Abstract
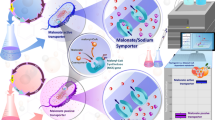
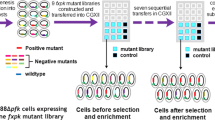
Our previous study’s introduction of the malonic acid assimilation pathway into Escherichia coli enabled biosynthesis of 3-Hydroxypropionate (3-HP) from malonate. However, the relatively low uptake activity of tripartite ATP-independent periplasmic (TRAP) malonic acid transporter (MatPQM) is considered rate-limiting in malonate utilization. Here, to improve the transport performance of this importer, MatP variants were obtained via directed evolution and a novel developed enzyme-inhibition-based high throughput screening approach. This plate chromogenic screening method is based on the fact that malonic acid inhibits both of succinate dehydrogenase activity and further the capability of the reduction of methylene-blue to methylene-white. The best mutant E103G/S194G/Y218H/L235P/N272S showed twofold increased transport efficiency compared to the wild-type. ITC assay and structural analysis revealed that increased binding affinity of the mutant to the ligand was the reason for improved uptake activity of MatPQM. Finally, the engineered strain harboring the evolved mutant produced 20.08 g/L 3-HP with the yield of 0.87 mol/mol malonate in a bioreactor. Therefore, the well-established directed evolution strategy can be regarded as the reference work for other TRAP-type transporters engineering. And, this transporter mutant with enhanced malonic acid uptake activity has broad applications in the microbial biosynthesis of malonyl-CoA-derived valuable compounds in bacteria.
Key points
• We reported directed evolution of a TRAP-type malonic acid transporter.
• We found the enhanced malonate uptake activity of mutant lies in improved affinity.
• We enhanced 3-HP bioproduction with high yield by employing the best mutant.






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Bali AP, Genee HJ, Sommer MOA (2018) Directed evolution of membrane transport using synthetic selections. ACS Synth Biol 7(3):789–793. https://doi.org/10.1021/acssynbio.7b00407
Bisson C, Salmon RC, West L, Rafferty JB, Hitchcock A, Thomas GH, Kelly DJ (2022) The structural basis for high-affinity uptake of lignin-derived aromatic compounds by proteobacterial TRAP transporters. FEBS J 289(2):436–456. https://doi.org/10.1111/febs.16156
Chang Z, Dai W, Mao Y, Cui Z, Zhang Z, Wang Z, Ma H, Chen T (2022) Enhanced 3-hydroxypropionic acid production from acetate via the malonyl-CoA pathway in Corynebacterium glutamicum. Front Bioeng Biotech 9:808258. https://doi.org/10.3389/fbioe.2021.808258
Chen AM, Wang YB, Jie S, Yu AY, Luo L, Yu GQ, Zhu JB, Wang YZ (2010) Identification of a TRAP transporter for malonate transport and its expression regulated by GtrA from Sinorhizobium meliloti. Res Microbiol 161(7):556–564. https://doi.org/10.1016/j.resmic.2010.05.004
Chen Z, Huang JH, Wu Y, Wu WJ, Zhang Y, Liu DH (2017) Metabolic engineering of corynebacterium glutamicum for the production of 3-hydroxypropionic acid from glucose and xylose. Metab Eng 39:151–158. https://doi.org/10.1016/j.ymben.2016.11.009
Cheng Z, Jiang J, Wu H, Li Z, Ye Q (2016) Enhanced production of 3-hydroxypropionic acid from glucose via malonyl-CoA pathway by engineered Escherichia coli. Bioresour Technol 200:897–904. https://doi.org/10.1016/j.biortech.2015.10.107
De Fouchécour F, Sánchez-Castañeda A-K, Saulou-Bérion C, Spinnler HE (2018) Process engineering for microbial production of 3-hydroxypropionic acid. Biotechnol Adv 36(4):1207–1222. https://doi.org/10.1016/j.biotechadv.2018.03.020
Della Pina C, Falletta E, Rossi M (2011) A green approach to chemical building blocks The case of 3-hydroxypropanoic acid. Green Chem 13(7):1624–1632. https://doi.org/10.1039/C1GC15052A
Deng X, Shi B, Ye Z, Huang M, Chen R, Cai Y, Kuang Z, Sun X, Bian G, Deng Z, Liu T (2022) Systematic identification of ocimum sanctum sesquiterpenoid synthases and (-)-eremophilene overproduction in engineered yeast. Metab Eng 69:122–133. https://doi.org/10.1016/j.ymben.2021.11.005
