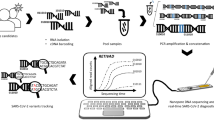
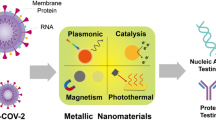
Abstract
The COVID-19, MERS-CoV, and SARS-CoV are hazardous epidemics that have resulted in many deaths which caused a worldwide debate. Despite control efforts, SARS-CoV-2 continues to spread, and the fast spread of this highly infectious illness has posed a grave threat to global health. The effect of the SARS-CoV-2 mutation, on the other hand, has been characterized by worrying variations that modify viral characteristics in response to the changing resistance profile of the human population. The repeated transmission of virus mutation indicates that epidemics are likely to occur. Therefore, an early identification system of ongoing mutations of SARS-CoV-2 will provide essential insights for planning and avoiding future outbreaks. This article discussed the following highlights: First, comparing the omicron mutation with other variants; second, analysis and evaluation of the spread rate of the SARS-CoV 2 variations in the countries; third, identification of mutation areas in spike protein; and fourth, it discussed the photonics approaches enabled with artificial intelligence. Therefore, our goal is to identify the SARS-CoV 2 virus directly without the need for sample preparation or molecular amplification procedures. Furthermore, by connecting through the optical network, the COVID-19 test becomes a component of the Internet of healthcare things to improve precision, service efficiency, and flexibility and provide greater availability for the evaluation of the general population.
Key points
• A proposed framework of photonics based on AI for identifying and sorting SARS-CoV 2 mutations.
• Comparative scatter rates Omicron variant and other SARS-CoV 2 variations per country.
• Evaluating mutation areas in spike protein and AI enabled by photonic technologies for SARS-CoV 2 virus detection.
Similar content being viewed by others
Introduction
In 2020, the severe acute respiratory syndrome coronavirus (SARS-CoV 2) caused a global epidemic. The original virus strain was identified in Wuhan, Hubei Province, China (Loo et al. 2021). Although vaccinations have been introduced throughout the world, the number of infected cases continues to increase due to the effects of new SARS-CoV-2 variations. From an evolutionary point of view, Variations of Concern (VOCs) are variants with selected evolutionary advantages (Lauring and Hodcroft 2021). These new variations appeared in different places at the same time in September 2020 but are not related to each other. A mutate called B.1.1.7 first appeared in the UK (Tang et al. 2021). Then, it was the B.1.351 in South Africa, B.1.617 in India, and P.1 in Brazil (Samarasekera 2021; Thye et al. 2021). Consequently, the World Health Organization (WHO) recognized variant B.1.1.529 as a worrying variant on November 26, 2021 and renamed it as Omicron. Figure 1 describes the coronavirus disease (COVID-19) mutations that occur. An initial imaging of the newly discovered COVID-19 variant Omicron, manufactured and published by the famous Bambino Gesu Hospital in Rome, was recently reported in the literature. Different strains, on the other hand, contain several changes in their spike protein, suggesting that they are more pathogenic. Mutations in the gene can potentially alter receptors to achieve cell entry and viral immune evasion and immunogenicity, as the SARS-CoV 2 spike protein binds to the angiotensin converting enzyme 2 (ACE2) (Watanabe et al. 2020). Interestingly, changes in the spike protein are of particular concern as vaccination encourages the formation of antibodies against its contents. SARS-CoV 2 identities are based on spike sequences generated from the early Wuhan strain and include recombinant protein, inactivated virus, RNA, and virally vectorized platforms (Krammer 2020). This current version contains significantly more mutations than the delta version. Preliminary research states that Omicron latent infection may be more difficult for those who previously had COVID-19 than other worrying variations. On the other hand, the virus has shown us in the past that different strains can cause different degrees of disease and different symptoms. For example, Alpha, which was discovered in the UK, was more transmissible than previous variations, but the symptoms were the same. The Delta which was originally discovered in India is much more contagious and deadly than COVID-19. The genomes and proteins of numerous people were also compared and contrasted using various bioinformatic methods (Srinivasan et al. 2020). This research helped identify the virus’ close association with other people using a SARS-CoV 2 protein information database in open sources. It studied numerous mutations in these proteins in isolates from various geographical regions. Understanding the alterations in the virus’ multiple proteins can aid in unraveling the riddle of the increased COVID-19 transmission rates that resulted in the pandemic and give direction for targeted viral management (Consortium 2021). However, two of the most common symptoms of COVID-19, chronic cough and loss of smell and taste, are less common, but more people are reporting them. Omicron virus has been identified as a worrying variant to the extent that the WHO issues recommendations for nations around the globe, including increased surveillance and case sequencing. Collaboration in the publication of genomic sequences in publicly accessible databases has been done. However, future pandemic threats will require the development of diagnostics, drugs, and vaccinations to counter them (Jakhmola et al. 2021). Furthermore, not every healthcare center or hospital, particularly smaller organizations with limited resources, can handle the massive workload caused by increased demand due to insufficient PCR testing capabilities. Finally, the number of kits and reagents available is inadequate to meet spiking demand in general (Wan et al. 2021). Medical telemonitoring provided help to the caregivers staff to monitor and decipher the patient’s therapeutic indications. Practical or automated methods can be used to perform these tasks. Therapeutic demonstrations which can be helpful towards doctors can be conducted remotely with the assistance of a doctor/health practitioner. Patients in need of medical attention can seek advice from a doctor by phone or computer screenshot.
COVID-19 mutations
Many viruses, including SARS-CoV-2, which is responsible for COVID-19, evolve over time. Most changes have little to no effect on the properties of the virus. However, specific changes can affect the characteristics of the virus, such as the speed of spread, the severity of comorbidities or the performance of vaccinations, therapeutic drugs, diagnostic tools, or other public health and welfare measures. Although various systems for detecting COVID-19 particles come in handy, many problems limit their usefulness. In addition, there are the following challenges: Sensitivity and accuracy are reduced, preparation and cleaning of the samples require time, and the complex operation of the devices. Hence, detection methods for COVID-19 need to be improved urgently. These methods need to be implemented for greater precision, service efficiency, and flexibility, as well as widespread availability for the assessment of the general population (Taha et al. 2021a, b). A mutation is a single change in the genome or genetic code of a virus. Mutations are common but rarely affect the properties of the virus. Many mutations in SARS-CoV 2 cause changes in the properties of the virus and affect the response to the pandemic. Preliminary analysis shows that at least one mutation, common to three new worrying variants, appears to increase transmission efficiency (Unlu et al. 2021). It is imperative that vaccines are produced as soon as possible and that genetic monitoring needs to be stepped up to detect variations when they first emerge and wander around the world. Since its discovery in December 2020, the Delta variety has become the most common variety in India and the UK. In addition, the CDC estimates that more than 20% of cases in the USA have been assigned to this strain. However, medical professionals assume that the Delta strain is more contagious than the other variants of the virus. WHO organization has identified five mutations in the SARS-CoV 2 virus. They were named after their Greek alphabet, Beta, Gamma, and Delta, starting with Alpha. However, in the Omicron variant, the 15th letter was used instead, which classifies it as a very risky variant worldwide. Delta variation, while being the most common, accounts for more than 99% of sequenced cases worldwide (Almubaid and Al-Mubaid 2021; Teng et al. 2020). In their thorough study of the mechanisms of viral mutation, Sanjuan and colleagues found that RNA viruses mutate faster than DNA viruses (Sanjuán and Domingo-Calap 2016). The D614G mutation makes a particular cavity between the spike protein subunits of the repeating unit. On the contrary, the intermolecular hydrogen bonding potential between the spike protein subunits at that location is abolished when S982A is substituted. Therefore, mutations in B.1.1.7 increase the affinity of SARS-CoV 2 for ACE2, and substitutions for A570D, D614G and S982A can improve the dynamic viral fusion process by reducing the intermolecular stability of the spike protein subunits (Ostrov 2021). The spike protein D614G mutation can be identified based on quantitative PCR high-resolution melting (qPCR-HRM). Six SARS-CoV-2 RNA samples were converted to DNA and analyzed using the qPCR-HRM method to determine whether the spike protein contained the D614G mutation. The primers designed are to target the D614G mutated spike region selectively (Gazali et al. 2021). A study in England reported that variant B.1.1.7 had a 4390% higher reproduction rate than previous forms (Davies Nicholas et al. 2021). Compared to the precursor isolate with D614G exchange, Hoffman et al. observed that B.1.1.7, B.1.351, and P.1 show no significant differences in spike protein stability or entry kinetics (Hoffmann et al. 2021). As seen in Fig. 2, SARS-CoV 2 has been changing, as have all viruses, since its discovery in late 2019. Changes in the genetic coding of the spike protein can affect its capacity to infect cells.
