6803 or the gghA-deficient mutant (QL514), the heterologous expression of ggp indeed caused a great suppression of the salt-induced GG accumulation (Fig. 7b). A direct depletion of accumulated GG was also observed with the induced expression of GGP (Fig. 7c). Because this analysis was conducted in the gghA-deficient background, the influence of the native catabolizing pathway could be eliminated. These results confirmed the GG-catabolizing activity of GGP in the living cells. On the other hand, the analysis in Marinobacter was limited because of the failed induction of intracellular GG accumulation. No GG was detected in the wild-type cells of M. salinexigens ZYF650T after salt treatment at all tested concentrations (up to 12% NaCl). (i) One explanation can be that GG is not the main compatible solute of this bacterium and stays at a non-detectable level due to the fast turnover. Putative anabolic pathways of ectoine (EctBAC), GB (BetAB), and GGA (GPGS and GPGP), the other three well-identified compatible solutes in many bacteria, were found in 228, 229, and 199 cases of 250 MarinobacterS isolates, respectively (Table S1 and Fig. 2). They were more widely distributed among Marinobacter than the putative GG metabolic pathways (49/250). However, we also noticed that two results did not support the above conjecture: GG was not accumulated in the wild-type M. salinexigens ZYF650T at relatively high NaCl concentrations (such as 9% and 12%, equivalent to 1.5 M and 2.1 M, respectively), at which GGP did not exhibit activity (Fig. 5a); and GG accumulation was also not observed when the ggp gene was inactivated (Fig. 6a). (ii) Another possibility is that the tested conditions are not suitable to induce GG synthesis. Many investigations have reported that bacteria produce compatible solutes (e.g., ectoine, 5-hydroxyectoine, trehalose, GGA, and sucrose) under diverse stress conditions such as high salinity, high or low temperature, nitrogen starvation, and desiccation (Alarico et al. 2014; Bursy et al. 2008; Cánovas et al. 2001; de Alvarenga et al. 2020; Kuhlmann et al. 2008). In addition to salt treatment, in the present study, the effect of relatively high temperatures (e.g., 45 °C) was also evaluated (Fig. S4b). Unfortunately, M. salinexigens ZYF650T did not produce detectable GG at all these temperatures. Further investigations on more stress conditions need to be performed.
GGP is a GG-specific phosphorylase that belongs to the GH13_18 family in the carbohydrate-active enzyme database (CAZy, http://cazy.org). It is able to catalyze reversible phosphorolysis of GG under proper conditions. In terms of biotechnology, this provides the possibility to synthesize GG by the utilization of the reverse phosphorolytic reaction. Several efforts to produce GG from αG1P and glycerol using M. adhaerens HP15 GGP have been conducted (Sun et al. 2023; Zhang et al. 2020b). To reduce production cost, αG1P was provided from low-cost sucrose (or maltodextrin) instead of direct addition by a coupling use of sucrose phosphorylase (or α-glucan phosphorylase). Under the optimized condition, a production of 61 g/l GG was achieved after 24 h using maltodextrin and glycerol as substrates. The results of Sun et al. also demonstrated that the GGPs of different bacterial sources presented distinct activities to catalyze GG synthesis. Hence, the further identification of GGPs will provide more options of enzyme catalysts in GG biosynthesis.
Data availability
Data will be made available on request.