Eberhardt J, Santos-Martins D, Tillack AF, Forli S (2021) AutoDock vina 1.2.0: new docking methods expanded force field and python bindings. J Chem Inf Model 61(8):3891–3898
Fischer M, Zhang QY, Hubbard RE, Thomas GH (2010) Caught in a TRAP: substrate-binding proteins in secondary transport. Trends Microbiol 18(10):471–478. https://doi.org/10.1016/j.tim.2010.06.009
Fischer M, Hopkins AP, Severi E, Hawkhead J, Bawdon D, Watts AG, Hubbard RE, Thomas GH (2015) Tripartite ATP-independent periplasmic (TRAP) transporters use an arginine-mediated selectivity filter for high affinity substrate binding. J Biol Chem 290(45):27113–27123. https://doi.org/10.1074/jbc.M115.656603
Gargiulo S, Soumillion P (2021) Directed evolution for enzyme development in biocatalysis. Curr Opin Chem Biol 61:107–113. https://doi.org/10.1016/j.cbpa.2020.11.006
Jers C, Kalantari A, Garg A, Mijakovic I (2019) Production of 3-hydroxypropanoic acid from glycerol by metabolically engineered bacteria. Front Bioeng Biotechnol 7:124. https://doi.org/10.3389/fbioe.2019.124
Kumar V, Ashok S, Park S (2013) Recent advances in biological production of 3-hydroxypropionic acid. Biotechnol Adv 31(6):945–961. https://doi.org/10.1016/j.biotechadv.2013.02.008
Lai N, Luo Y, Fei P, Hu P, Wu H (2021) One stone two birds: Biosynthesis of 3-hydroxypropionic acid from CO(2) and syngas-derived acetic acid in Escherichia coli. Synthetic and Systems Biotechnology 6(3):144–152. https://doi.org/10.1016/j.synbio.2021.06.003
Lama S, Kim Y, Nguyen DT, Im CH, Sankaranarayanan M, Park S (2021) Production of 3-hydroxypropionic acid from acetate using metabolically-engineered and glucose-grown Escherichia coli. Bioresour Technol 320:124362. https://doi.org/10.1016/j.biortech.2020.124362
Lashkov AA, Tolmachev IV, Eistrikh-Heller PA, Rubinsky SV (2021) PyFepRestr: plugin to pymol molecular graphics system for calculating the free energy of ligand-receptor binding. Crystallogr Rep+ 66(5):861–865. https://doi.org/10.1134/S1063774521050126
Lian JZ, Li YL, HamediRad M, Zhao HM (2014) Directed evolution of a cellodextrin transporter for improved biofuel production under anaerobic conditions in Saccharomyces cerevisiae. Biotechnol Bioeng 111(8):1521–1531. https://doi.org/10.1002/bit.25214
Liang B, Sun G, Wang Z, Xiao J, Yang J (2019) Production of 3-hydroxypropionate using a novel malonyl-CoA-mediated biosynthetic pathway in genetically engineered E coli strain. Green Chem 21(22):6103–6115. https://doi.org/10.1039/c9gc02286d
Lindlbauer KA, Marx H, Sauer M (2017) 3-hydroxypropionaldehyde production from crude glycerol by lactobacillus diolivorans with enhanced glycerol uptake. Biotechnol Biofuels 10:295. https://doi.org/10.1186/S13068-017-0982-Y
Liu C, Ding Y, Zhang R, Liu H, Xian M, Zhao G (2016) Functional balance between enzymes in malonyl-CoA pathway for 3-hydroxypropionate biosynthesis. Metab Eng 34:104–111. https://doi.org/10.1016/j.ymben.2016.01.001
Liu C, Ding Y, Xian M, Liu M, Liu H, Ma Q, Zhao G (2017) Malonyl-CoA pathway: a promising route for 3-hydroxypropionate biosynthesis. Crit Rev Biotechnol 37(7):933–941. https://doi.org/10.1080/07388551.2016.1272093
Liu B, Xiang S, Zhao G, Wang B, Ma Y, Liu W, Tao Y (2019) Efficient production of 3-hydroxypropionate from fatty acids feedstock in Escherichia coli. Metab Eng 51:121–130. https://doi.org/10.1016/j.ymben.2018.10.003
Liu CL, Xue K, Yang Y, Liu X, Li Y, Lee TS, Bai Z, Tan T (2022) Metabolic engineering strategies for sesquiterpene production in microorganism. Crit Rev Biotechnol 42(1):73–92. https://doi.org/10.1080/07388551.2021.1924112
Matsakas L, Hrůzová K, Rova U, Christakopoulos P (2018) Biological production of 3-hydroxypropionic acid: an update on the current status. Fermentation 4(1):13. https://doi.org/10.3390/fermentation4010013
Mulligan C, Geertsma ER, Severi E, Kelly DJ, Poolman B, Thomas GH (2009) The substrate-binding protein imposes directionality on an electrochemical sodium gradient-driven TRAP transporter. Proc Natl Acad Sci USA 106(6):1778–1783. https://doi.org/10.1073/pnas.0809979106