Omicron contrast to other variants
B.1.1.529, a novel SARS-CoV 2 variant, was discovered in Botswana and South Africa at the beginning of November 2021 and classified by the WHO as VOC Omicron on November 26, 2021. This is in order to determine the transferability, severity, and ability to bypass the immune system in this strain, due to its improvement to the spike protein (Gu et al. 2021). The spike protein, the main focus of the immune response, has been modified and highlighted in this strain. Changes in invariants such as Delta and Alpha have increased infectivity and show a better ability to avoid infection-blocking antibodies (Callaway 2021). The latest finding of the SARS-CoV 2 Omicron variant (B.1.1.529) is contributing to the apparently ongoing wildfire of the almost 2-year-old global COVID-19 pandemic. At least 32 mutations occur in the spike protein alone compared to the sixteen mutations in the already highly infectious Delta version. Additional proteins such as NSP12 and NSP14, which are important for virus replication, are also found. It is estimated to be at least three times as infectious as the original SARS-CoV 2 strain and maybe even more so than the Delta strain. There have been reports of it in many nations, including Australia, Belgium, Botswana, Germany, Hong Kong, Israel, Italy, the Netherlands, and the UK. Additionally, numerous countries have implemented tight quarantine policies for patients (Gao et al. 2021; Rao and Singh 2021). Recent published research has highlighted the variant mutated in the Delta variant in the literature. It is discovered that a spike mutation E465A was present in 15 Delta variation sequences and that n = 1756 of Delta (B.1.617.2) variant mutations comprise 20% of the viral genome (Kannan et al. 2021). Furthermore, it has several mutated forms, but the Omicron variant is unique in that it is the heaviest of them all. Table 1 shows a comprehensive comparison of Omicron with other variants. Although SARS-CoV 2 vaccines are developing rapidly, little is known about the immunological correlates of protection or the virus’ ability to escape the human immune response through mutation and recombination (Kottier et al. 1995). According to a report of the B.1.1.7 variant in England, this variation has a 43–90% greater reproduction rate than prior forms.
Analysis for SARS-CoV 2 variants
Viruses are subject to mutation and evolution in response to selection forces from their environment, resulting in variations that can be more virulent. Health professionals are particularly concerned about the spread of these novel strains and their reinfection rates, the severity of the disease, and the effectiveness of vaccination (Volz et al. 2021; Zhou et al. 2021). Omicron GR/484A (B.1.1.529) exhibits a particularly interesting combination of spike amino acid changes as it includes those previously identified as affecting receptor binding and antibody loss. It must constantly monitor all low-frequency variations with potentially important changes to see whether the distribution of these variants is influenced by immune escape or altered receptor contacts (Yurkovetskiy et al. 2020). Researchers in Botswana, Hong Kong, and South Africa made important contributions to the rapid discovery of Omicron protein subunits SARS-CoV 2 spike glycoprotein trimer in combination with the human host cell receptor ACE2 (1 unit shown, green band). Figure 3 shows the protein subunits SARS-CoV 2 spike glycoprotein trimer in combination with the human host cell receptor ACE2 (1 unit shown, green band). The colored spheres mark the places where the B.1.1.529 line has changed. In addition, bright orange or cyan is used for deletions and green for inserts to distinguish between changes that have phenotypic effects and those that do not. We collected information from the GISAID website, which facilitates the quick exchange of information on all influenza viruses, including COVID-19. It contains genetic sequences and associated clinical and epidemiological data for human viruses and regional and species-specific data for avian and other animal viruses. This information helps researchers understand how viruses change and spread during epidemics and pandemics. Data is available on GISAID (https://www.gisaid.org/). However, the functional characterization of these mutations remains unclear.
Shows the spike protein to the Omicron variant and the rate of spread with other variants: A. Amino acid changes change the 3-D structure of a spike. B. Monitoring of Omicron by country penetration rate. C. Representation of the comparative scatter rates SARS-CoV 2 variations: Alpha, Beta, Gamma, and Delta sequentially. Data available on GISAID (https://www.gisaid.org/)
D614G mutation of the spike protein
Similarity between SARS-CoV 2, severe acute respiratory syndrome coronavirus (SARS-CoV), and Middle Eastern respiratory syndrome coronavirus (MERS-CoV) have been discovered, all of which induce life-threatening respiratory diseases. The ORFs genome is roughly 30 kb in length and encodes around 16 non-structural and four structural proteins, including the envelope (E), the membrane (M), the nucleocapsid (N), and the spike (S) (Dearlove et al. 2020). However, due to the rapid spread of virus mutations, there are uncertainties about whether vaccination will be effective worldwide. When the virus mutates, it can spread these varieties in populations and over long periods of time. Furthermore, the ACE2 receptors on the surface of host cells are occupied by the receptor binding domain (RBD) of the S1 subunit of the S protein, and this interaction facilitates the entry of virus into the host cell. It has been suggested that it affects the infectivity of the virus. The rapid spread of the mutated D614G virus was related to the increased infectivity of SARS-CoV 2, which carries this mutation (Grubaugh et al. 2020), (Weissman et al. 2020). The biology of SARS-CoV 2 is significantly affected by the D614G mutation, and the development of new tools to identify it quickly and accurately is crucial. SARS-CoV 2 and its D614G mutation are essential to contain the pandemic. Researchers have used surface plasmon resonance (SPR) as a potential tool to study how the binding of the protein from SARS-CoV 2 to human ACE2 is affected by the D614G mutation (Yurkovetskiy et al. 2020; Zhang et al. 2021). Like antibodies, the N protein of SARS-CoV 2 can be detected using aptamers in ELISA and colloidal immunochromatographic gold strips (Zhang et al. 2020b). According to a study using fiber optic biosensors, SARS-CoV 2 may be identified in vivo. The suggested catheter-like probe is designed to eliminate the requirement for sample storage and other ancillary issues (Biswas 2021). Figure 4 shows the D614G mutation of SARS-CoV 2 and the surface spike (S) protein structure.
Spike protein and infection
The coronavirus infection cycle begins with receptor binding and membrane fusion, which are critical. SARS-CoV 2 is covered with spike proteins. The S protein facilitates viral attachment to ACE2 as a cellular receptor during virus entry. Then, a type 2 TM serine protease helps the virus enter the host cell. Polyproteins are formed when the virus’ RNA genome enters the cell and is translated into single-stranded RNA. The viral genome replication and transcription take place in the following stage by cleaving viral proteins and assembling the transcription complex (RTC). Ultimately, the host cell assembles the viral DNA into proteins, which are then packaged and released into the environment. In addition, virions from infected cells can infect new cells (Zandi et al. 2021; Boopathi et al. 2021; Hasan et al. 2021). However, several well-designed trials and promising results are required to ensure such a procedure. Indeed, investigating the virus’ propagation is critical for avoiding future outbreaks. The COVID-19 virus spreads faster than the SARS-CoV virus and quickly attacks those previously infected (Fu et al. 2020). Figure 5 illustrates the mechanism by which the SARS-CoV 2 spike protein enters the host cell.