References
Ahmad W, Zheng Y, Li Y, Sun W, Hu Y, He X, Liu R, Xue CX, Zhang XH (2020) Marinobacter salinexigens sp. nov., a marine bacterium isolated from hadal seawater of the Mariana Trench. Int J Syst Evol Microbiol 70:3794–3800. https://doi.org/10.1099/ijsem.0.004236
Alarico S, Costa M, Sousa MS, Maranha A, Lourenco EC, Faria TQ, Ventura MR, Empadinhas N (2014) Mycobacterium hassiacum recovers from nitrogen starvation with up-regulation of a novel glucosylglycerate hydrolase and depletion of the accumulated glucosylglycerate. Sci Rep 4:6766. https://doi.org/10.1038/srep06766
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1006/abio.1976.9999
Brown AD (1976) Microbial water stress. Bacteriol Rev 40:803–846. https://doi.org/10.1128/br.40.4.803-846.1976
Bursy J, Kuhlmann AU, Pittelkow M, Hartmann H, Jebbar M, Pierik AJ, Bremer E (2008) Synthesis and uptake of the compatible solutes ectoine and 5-hydroxyectoine by Streptomyces coelicolor A3(2) in response to salt and heat stresses. Appl Environ Microbiol 74:7286–7296. https://doi.org/10.1128/AEM.00768-08
Cánovas D, Fletcher SA, Hayashi M, Csonka LN (2001) Role of trehalose in growth at high temperature of Salmonella enterica serovar Typhimurium. J Bacteriol 183:3365–3371. https://doi.org/10.1128/JB.183.11.3365-3371.2001
Cánovas D, Vargas C, Calderón MI, Ventosa A, Nieto JJ (1998) Characterization of the genes for the biosynthesis of the compatible solute ectoine in the moderately halophilic bacterium Halomonas elongata DSM 3043. Syst Appl Microbiol 21:487–497. https://doi.org/10.1016/S0723-2020(98)80060-X
Costa J, Empadinhas N, da Costa MS (2007) Glucosylglycerate biosynthesis in the deepest lineage of the Bacteria: characterization of the thermophilic proteins GpgS and GpgP from Persephonella marina. J Bacteriol 189:1648–1654. https://doi.org/10.1128/JB.00841-06
da Costa MS, Santos H, Galinski EA (1998) An overview of the role and diversity of compatible solutes in Bacteria and Archaea. Adv Biochem Eng Biotechnol 61:117–153. https://doi.org/10.1007/BFb0102291
Dandapath I, Chatterjee M, Sarkar D, Gupta A, Rabbani G, Minakshi R (2017) Bacterial osmolyte system and its physiological roles. In: Singh LR, Dar TA (eds) Cellular osmolytes. Springer, Singapore, 229–249. https://doi.org/10.1007/978-981-10-3707-8
de Alvarenga LV, Lucius S, Vaz M, Araújo WL, Hagemann M (2020) The novel strain Desmonostoc salinum CCM-UFV059 shows higher salt and desiccation resistance compared to the model strain Nostoc sp. PCC 7120. J Phycol 56:496–506. https://doi.org/10.1111/jpy.12968
Franceus J, DecuyperlL DM, Desmet T (2018) Exploring the sequence diversity in glycoside hydrolase family 13_18 reveals a novel glucosylglycerol phosphorylase. Appl Microbiol Biotechnol 102:3183–3191. https://doi.org/10.1007/s00253-018-8856-1
Franceus J, Pinel D, Desmet T (2017) Glucosylglycerate phosphorylase, an enzyme with novel specificity involved in compatible solute metabolism. Appl Environ Microbiol 83:e01434-e1517. https://doi.org/10.1128/AEM.01434-17
Goedl C, Schwarz A, Minani A, Nidetzky B (2007) Recombinant sucrose phosphorylase from Leuconostoc mesenteroides: characterization, kinetic studies of transglucosylation, and application of immobilised enzyme for production of α-D-glucose 1-phosphate. J Biotechnol 129:77–86. https://doi.org/10.1016/j.jbiotec.2006.11.019
Hagemann M, Erdmann N (1994) Activation and pathway of glucosylglycerol synthesis in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology-SGM 140:1427–1431. https://doi.org/10.1099/00221287-140-6-1427
Hagemann M, Ribbeck-Busch K, Klähn S, Hasse D, Steinbruch R, Berg G (2008) The plant-associated bacterium Stenotrophomonas rhizophila expresses a new enzyme for the synthesis of the compatible solute glucosylglycerol. J Bacteriol 190:5898–5906. https://doi.org/10.1128/JB.00643-08