Park SR, Ahn MS, Han AR, Park JW, Yoon YJ (2011) Enhanced flavonoid production in Streptomyces venezuelae via metabolic engineering. J Microbiol Biotechnol 21(11):1143–1146. https://doi.org/10.4014/jmb.1108.08012
Suyama A, Higuchi Y, Urushihara M, Maeda Y, Takegawa K (2017) Production of 3-hydroxypropionic acid via the malonyl-CoA pathway using recombinant fission yeast strains. J Biosci Bioeng 124(4):392–399. https://doi.org/10.1016/j.jbiosc.2017.04.015
Takayama S, Ozaki A, Konishi R, Otomo C, Kishida M, Hirata Y, Matsumoto T, Tanaka T, Kondo A (2018) Enhancing 3-hydroxypropionic acid production in combination with sugar supply engineering by cell surface-display and metabolic engineering of Schizosaccharomyces pombe. Microb Cell Fact 17(1):176. https://doi.org/10.1186/s12934-018-1025-5
Valdehuesa KNG, Liu H, Nisola GM, Chung W-J, Lee SH, Park SJ (2013) Recent advances in the metabolic engineering of microorganisms for the production of 3-hydroxypropionic acid as C3 platform chemical. Appl Microbiol Biotechnol 97(8):3309–3321. https://doi.org/10.1007/s00253-013-4802-4
Vetting MW, Al-Obaidi N, Zhao S, San Francisco B, Kim J, Wichelecki DJ, Bouvier JT, Solbiati JO, Vu H, Zhang X (2015) Experimental strategies for functional annotation and metabolism discovery: targeted screening of solute binding proteins and unbiased panning of metabolomes. Biochemistry 54(3):909–931. https://doi.org/10.1021/bi501388y
Wang Q, Yang P, Xian M, Feng L, Wang J, Zhao G (2014) Metabolic engineering of Escherichia coli for poly (3-hydroxypropionate) production from glycerol and glucose. Biotechnol Lett 36(11):2257–2262. https://doi.org/10.1007/s10529-014-1600-8
Wang S, Jin X, Jiang W, Wang Q, Qi Q, Liang Q (2022) The expression modulation of the key enzyme Acc for highly efficient 3-hydroxypropionic acid production. Front Microbiol 13:902848. https://doi.org/10.3389/fmicb.2022.902848
Werpy T, Petersen G (2004) Top value added chemicals from biomass office of energy efficiency and renewable energy. US Department of Energy, Washington DC
Yang JY, Zhang Y (2015) I-tasser server: new development for protein structure and function predictions. Nucleic Acids Res 43(W1):W174–W181. https://doi.org/10.1093/nar/gkv342
Yoon J, Oh MK (2022) Strategies for biosynthesis of C1 gas-derived polyhydroxyalkanoates: a review. Bioresour Technol 344(Pt B):126307. https://doi.org/10.1016/j.biortech.2021.126307
Yu W, Cao X, Gao J, Zhou YJ (2022) Overproduction of 3-hydroxypropionate in a super yeast chassis. Biores Technol 361:127690. https://doi.org/10.1016/j.biortech.2022.127690
Zhao P, Tian PF (2021) Biosynthesis pathways and strategies for improving 3-hydroxypropionic acid production in bacteria. World J Microb Biot 37(7):117. https://doi.org/10.1007/s11274-021-03091-6
Funding
This work was supported by grants from the National Natural Science Foundation of China (22278233,31860011), the Natural Science Foundation of Shandong Province, China (ZR2020MC009), the “First class grassland science discipline” program in Shandong Province. The authors would also like to thank the “Laboratory for Agricultural Molecular Biology of Qingdao Agricultural University” for providing laboratory apparatus.
Author information
Authors and Affiliations
Contributions
BL and JY designed the study. BL, XZ, CM, and LW performed the study. BL and XZ analyzed the data. BL, XZ, and JY wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants performed by any of the authors. The authors have no relevant financial or non-financial interests to disclose.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, B., Zhang, X., Meng, C. et al. Directed evolution of tripartite ATP-independent periplasmic transporter for 3-Hydroxypropionate biosynthesis. Appl Microbiol Biotechnol 107, 663–676 (2023). https://doi.org/10.1007/s00253-022-12330-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12330-1