Distribution mutations of SARS-CoV 2 spike
The spiking protein consists of an N-terminal S1 subunit and a C-terminal S2 subunit located near the membrane. The S1 subunit comprises the S1A, S1B, S1C, and S1D domains. The S1A domain, sometimes referred to as the N-terminal domain (NTD), detects carbohydrates such as sialic acid required for viral attachment to the host cell membrane. The SARS-CoV 2 spike protein’s S1B domain, commonly referred to as the receptor-binding domain (RBD), interacts with the human ACE-2 receptor (Zhang et al. 2020a; Wang et al. 2020a). Human SARS-CoV 2 spike proteins have been shown to contain mutations and changes at the glycosylation site (Kaushal et al. 2020). Research indicates that the D614G mutation is a proportionally more common mutation (Korber et al. 2020; Plante et al. 2021; Yang et al. 2021). Figure 6 shows the mutation density calculated as a function of the number of unique mutations found for each sequence length corresponding to the various regions of the spike protein as a function of the length of the sequence. In the spike protein, the protease cleavage site (between residues 675 and 692) is associated with the highest possible mutation density. Viruses that are proteolytically cleaved by a large number of host enzymes can benefit from changes that occur at this point in the spike protein during their evolutionary development. Furthermore, the NTD (S1A domain) is another part of the spike protein in which mutations have accumulated in a more significant number than the rest of the sequence of the spike protein. Despite the significant number of SARS-CoV 2 spike protein sequences, it is now accessible in the NCBI virus database to better understand the current spike protein mutation scenario (https://www.ncbi.nlm.nih.gov/labs/virus/vssi/).
AI enabled by photonics technologies
Artificial neural networks (ANN) that can mimic the structural, functional, and biological properties of human neural networks are urgently needed to meet the growing needs of brain research and artificial intelligence (Zhang et al. 2019). Nanophotonics is a potential technique for studying biological neural networks (BNNs) using optical imaging. The advancement of optical imaging, in particular the development of super-resolution optical far-field microscopy (awarded the Nobel Prize in Chemistry in 2014), has sparked interest in neural networks on the nanoscale (Hell and Wichmann 1994; Betzig et al. 2006). Photonic biosensor technologies have high precision, low-cost, ease of use, and ready-to-use mode. Thus, it can be necessary to rapidly detect harmful viruses like COVID-19 and give creative techniques to suppress emerging viral outbreaks (Lukose et al. 2021; Taha et al. 2020; Taha 2021; Samarrai et al. 2021; Serag and El-Zeftawy 2021). The development of intelligent systems such as biological neural networks (BNNs) continued shortly after the invention of the modern computer. Most research on artificial neural networks is carried out through computer software simulations. Von Neumann computers are used in most of this work. The idea of using electronic or photonic hardware to look like BNNs was first suggested in the late 1980s (Psaltis et al. 1990; Mead 1990). Light has unique characteristics such as maximum transmission speed, no mutual interference side effects, and optical signals can be time-multiplexed (Deng and Liu 2014). Integrated optical circuits on a chip provide an appropriate platform for high-compactness, high-stability ANNs. It can fabricate integrated lasers, photodetectors, and non-linear optical devices (Mesaritakis et al. 2016). It provides several advantages for establishing free space and waveguide interconnections, notably high bandwidth, low loss, and low crosstalk. Photodetectors can form photonic somas to transfer optoelectrical and electro-optical signals (Tait et al. 2017; Shen et al. 2017). In addition, modulators and light sources (LED and laser) include lasers, optical amplifiers, saturable absorbers. Optical switches such as holograms and Mach Zehnder interferometers (MZI) can perform the weighing function (Rosenbluth et al. 2009; Li and Cai 2010; Muhanad Fadhel et al. 2021). Artificial intelligence-based face mask classification algorithms have been developed to help manage the COVID-19 pandemic. It used a mask detector that evaluates whether or not a person is wearing a mask using a machine learning face classification algorithm. It can be connected to a surveillance system to guarantee that only mask-wearing individuals are admitted (Gupta et al. 2021). A smart sensor system based on machine learning techniques and data collection devices has also been designed. Computational sensing systems reduce the data load and at the same time improve sensing capabilities, allowing inexpensive and compact sensors (Ballard et al. 2021). On the other hand, there are many applications of photonics (Xin et al. 2020), such as optical fiber tweezers were devised for the trapping and manipulation of small objects ranging from dielectric particles (Constable et al. 1993; Zhao et al. 2020). Carbon nanotubes and metallic particles are implanted in biological targets such as single cells, viruses, and bacteria (Zhang et al. 2018; Li et al. 2015). Furthermore, evanescent fields around optical waveguides/subwavelength optical fibers allow long-distance propagation and detection of nanoparticles and microbes (Yu et al. 2018; Muda et al. 2018). The combination of fiber optic and fluidic forces in an optofluidic platform enables size-dependent sorting of nanoparticles and shape-selective bacterial sieving (Xin et al. 2013; Shi et al. 2018a). A strategy to search for bacterial binding agents such as antibodies is also used at the single-bacterial level (Shi et al. 2018b).
COVID-19 diseases monitoring system
The rural area of many developing nations lacks access to high-quality health services. Thus, it causes early death due to several undiagnosed and untreated conditions. Detecting infectious diseases early is critical to contain disease transmission through increased self-isolation and early treatment. Currently, most diagnostic techniques involve collecting nasal secretions, saliva, or blood, followed by a nucleic acid-based test to detect ongoing infections or blood-based serological detection for previous illnesses. For nucleic acid-based diagnostics, despite their remarkable sensitivity, samples may be required that are collected several days after exposure to obtain clear positive evidence (Sethuraman et al. 2020). In addition, they cannot be used cost-effectively over the long term and are hampered by the increasing scarcity of critical reagents. Disposable wearable devices are an accurate and widely used technology for determining individual baseline health indicators to detect significant deviations from baseline physiology at the onset of an infection (Li et al. 2017; Dunn et al. 2018). Given the above, we recommend the following: First, understanding the virus better is needed to identify, diagnose, and predict mutations. Second, monitoring and contact tracing help prevent or delay the transmission of the virus. Third, coping with health crises through individualized information and education. Fourth, assessing the recovery process and improving early warning systems. A specific study has shown that the risk of SARS-CoV 2 infection from a single point of contact is low. It is mentioned that repeated testing of such surfaces could give an early warning of impending breakouts (Harvey et al. 2021). In other studies, SARS-CoV 2 has been shown to be viable on surfaces for up to 28 days in laboratory tests with high initial viral loads and under optimal environmental conditions, with half-lives of hours to days on plastic and stainless steel (Kampf et al. 2020; Zou et al. 2020; Aboubakr et al. 2021). Although eye cells express the ACE2 receptor, these researchers believe that eye infection or tear-induced virus spread would be quite rare. It is hypothesized that tears would wash away the virus and that an immune response in the eyes led by antibodies and a protein called lactoferrin would prevent widespread infection (Liu and Sun 2020). Finally, in most COVID-19 vaccines, spike proteins stimulate antibody production, either made with an adjuvant or encoded in messenger RNA or DNA encoded in an adenoviral vector. The difficulty with this strategy is a potential problem. On the other hand, spike proteins are key to stopping the pandemic, and our approach has been summarized below.
-
Pillar 1: A. Photonics detection: Nano-sized scatter signals are weak due to their tiny size and optical wavelength, making it impossible to observe nano-sized objects with light methods directly. Various techniques have been employed to address this issue. Many papers have been published on bio-sensing for SARS-CoV-2, which uses direct immunoassays for amplification-free DNA or RNA biosensing (Pinheiro et al. 2021). Furthermore, many studies indicate single-molecule and particle detection on actual portable microscopy platforms such as a smartphone based on fluorescence (Wei et al. 2013), a 3D printed platform for free lens holographic imaging (McLeod et al. 2014), a smartphone based on fluorescence spectroscopy (Yu et al. 2014), a smartphone based on plasmonic nanohole array (Cetin et al. 2014), a smartphone based on dark-filed microscopy (Sun and Hu 2018), a smartphone based on surface-enhanced Raman spectroscopy with cloud network (Mu et al. 2019), homemade smartphone using total internal reflection microscopy (Varra et al. 2020), and multi-modal smartphone (bright-field, oblique illumination dark-field, and total internal reflection dark-field) on a single platform (Rabha et al. 2021). An optical sensor device connecting through the 5 G network allows the Internet of Things (IoT) to identify single nucleic acids SARS-CoV 2 (Guo et al. 2021). Microfluidics is a significant advantage for portable optical detection employing fluorescent tags. It allows for quick mixing, filtration, and other liquid sample preparation on tiny amounts in the field. Mobile microscopy and spectroscopy systems can be combined with optofluidic chips for assay methods (Yang et al. 2019). There are also optical waveguides method using spectroscopy (Wang et al. 2020b).