Hagemann M, Richter S, Zuther E, Schoor A (1996) Characterization of a glucosylglycerol-phosphate-accumulating, salt-sensitive mutant of the cyanobacterium Synechocystis sp. strain PCC 6803. Arch Microbiol 166:83–91. https://doi.org/10.1007/s002030050360
Hagemann M, Schoor A, Jeanjean R, Zuther E, Joset F (1997) The stpA gene from Synechocystis sp. strain PCC 6803 encodes the glucosylglycerol-phosphate phosphatase involved in cyanobacterial osmotic response to salt shock. J Bacteriol 179:1727–1733. https://doi.org/10.1128/jb.179.5.1727-1733.1997
Kempf B, Bremer E (1998) Uptake and synthesis of compatible solutes as microbial stress responses to high-osmolality environments. Arch Microbiol 170:319–330. https://doi.org/10.1007/s002030050649
Kirsch F, Klähn S, Hagemann M (2019) Salt-regulated accumulation of the compatible solutes sucrose and glucosylglycerol in cyanobacteria and its biotechnological potential. Front Microbiol 10:2139. https://doi.org/10.3389/fmicb.2019.02139
Kirsch F, Luo Q, Lu X, Hagemann M (2018) Inactivation of invertase enhances sucrose production in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology-SGM 164:1220–1228. https://doi.org/10.1099/mic.0.000708
Kirsch F, Pade N, Klähn S, Hess WR, Hagemann M (2017) The glucosylglycerol-degrading enzyme GghA is involved in acclimation to fluctuating salinities by the cyanobacterium Synechocystis sp. strain PCC 6803. Microbiology-SGM 163:1319–1328. https://doi.org/10.1099/mic.0.000518
Klähn S, Hagemann M (2011) Compatible solute biosynthesis in cyanobacteria. Environ Microbiol 13:551–562. https://doi.org/10.1111/j.1462-2920.2010.02366.x
Klähn S, Steglich C, Hess WR, Hagemann M (2010) Glucosylglycerate: a secondary compatible solute common to marine cyanobacteria from nitrogen-poor environments. Environ Microbiol 12:83–94. https://doi.org/10.1111/j.1462-2920.2009.02045.x
Kuhlmann AU, Bursy J, Gimpel S, Hoffmann T, Bremer E (2008) Synthesis of the compatible solute ectoine in Virgibacillus pantothenticus is triggered by high salinity and low growth temperature. Appl Environ Microbiol 74:4560–4563. https://doi.org/10.1128/AEM.00492-08
Lamark T, Kaasen I, Eshoo MW, Falkenberg P, McDougall J, Strøm AR (1991) DNA sequence and analysis of the bet genes encoding the osmoregulatory choline-glycine betaine pathway of Escherichia coli. Mol Microbiol 5:1049–1064. https://doi.org/10.1111/j.1365-2958.1991.tb01877.x
Luo Q, Duan Y, Lu X (2022) Biological sources, metabolism, and production of glucosylglycerols, a group of natural glucosides of biotechnological interest. Biotechnol Adv 59:107964. https://doi.org/10.1016/j.biotechadv.2022.107964
Marin K, Huckauf J, Fulda S, Hagemann M (2002) Salt-dependent expression of glucosylglycerol-phosphate synthase, involved in osmolyte synthesis in the cyanobacterium Synechocystis sp. strain PCC 6803. J Bacteriol 184:2870–2877. https://doi.org/10.1128/JB.184.11.2870-2877.2002
Marin K, Zuther E, Kerstan T, Kunert A, Hagemann M (1998) The ggpS gene from Synechocystis sp. strain PCC 6803 encoding glucosyl-glycerol-phosphate synthase is involved in osmolyte synthesis. J Bacteriol 180:4843–4849. https://doi.org/10.1128/JB.180.18.4843-4849.1998
Mikkat S, Hagemann M, Schoor A (1996) Active transport of glucosylglycerol is involved in salt adaptation of the cyanobacterium Synechocystis sp. strain PCC 6803. Microbiology 142:1725–1732. https://doi.org/10.1099/13500872-142-7-1725
Nakahira Y, Ogawa A, Asano H, Oyama T, Tozawa Y (2013) Theophylline-dependent riboswitch as a novel genetic tool for strict regulation of protein expression in cyanobacterium Synechococcus elongatus PCC 7942. Plant Cell Physiol 54:1724–1735. https://doi.org/10.1093/pcp/pct115
Nie Y, Su X, Wu D, Zhang R, Wang R, Zhao Z, Xamxidin M, Sun C, Wu M (2021) Marinobacter caseinilyticus sp. nov., isolated from saline soil. Curr Microbiol 78:1045–1052. https://doi.org/10.1007/s00284-021-02351-w