-
B. Optical band pass filter ( pinhole): pinhole collimators or practical diameter concept is used to image nanoparticles and tiny biological (Metzler et al. 2001). This study aimed to determine whether pinhole micro-single-photon emission computed tomography (PM-SPECT) can accurately diagnose certain medical conditions. The results show an exact localization and quantification of the viral infection and are able to replace more time-consuming and expensive analyses (Penheiter et al. 2012). Other than that, single-photon emission through a pinhole computed tomography are used to support the development of novel equipment for biological assessment (Wu et al. 2003). Pinhole collimator for imaging of tiny animals of different sizes was also successfully designed. The results indicated the high resolution and efficiency of the detection (Seng Peng et al. 2005). Multi-pinhole fluorescence x-ray computed tomography using a 2-D detector and full-field volumetric beam are used to expedite data acquisition and improve signal-to-noise ratios for molecular imaging projections. The system detected a concentration of 0.038 mg/ml in vivo imaging (Sasaya et al. 2017).
-
C. CMOS/APS technology: CMOS image sensors are used in various medical applications; CMOS imaging systems have attracted significant interest from companies and universities due to the growing demand for compact and low-power imaging systems. In addition, CMOS image sensors have better integration capabilities compared to charge-coupled devices (CCDs) (Campos 2011). DNA microarray detection based on CMOS technology is reported. The methodology used an integrated active-pixel sensor (APS) to detect a change in thickness when an invisible nanoparticle was at the same spot as precisely matched DNA samples. In addition, the platform sensitivity may be increased by using a longer silver deposition duration and can detect a low concentration of 10 pM (Wang et al. 2005). A study showed tested an Array Pixel Sensor (APS) to build a detector for transmission electron microscopy (TEM). The result was that the APS could make a short readout time, thereby providing data collection at a much higher rate than a standard CCD-based camera (Milazzo et al. 2005). Researchers present a neuromorphic active pixel image sensor array (NAPISA) chip using an oxide semiconductor that emulates human visual memory. The methods may successfully show the visual memory and forgetting characteristics utilizing the pulsed light stencil approach without any software or simulation. It will be helpful to other neuromorphic devices and systems for next-generation artificial intelligence technologies (Hong et al. 2021). A novel active-pixel sensor for electron counting in cryo-EM has successfully integrated the camera to acquire single-particle and tomographic cryo-EM data automatically. The results show that enabling the high-speed gathering of high signal noise ratio and ease of use of on-chip electron counting lowers the overall cost of this camera, making high-resolution cryo-EM equipment more affordable (Bammes and Bilhorn 2021).
-
Pillar 2: Connection: Fiber optic technologies enable remote detection and control of virus epidemics through surface/environmental testing and in vitro diagnosis of human diseases (Taha et al. 2021b). Our proposed photonic system, based on artificial intelligence to track and detect COVID-19 on surfaces, will assist medical teams by providing real-time monitoring and detection, improving the efficiency and accuracy of optical biosensors, and improving the quality of healthcare in public spaces such as Universities, Supermarkets Ports, and Food Quality Checks.
-
Pillar 3: Big data empowering: Expanding medical procedures and activities during the COVID-19 pandemic, including remote monitoring, remote screens, remote consultation, and remote management, can improve the estimation and control of SARS-CoV2 mutations. Optical fiber cables are suitable for extended transmission operations, data, and communication networks. Optical fiber is the transmission medium of choice for most backbone providers in developed countries due to its key advantages: high bandwidth, higher speed, high reliability, and security. Intelligent optical networks that collect vast amounts of data from detection lasers are the link between hospitals and endpoint locations. In addition, laser-based optical detection can help in the future with environmental monitoring, measuring virus concentrations in the air, and checking food quality. For example, data for COVID-19 tracking can be collected via an intelligent network of medical biosensors, which improves the quality of healthcare. Integrating telehealth services to meet COVID-19 requirements and validating healthcare technologies are critical steps to improve utility and complete the implementation of clinical virus assessment systems. Telemedicine technology is being discussed to track COVID-19 patients with mild symptoms who are sent home to recover, to monitor COVID-19 patients at home and at risk of deterioration due to disease progression. Telemedicine systems have reduced the risk of hospitalization delays due to the spread of the disease and helped patients self-assess. The telemedicine approach lowered the danger of late admission owing to illness spread and aided patients in self-evaluation. Patients can keep the multidisciplinary team informed via online telehealth forms (Xu et al. 2020). Developing a system to manage emergency dental treatment patients using recent research and previous experiences, the authors offered general information about the COVID-19 illness and recommendations for treating emergency dental operations to prevent cross-infections in the dental office. They also discussed their experience and the possibility of telemedicine for dental practitioners (Giudice et al. 2020). Telehealth was advised to screen suspicious patients to reduce exposure risks and maximize medical staff protection. This study devised a method to decrease COVID-19 exposure in the emergency room. First, in compliance with a telehealth screening policy, the patient examination room is equipped with an intercom and iPad tablets for communication. Second, through intercom or video conferencing, physicians are able to make visual assessment of patients (Chou et al. 2020). The possible dangers in teleconsultation services during the COVID-19 epidemic and how to avoid them are highlighted. They examined the future of telehealth in health care systems and proposed acceptable settings with improved documentation, adequate training, information exchange, and observation criteria to minimize remote consultation concerns (Iyengar et al. 2020). The assessed teleconsultation system was in plastic surgery clinics during the COVID-19 epidemic. Plastic surgeons received a survey to evaluate the efficacy, modality, safety, and usability of virtual consultations. The data suggested that teleconsultation was both time and cost-effective, allowing patients to continue receiving therapy (Sinha et al. 2021). Finally, some challenges include telehealth data privacy and security, a lack of integration of telehealth systems, telehealth infrastructure, tele examination equipment, and a clear path to use telehealth after COVID-19 (Negrini et al. 2020). Improving telehealth infrastructure and systems will help provide more excellent quality treatment and control to many patients. Our objective is to obtain direct detection of the SARS-CoV 2 virus without the requirement for sample preparation or molecular amplification techniques. Furthermore, by connecting via the 5G network, testing for COVID-19 becomes a part of the Internet of Healthcare Things as shown in Fig. 7.
Conclusions
The COVID-19 epidemic has spread rapidly around the world since 2019, and people’s lives are becoming more difficult and uncomfortable because of the epidemic. Numerous people have died as a result of this infection. One of the reasons the virus is spreading is the scarcity of antiviral drugs. This disease spreads constantly and quickly from infected people through common acts such as breathing, coughing, and sneezing. The main symptom is regular flu and loss of taste and smell. In summary, the integration of lighting technologies into artificial intelligence systems contributes to the three main benefits: the ability to identify and isolate infected individuals, track viral mutations by detecting genetic sequencing changes, and monitor contaminated surfaces. In addition, photonic technology can enable the capture and transmission of big data, where data can be transmitted from anywhere. As a result, future biological applications will benefit from the use of photonics. Photonics has already made significant advances in biological research from in vitro to in vivo. It is forecast that optical technology will continue to have many new positive effects in various fields in the coming years.
Data availability
Data availability is available on GISAID: (https://www.gisaid.org/) and SARS-CoV-2 spike protein sequences, which are now accessible in the NCBI virus database to better understand the current spike protein mutation scenario: https://www.ncbi.nlm.nih.gov/labs/virus/vssi/).
References
Aboubakr HA, Sharafeldin TA, Goyal SM (2021) Stability of SARS-CoV-2 and other coronaviruses in the environment and on common touch surfaces and the influence of climatic conditions: a review. Transbound Emerg Dis 68(2):296–312. https://doi.org/10.1111/tbed.13707
Almubaid Z, Al-Mubaid H (2021) Analysis and comparison of genetic variants and mutations of the novel coronavirus SARS-CoV-2. Gene Rep 23:101064
Ballard Z, Brown C, Madni AM, Ozcan A (2021) Machine learning and computation-enabled intelligent sensor design. Nat Mach Intell 3(7):556–565. https://doi.org/10.1038/s42256-021-00360-9
Bammes B, Bilhorn R (2021) A novel event-based active pixel sensor for cryo-EM electron counting. Microsc Microanal 27(S1):1334–1336. https://doi.org/10.1017/S1431927621004979
Betzig E, Patterson GH, Sougrat R, Lindwasser OW, Olenych S, Bonifacino JS, Davidson MW, Lippincott-Schwartz J, Hess HF (2006) Imaging intracellular fluorescent proteins at nanometer resolution. Science 313(5793):1642–1645. https://doi.org/10.1126/science.1127344
Biswas R (2021) Catheter like in vivo fiber optic probe for rapid diagnosis of SARS-CoV-2. Results Opt 5:100180. https://doi.org/10.1016/j.rio.2021.100180
Boopathi S, Poma AB, Kolandaivel P (2021) Novel 2019 coronavirus structure, mechanism of action, antiviral drug promises and rule out against its treatment. J Biomol Struct Dyn 39(9):3409–3418. https://doi.org/10.1080/07391102.2020.1758788
Callaway E (2021) Heavily mutated Omicron variant puts scientists on alert. Nat Mag 2021
Campos FDS (2011) Active pixel sensor CMOS operating multi-sampled in time domain Photodiodes-World Activities in 2011. IntechOpen
Cetin AE, Coskun AF, Galarreta BC, Huang M, Herman D, Ozcan A, Altug H (2014) Handheld high-throughput plasmonic biosensor using computational on-chip imaging. Light Sci Appl 3(1):e122–e122. https://doi.org/10.1038/lsa.2014.3
Chou E, Hsieh Y-L, Wolfshohl J, Green F, Bhakta T (2020) Onsite telemedicine strategy for coronavirus (COVID-19) screening to limit exposure in ED. Emerg Med J 37(6):335. https://doi.org/10.1136/emermed-2020-209645
Consortium WST (2021) Repurposed antiviral drugs for COVID-19: interim results of the WHO SOLIDARITY trial. N Engl J Med 384(6):497–511
Constable A, Kim J, Mervis J, Zarinetchi F, Prentiss M (1993) Demonstration of a fiber-optical light-force trap. Opt Lett 18(21):1867–1869. https://doi.org/10.1364/OL.18.001867
Davies NG, Abbott S, Barnard RC, Jarvis CI, Kucharski AJ, Munday JD, Pearson CAB, Russell TW, Tully DC, Washburne AD, Wenseleers T, Gimma A, Waites W, Wong KLM, van Zandvoort K, Silverman JD, CMMID COVID-19 Working Group; COVID-19 Genomics UK (COG-UK) Consortium, Diaz-Ordaz K, Keogh R, Eggo RM, Funk S, Jit M, Atkins KE, Edmunds WJ (2021) Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372(6538):3055. https://doi.org/10.1126/science.abg3055
Dearlove B, Lewitus E, Bai H, Li Y, Reeves DB, Joyce MG, Scott PT, Amare MF, Vasan S, Michael NL, Modjarrad K, Rolland M (2020) A SARS-CoV-2 vaccine candidate would likely match all currently circulating variants. Proc Natl Acad Sci 117(38):23652. https://doi.org/10.1073/pnas.2008281117
Deng R, Liu X (2014) Tunable lifetime nanocrystals. Nat Photon 8(1):10–12. https://doi.org/10.1038/nphoton.2013.353
Dunn J, Runge R, Snyder M (2018) Wearables and the medical revolution. J Pers Med 15(5):429–448. https://doi.org/10.2217/pme-2018-0044
Fu L, Wang B, Yuan T, Chen X, Ao Y, Fitzpatrick T, Li P, Zhou Y, Lin Y-f, Duan Q, Luo G, Fan S, Lu Y, Feng A, Zhan Y, Liang B, Cai W, Zhang L, Du X, Li L, Shu Y, Zou H (2020) Clinical characteristics of coronavirus disease 2019 (COVID-19) in China: a systematic review and meta-analysis. J Infect 80(6):656–665. https://doi.org/10.1016/j.jinf.2020.03.041
Gao S-J, Guo H, Luo G (2021) Omicron variant (B.1.1.529) of SARS-CoV-2, a global urgent public health alert! J Med Virol. https://doi.org/10.1002/jmv.27491
Gazali FM, Nuhamunada M, Nabilla R, Supriyati E, Hakim MS, Arguni E, Daniwijaya EW, Nuryastuti T, Haryana SM, Wibawa T, Wijayanti N (2021) Detection of SARS-CoV-2 spike protein D614G mutation by qPCR-HRM analysis. Heliyon 7(9):e07936. https://doi.org/10.1016/j.heliyon.2021.e07936
Giudice A, Bennardo F, Antonelli A, Barone S, Fortunato L (2020) COVID-19 is a new challenge for dental practitioners: advice on patients’ management from prevention of cross infections to telemedicine. Open Dent J 14(1)
Grubaugh ND, Hanage WP, Rasmussen AL (2020) Making sense of mutation: what D614G means for the COVID-19 pandemic remains unclear. Cell 182(4):794–795. https://doi.org/10.1016/j.cell.2020.06.040
Gu H, Krishnan P, Ng DY, Chang LD, Liu GY, Cheng SS, Hui MM, Fan MC, Wan JH, Lau LH (2021) Probable transmission of SARS-CoV-2 omicron variant in quarantine hotel, Hong Kong, China, November 2021. Emerg Infect Dis 28(2)
Guo J, Chen S, Tian S, Liu K, Ni J, Zhao M, Kang Y, Ma X, Guo J (2021) 5G-enabled ultra-sensitive fluorescence sensor for proactive prognosis of COVID-19. Biosens Bioelectron 181:113160. https://doi.org/10.1016/j.bios.2021.113160
Gupta S, Sreenivasu SVN, Chouhan K, Shrivastava A, Sahu B, Manohar Potdar R (2021) Novel face mask detection technique using machine learning to control COVID’19 pandemic. Mater Today: Proc. https://doi.org/10.1016/j.matpr.2021.07.368
Harvey AP, Fuhrmeister ER, Cantrell ME, Pitol AK, Swarthout JM, Powers JE, Nadimpalli ML, Julian TR, Pickering AJ (2021) Longitudinal monitoring of SARS-CoV-2 RNA on high-touch surfaces in a community setting. Environ Sci Technol Lett 8(2):168–175. https://doi.org/10.1021/acs.estlett.0c00875
Hasan A, Paray BA, Hussain A, Qadir FA, Attar F, Aziz FM, Sharifi M, Derakhshankhah H, Rasti B, Mehrabi M, Shahpasand K, Saboury AA, Falahati M (2021) A review on the cleavage priming of the spike protein on coronavirus by angiotensin-converting enzyme-2 and furin. J Biomol Struct Dyn 39(8):3025–3033. https://doi.org/10.1080/07391102.2020.1754293
Hell SW, Wichmann J (1994) Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt Lett 19(11):780–782. https://doi.org/10.1364/OL.19.000780