Novak JF, Stirnberg M, Roenneke B, Marin K (2011) A novel mechanism of osmosensing, a salt-dependent protein-nucleic acid interaction in the cyanobacterium Synechocystis species PCC 6803. J Biol Chem 286:3235–3241. https://doi.org/10.1074/jbc.M110.157032
Nyyssola A, Kerovuo J, Kaukinen P, von Weymarn N, Reinikainen T (2000) Extreme halophiles synthesize betaine from glycine by methylation. J Biol Chem 275:22196–22201. https://doi.org/10.1074/jbc.M910111199
Qiao C, Duan Y, Zhang M, Hagemann M, Luo Q, Lu X (2018) Effects of reduced and enhanced glycogen pools on salt-induced sucrose production in a sucrose-secreting strain of Synechococcus elongatus PCC 7942. Appl Environ Microbiol 84:e02023-e2117. https://doi.org/10.1128/AEM.02023-17
Qiao C, Zhang M, Luo Q, Lu X (2019) Identification of two two-component signal transduction mutants with enhanced sucrose biosynthesis in Synechococcus elongatus PCC 7942. J Basic Microbiol 59:465–476. https://doi.org/10.1002/jobm.201800676
Savakis P, Tan X, Qiao C, Song K, Lu X, Hellingwerf KJ, Branco Dos Santos F (2016) Slr1670 from Synechocystis sp. PCC 6803 is required for the re-assimilation of the osmolyte glucosylglycerol. Front Microbiol 7:1350. https://doi.org/10.3389/fmicb.2016.01350
Schwibbert K, Marin-Sanguino A, Bagyan I, Heidrich G, Lentzen G, Seitz H, Rampp M, Schuster SC, Klenk HP, Pfeiffer F, Oesterhelt D, Kunte HJ (2011) A blueprint of ectoine metabolism from the genome of the industrial producer Halomonas elongata DSM 2581T. Environ Microbiol 13:1973–1994. https://doi.org/10.1111/j.1462-2920.2010.02336.x
Sun X, Zhang T, Liu Y, Chen P, Qin H, Yang J, Sun Y (2023) Self-assembled multienzyme complex facilitates synthesis of glucosylglycerol from maltodextrin and glycerol. J Sci Food Agric 104:266–272. https://doi.org/10.1002/jsfa.12907
Yoo Y, Lee H, Kwon BO, Khim JS, Baek S, Pathiraja D, Park B, Choi IG, Kim GH, Kim BS, Kim JJ (2020) Marinobacter halodurans sp. nov., a halophilic bacterium isolated from sediment of a salt flat. Int J Syst Evol Microbiol 70:6294–6300. https://doi.org/10.1099/ijsem.0.004530
Zhang DC, Li HR, Xin YH, Chi ZM, Zhou PJ, Yu Y (2008) Marinobacter psychrophilus sp. nov., a psychrophilic bacterium isolated from the Arctic. Int J Syst Evol Microbiol 58:1463–1466. https://doi.org/10.1099/ijs.0.65690-0
Zhang MX, Li AZ, Wu Q, Yao Q, Zhu HH (2020a) Marinobacter denitrificans sp. nov., isolated from marine sediment of southern Scott Coast. Antarctica Int J Syst Evol Microbiol 70:2918–2924. https://doi.org/10.1099/ijsem.0.004120
Zhang T, Yang J, Tian C, Ren C, Chen P, Men Y, Sun Y (2020b) High-yield biosynthesis of glucosylglycerol through coupling phosphorolysis and transglycosylation reactions. J Agric Food Chem 68:15249–15256. https://doi.org/10.1021/acs.jafc.0c04851
Zhou J, Zhang H, Meng H, Zhu Y, Bao G, Zhang Y, Li Y, Ma Y (2014) Discovery of a super-strong promoter enables efficient production of heterologous proteins in cyanobacteria. Sci Rep 4:4500. https://doi.org/10.1038/srep04500
Acknowledgements
Many thanks to Marine Culture Collection of China for providing the Marinobacter strain.
Funding
We acknowledge the financial support from the Shandong Provincial Natural Science Foundation project (ZR2023MC175 to QL), the Shandong Taishan Scholarship (to XL), and the National Science Foundation of China project (31872622 to QL).
Author information
Authors and Affiliations
Contributions
QL conceived and designed research. LC and ZZ conducted experiments. QL and DZ analyzed data. QL and LC wrote the manuscript. DZ and XL revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
All authors mentioned in the manuscript have given consent for submission and subsequent publication of the manuscript.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cheng, L., Zhang, Z., Zhu, D. et al. Glucosylglycerol phosphorylase, a potential novel pathway of microbial glucosylglycerol catabolism. Appl Microbiol Biotechnol 108, 214 (2024). https://doi.org/10.1007/s00253-024-13035-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13035-3