Hoffmann M, Arora P, Groß R, Seidel A, Hörnich BF, Hahn AS, Krüger N, Graichen L, Hofmann-Winkler H, Kempf A, Winkler MS, Schulz S, Jäck H-M, Jahrsdörfer B, Schrezenmeier H, Müller M, Kleger A, Münch J, Pöhlmann S (2021) SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 184(9):2384-2393.e12. https://doi.org/10.1016/j.cell.2021.03.036
Hong S, Cho H, Kang BH, Park K, Akinwande D, Kim HJ, Kim S (2021) Neuromorphic active pixel image sensor array for visual memory. ACS Nano 15(9):15362–15370. https://doi.org/10.1021/acsnano.1c06758
Iyengar K, Jain VK, Vaishya R (2020) Pitfalls in telemedicine consultations in the era of COVID 19 and how to avoid them. Diabetes Metab Syndr 14(5):797–799. https://doi.org/10.1016/j.dsx.2020.06.007
Jakhmola S, Indari O, Kashyap D, Varshney N, Das A, Manivannan E, Jha HC (2021) Mutational analysis of structural proteins of SARS-CoV-2. Heliyon 7(3):e06572. https://doi.org/10.1016/j.heliyon.2021.e06572
Kampf G, Todt D, Pfaender S, Steinmann E (2020) Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J Hosp Infect 104(3):246–251. https://doi.org/10.1016/j.jhin.2020.01.022
Kannan SR, Spratt AN, Cohen AR, Naqvi SH, Chand HS, Quinn TP, Lorson CL, Byrareddy SN, Singh K (2021) Evolutionary analysis of the Delta and Delta Plus variants of the SARS-CoV-2 viruses. J Autoimmun 124:102715. https://doi.org/10.1016/j.jaut.2021.102715
Kaushal N, Gupta Y, Goyal M, Khaiboullina SF, Baranwal M, Verma SC (2020) Mutational frequencies of SARS-CoV-2 genome during the beginning months of the outbreak in USA. Pathogens 9(7):565
Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, Hengartner N, Giorgi EE, Bhattacharya T, Foley B, Hastie KM, Parker MD, Partridge DG, Evans CM, Freeman TM, de Silva TI, Angyal A, Brown RL, Carrilero L, Green LR, Groves DC, Johnson KJ, Keeley AJ, Lindsey BB, Parsons PJ, Raza M, Rowland-Jones S, Smith N, Tucker RM, Wang D, Wyles MD, McDanal C, Perez LG, Tang H, Moon-Walker A, Whelan SP, LaBranche CC, Saphire EO, Montefiori DC (2020) Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182(4):812-827.e19. https://doi.org/10.1016/j.cell.2020.06.043
Kottier SA, Cavanagh D, Britton P (1995) Experimental evidence of recombination in coronavirus infectious bronchitis virus. Virol 213(2):569–580. https://doi.org/10.1006/viro.1995.0029
Krammer F (2020) SARS-CoV-2 vaccines in development. Nature 586(7830):516–527. https://doi.org/10.1038/s41586-020-2798-3
Lauring AS, Hodcroft EB (2021) Genetic variants of SARS-CoV-2—what do they mean? JAMA 325(6):529–531. https://doi.org/10.1001/jama.2020.27124
Li S, Cai X (2010) High-contrast all optical bistable switching in coupled nonlinear photonic crystal microcavities. Appl Phys Lett 96(13):131114. https://doi.org/10.1063/1.3378812
Li Y, Xin H, Liu X, Li B (2015) Non-contact intracellular binding of chloroplasts in vivo. Sci Rep 5(1):10925. https://doi.org/10.1038/srep10925
Li X, Dunn J, Salins D, Zhou G, Zhou W, Schüssler-Fiorenza Rose SM, Perelman D, Colbert E, Runge R, Rego S (2017) Digital health: tracking physiomes and activity using wearable biosensors reveals useful health-related information. PLoS Biol 15(1):e2001402
Liu Z, Sun C-B (2020) Conjunctiva is not a preferred gateway of entry for SARS-CoV-2 to infect respiratory tract. J Med Virol 92(9):1410–1412. https://doi.org/10.1002/jmv.25859
Loo K-Y, Letchumanan V, Ser H-L, Teoh SL, Law JW-F, Tan LT-H, Ab Mutalib N-S, Chan K-G, Lee L-H (2021) COVID-19: insights into potential vaccines. Microorganisms 9(3):605
Lukose J, Chidangil S, George SD (2021) Optical technologies for the detection of viruses like COVID-19: Progress and prospects. Biosens Bioelectron 178:113004. https://doi.org/10.1016/j.bios.2021.113004
McLeod E, Nguyen C, Huang P, Luo W, Veli M, Ozcan A (2014) Tunable vapor-condensed nanolenses. ACS Nano 8(7):7340–7349. https://doi.org/10.1021/nn502453h
Mead C (1990) Neuromorphic electronic systems. Proc IEEE 78(10):1629–1636. https://doi.org/10.1109/5.58356
Mesaritakis C, Kapsalis A, Bogris A, Syvridis D (2016) Artificial neuron based on integrated semiconductor quantum dot mode-locked lasers. Sci Rep 6(1):39317. https://doi.org/10.1038/srep39317
Metzler SD, Bowsher JE, Smith MF, Jaszczak RJ (2001) Analytic determination of pinhole collimator sensitivity with penetration. IEEE Trans Med Imaging 20(8):730–741. https://doi.org/10.1109/42.938241
Milazzo A-C, Leblanc P, Duttweiler F, Jin L, Bouwer JC, Peltier S, Ellisman M, Bieser F, Matis HS, Wieman H, Denes P, Kleinfelder S, Xuong N-H (2005) Active pixel sensor array as a detector for electron microscopy. Ultramicroscopy 104(2):152–159. https://doi.org/10.1016/j.ultramic.2005.03.006
Mu T, Li S, Feng H, Zhang C, Wang B, Ma X, Guo J, Huang B, Zhu L (2019) High-sensitive smartphone-based Raman system based on cloud network architecture. IEEE J Sel Top Quantum Electron 25(1):1–6. https://doi.org/10.1109/JSTQE.2018.2832661
Muda R, Arsad N, Ariffin N, Hadi A (2018) A compact optical fiber based gas sensing system for carbon dioxide monitoring. J Telecommun Electron Comput Eng 10(1–3):11–14
Muhanad Fadhel M, Rashid H, Essa Hamzah A, Dzulkefly Zan MS, Abd Aziz N, Arsad N (2021) Flat frequency comb generation employing cascaded single-drive Mach-Zehnder modulators with a simple analogue driving signal. J Mod Opt 68(10):536–541. https://doi.org/10.1080/09500340.2021.1925764
Negrini S, Kiekens C, Bernetti A, Capecci M, Ceravolo MG, Lavezzi S, Zampolini M, Boldrini P (2020) Telemedicine from research to practice during the pandemic. “Instant paper from the field” on rehabilitation answers to the COVID-19 emergency. Eur J Phys Rehabil Med 56(3):327–330. https://doi.org/10.23736/s1973-9087.20.06331-5
Ostrov DA (2021) Structural consequences of variation in SARS-CoV-2 B. 1.1. 7. J Cell Immunol 3(2):103
Penheiter AR, Griesmann GE, Federspiel MJ, Dingli D, Russell SJ, Carlson SK (2012) Pinhole micro-SPECT/CT for noninvasive monitoring and quantitation of oncolytic virus dispersion and percent infection in solid tumors. Gene Ther 19(3):279–287. https://doi.org/10.1038/gt.2011.107
Pinheiro T, Cardoso AR, Sousa CEA, Marques AC, Tavares APM, Matos AM, Cruz MT, Moreira FTC, Martins R, Fortunato E, Sales MGF (2021) Paper-based biosensors for COVID-19: a review of innovative tools for controlling the pandemic. ACS Omega 6(44):29268–29290. https://doi.org/10.1021/acsomega.1c04012
Plante JA, Liu Y, Liu J, Xia H, Johnson BA, Lokugamage KG, Zhang X, Muruato AE, Zou J, Fontes-Garfias CR, Mirchandani D, Scharton D, Bilello JP, Ku Z, An Z, Kalveram B, Freiberg AN, Menachery VD, Xie X, Plante KS, Weaver SC, Shi P-Y (2021) Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592(7852):116–121. https://doi.org/10.1038/s41586-020-2895-3
Psaltis D, Brady D, Gu X-G, Lin S (1990) Holography in artificial neural networks. Nature 343(6256):325–330. https://doi.org/10.1038/343325a0
Rabha D, Biswas S, Chamuah N, Mandal M, Nath P (2021) Wide-field multi-modal microscopic imaging using smartphone. Opt Lasers Eng 137:106343. https://doi.org/10.1016/j.optlaseng.2020.106343
Rao S, Singh M (2021) The newly detected B.1.1.529 (Omicron) variant of SARS-CoV-2 with multiple mutations. DHR Proc 1:7–10
Rosenbluth D, Kravtsov K, Fok MP, Prucnal PR (2009) A high performance photonic pulse processing device. Opt Express 17(25):22767–22772. https://doi.org/10.1364/OE.17.022767
Samarasekera U (2021) India grapples with second wave of COVID-19. Lancet Microbe 2(6):e238. https://doi.org/10.1016/S2666-5247(21)00123-3
Samarrai R, Riccardi AC, Tessema B, Setzen M, Brown SM (2021) Continuation of telemedicine in otolaryngology post-COVID-19: applications by subspecialty. Am J Otolaryngol 42(3):102928. https://doi.org/10.1016/j.amjoto.2021.102928
Sanjuán R, Domingo-Calap P (2016) Mechanisms of viral mutation. Cell Mol Life Sci 73(23):4433–4448. https://doi.org/10.1007/s00018-016-2299-6
Sasaya T, Sunaguchi N, Hyodo K, Zeniya T, Yuasa T (2017) Multi-pinhole fluorescent x-ray computed tomography for molecular imaging. Sci Rep 7(1):5742. https://doi.org/10.1038/s41598-017-05179-2
Seng Peng M, Yuchuan W, Tsui BMW (2005) Design of a novel pinhole collimator system for SPECT imaging of small animals with different sizes. In: IEEE Nucl Sci Symp Conf Rec, 23–29 Oct. 2005. vol 5. p 2649–2652
Serag E, El-Zeftawy M (2021) Environmental aspect and applications of nanotechnology to eliminate COVID-19 epidemiology risk. Nanotechnol Environ Eng 6(1):11. https://doi.org/10.1007/s41204-021-00108-1
Sethuraman N, Jeremiah SS, Ryo A (2020) Interpreting diagnostic tests for SARS-CoV-2. JAMA 323(22):2249–2251. https://doi.org/10.1001/jama.2020.8259
Shen Y, Harris NC, Skirlo S, Prabhu M, Baehr-Jones T, Hochberg M, Sun X, Zhao S, Larochelle H, Englund D, Soljačić M (2017) Deep learning with coherent nanophotonic circuits. Nat Photon 11(7):441–446. https://doi.org/10.1038/nphoton.2017.93
Shi Y, Xiong S, Chin Lip K, Zhang J, Ser W, Wu J, Chen T, Yang Z, Hao Y, Liedberg B, Yap Peng H, Tsai Din P, Qiu C-W, Liu Ai Q (2018a) Nanometer-precision linear sorting with synchronized optofluidic dual barriers. Sci Adv 4(1):eaao0773. https://doi.org/10.1126/sciadv.aao0773
Shi YZ, Xiong S, Zhang Y, Chin LK, Chen YY, Zhang JB, Zhang TH, Ser W, Larrson A, Lim SH, Wu JH, Chen TN, Yang ZC, Hao YL, Liedberg B, Yap PH, Wang K, Tsai DP, Qiu CW, Liu AQ (2018b) Sculpting nanoparticle dynamics for single-bacteria-level screening and direct binding-efficiency measurement. Nat Commun 9(1):815. https://doi.org/10.1038/s41467-018-03156-5
Sinha V, Malik M, Nugent N, Drake P, Cavale N (2021) The role of virtual consultations in plastic surgery during COVID-19 lockdown. Aesthetic Plast Surg 45(2):777–783. https://doi.org/10.1007/s00266-020-01932-7
Srinivasan S, Cui H, Gao Z, Liu M, Lu S, Mkandawire W, Narykov O, Sun M, Korkin D (2020) Structural genomics of SARS-CoV-2 indicates evolutionary conserved functional regions of viral proteins. Viruses 12(4):360
Sun D, Hu TY (2018) A low cost mobile phone dark-field microscope for nanoparticle-based quantitative studies. Biosens Bioelectron 99:513–518. https://doi.org/10.1016/j.bios.2017.08.025
Taha BA (2021) Perspectives of photonics technology to diagnosis COVID–19 viruses: a short review. JASN 1(1):1–6. https://doi.org/10.53293/jasn.2021.11016
Taha BA, Al Mashhadany Y, Hafiz Mokhtar MH, Dzulkefly Bin Zan MS, Arsad N (2020) An analysis review of detection coronavirus disease 2019 (COVID-19) based on biosensor application. Sensors 20(23):6764
Taha BA, Al Mashhadany Y, Bachok NN, Ashrif A, Bakar A, Hafiz Mokhtar MH, Dzulkefly Bin Zan MS, Arsad N (2021a) Detection of COVID-19 virus on surfaces using photonics: challenges and perspectives. Diagnostics 11(6):1119
Taha BA, Ali N, Sapiee NM, Fadhel MM, Mat Yeh RM, Bachok NN, Al Mashhadany Y, Arsad N (2021b) Comprehensive review tapered optical fiber configurations for sensing application: trend and challenges. Biosensors 11(8):253
Tait AN, de Lima TF, Zhou E, Wu AX, Nahmias MA, Shastri BJ, Prucnal PR (2017) Neuromorphic photonic networks using silicon photonic weight banks. Sci Rep 7(1):7430. https://doi.org/10.1038/s41598-017-07754-z
Tang JW, Tambyah PA, Hui DSC (2021) Emergence of a new SARS-CoV-2 variant in the UK. J Infect 82(4):e27–e28. https://doi.org/10.1016/j.jinf.2020.12.024
Teng X, Li Q, Li Z, Zhang Y, Niu G, Xiao J, Yu J, Zhang Z, Song S (2020) Compositional variability and mutation spectra of monophyletic SARS-CoV-2 clades. Genomics Proteomics Bioinformatics 18(6):648–663. https://doi.org/10.1016/j.gpb.2020.10.003
Thye AY-K, Law JW-F, Pusparajah P, Letchumanan V, Chan K-G, Lee L-H (2021) Emerging SARS-CoV-2 variants of concern (VOCs): an impending global crisis. Biomedicines 9(10):1303
Unlu S, Uskudar-Guclu A, Basustaoglu A (2021) Predominant mutations of SARS-CoV-2: their geographical distribution and potential consequences. Mediterr J Infect Microbe Antimicrob. https://doi.org/10.4274/mjima.galenos.2021.2021.15
Varra T, Simpson A, Roesler B, Nilsson Z, Ryan D, Van Erdewyk M, Schuttlefield Christus JD, Sambur JB (2020) A homemade smart phone microscope for single-particle fluorescence microscopy. J Chem Educ 97(2):471–478. https://doi.org/10.1021/acs.jchemed.9b00670
Volz E, Hill V, McCrone JT, Price A, Jorgensen D, O’Toole Á, Southgate J, Johnson R, Jackson B, Nascimento FF, Rey SM, Nicholls SM, Colquhoun RM, da Silva FA, Shepherd J, Pascall DJ, Shah R, Jesudason N, Li K, Jarrett R, Pacchiarini N, Bull M, Geidelberg L, Siveroni I, Koshy C, Wise E, Cortes N, Lynch J, Kidd S, Mori M, Fairley DJ, Curran T, McKenna JP, Adams H, Fraser C, Golubchik T, Bonsall D, Moore C, Caddy SL, Khokhar FA, Wantoch M, Reynolds N, Warne B, Maksimovic J, Spellman K, McCluggage K, John M, Beer R, Afifi S, Morgan S, Marchbank A, Price A, Kitchen C, Gulliver H, Merrick I, Southgate J, Guest M, Munn R, Workman T, Connor TR, Fuller W, Bresner C, Snell LB, Charalampous T, Nebbia G, Batra R, Edgeworth J, Robson SC, Beckett A, Loveson KF, Aanensen DM, Underwood AP, Yeats CA, Abudahab K, Taylor BEW, Menegazzo M, Clark G, Smith W, Khakh M, Fleming VM, Lister MM, Howson-Wells HC, Berry L, Boswell T, Joseph A, Willingham I, Bird P, Helmer T, Fallon K, Holmes C, Tang J, Raviprakash V, Campbell S, Sheriff N, Loose MW, Holmes N, Moore C, Carlile M, Wright V, Sang F, Debebe J, Coll F, Signell AW, Betancor G, Wilson HD, Feltwell T, Houldcroft CJ, Eldirdiri S, Kenyon A, Davis T, Pybus O, du Plessis L, Zarebski A, Raghwani J, Kraemer M, Francois S, Attwood S, Vasylyeva T, Torok ME, Hamilton WL, Goodfellow IG, Hall G, Jahun AS, Chaudhry Y, Hosmillo M, Pinckert ML, Georgana I, Yakovleva A, Meredith LW, Moses S, Lowe H, Ryan F, Fisher CL, Awan AR, Boyes J, Breuer J, Harris KA, Brown JR, Shah D, Atkinson L, Lee JCD, Alcolea-Medina A, Moore N, Cortes N, Williams R, Chapman MR, Levett LJ, Heaney J, Smith DL, Bashton M, Young GR, Allan J, Loh J, Randell PA, Cox A, Madona P, Holmes A, Bolt F, Price J, Mookerjee S, Rowan A, Taylor GP, Ragonnet-Cronin M, Nascimento FF, Jorgensen D, Siveroni I, Johnson R, Boyd O, Geidelberg L, Volz EM, Brunker K, Smollett KL, Loman NJ, Quick J, McMurray C, Stockton J, Nicholls S, Rowe W, Poplawski R, Martinez-Nunez RT, Mason J, Robinson TI, O’Toole E, Watts J, Breen C, Cowell A, Ludden C, Sluga G, Machin NW, Ahmad SSY, George RP, Halstead F, Sivaprakasam V, Thomson EC, Shepherd JG, Asamaphan P, Niebel MO, Li KK, Shah RN, Jesudason NG, Parr YA, Tong L, Broos A, Mair D, Nichols J, Carmichael SN, Nomikou K, Aranday-Cortes E, Johnson N, Starinskij I, da Silva FA, Robertson DL, Orton RJ, Hughes J, Vattipally S, Singer JB, Hale AD, Macfarlane-Smith LR, Harper KL, Taha Y, Payne BAI, Burton-Fanning S, Waugh S, Collins J, Eltringham G, Templeton KE, McHugh MP, Dewar R, Wastenge E, Dervisevic S, Stanley R, Prakash R, Stuart C, Elumogo N, Sethi DK, Meader EJ, Coupland LJ, Potter W, Graham C, Barton E, Padgett D, Scott G, Swindells E, Greenaway J, Nelson A, Yew WC, Resende Silva PC, Andersson M, Shaw R, Peto T, Justice A, Eyre D, Crooke D, Hoosdally S, Sloan TJ, Duckworth N, Walsh S, Chauhan AJ, Glaysher S, Bicknell K, Wyllie S, Butcher E, Elliott S, Lloyd A, Impey R, Levene N, Monaghan L, Bradley DT, Allara E, Pearson C, Muir P, Vipond IB, Hopes R, Pymont HM, Hutchings S, Curran MD, Parmar S, Lackenby A, Mbisa T, Platt S, Miah S, Bibby D, Manso C, Hubb J, Chand M, Dabrera G, Ramsay M, Bradshaw D, Thornton A, Myers R, Schaefer U, Groves N, Gallagher E, Lee D, Williams D, Ellaby N, Harrison I, Hartman H, Manesis N, Patel V, Bishop C, Chalker V, Osman H, Bosworth A, Robinson E, Holden MTG, Shaaban S, Birchley A, Adams A, Davies A, Gaskin A, Plimmer A, Gatica-Wilcox B, McKerr C, Moore C, Williams C, Heyburn D, De Lacy E, Hilvers E, Downing F, Shankar G, Jones H, Asad H, Coombes J, Watkins J, Evans JM, Fina L, Gifford L, Gilbert L, Graham L, Perry M, Morgan M, Bull M, Cronin M, Pacchiarini N, Craine N, Jones R, Howe R, Corden S, Rey S, Kumziene-Summerhayes S, Taylor S, Cottrell S, Jones S, Edwards S, O’Grady J, Page AJ, Wain J, Webber MA, Mather AE, Baker DJ, Rudder S, Yasir M, Thomson NM, Aydin A, Tedim AP, Kay GL, Trotter AJ, Gilroy RAJ, Alikhan N-F, de Oliveira ML, Le-Viet T, Meadows L, Kolyva A, Diaz M, Bell A, Gutierrez AV, Charles IG, Adriaenssens EM, Kingsley RA, Casey A, Simpson DA, Molnar Z, Thompson T, Acheson E, Masoli JAH, Knight BA, Hattersley A, Ellard S, Auckland C, Mahungu TW, Irish-Tavares D, Haque T, Bourgeois Y, Scarlett GP, Partridge DG, Raza M, Evans C, Johnson K, Liggett S, Baker P, Essex S, Lyons RA, Caller LG, Castellano S, Williams RJ, Kristiansen M, Roy S, Williams CA, Dyal PL, Tutill HJ, Panchbhaya YN, Forrest LM, Niola P, Findlay J, Brooks TT, Gavriil A, Mestek-Boukhibar L, Weeks S, Pandey S, Berry L, Jones K, Richter A, Beggs A, Smith CP, Bucca G, Hesketh AR, Harrison EM, Peacock SJ, Palmer S, Churcher CM, Bellis KL, Girgis ST, Naydenova P, Blane B, Sridhar S, Ruis C, Forrest S, Cormie C, Gill HK, Dias J, Higginson EE, Maes M, Young J, Kermack LM, Hadjirin NF, Aggarwal D, Griffith L, Swingler T, Davidson RK, Rambaut A, Williams T, Balcazar CE, Gallagher MD, O’Toole Á, Rooke S, Jackson B, Colquhoun R, Ashworth J, Hill V, McCrone JT, Scher E, Yu X, Williamson KA, Stanton TD, Michell SL, Bewshea CM, Temperton B, Michelsen ML, Warwick-Dugdale J, Manley R, Farbos A, Harrison JW, Sambles CM, Studholme DJ, Jeffries AR, Darby AC, Hiscox JA, Paterson S, Iturriza-Gomara M, Jackson KA, Lucaci AO, Vamos EE, Hughes M, Rainbow L, Eccles R, Nelson C, Whitehead M, Turtle L, Haldenby ST, Gregory R, Gemmell M, Kwiatkowski D, de Silva TI, Smith N, Angyal A, Lindsey BB, Groves DC, Green LR, Wang D, Freeman TM, Parker MD, Keeley AJ, Parsons PJ, Tucker RM, Brown R, Wyles M, Constantinidou C, Unnikrishnan M, Ott S, Cheng JKJ, Bridgewater HE, Frost LR, Taylor-Joyce G, Stark R, Baxter L, Alam MT, Brown PE, McClure PC, Chappell JG, Tsoleridis T, Ball J, Gramatopoulos D, Buck D, Todd JA, Green A, Trebes A, MacIntyre-Cockett G, de Cesare M, Langford C, Alderton A, Amato R, Goncalves S, Jackson DK, Johnston I, Sillitoe J, Palmer S, Lawniczak M, Berriman M, Danesh J, Livett R, Shirley L, Farr B, Quail M, Thurston S, Park N, Betteridge E, Weldon D, Goodwin S, Nelson R, Beaver C, Letchford L, Jackson DA, Foulser L, McMinn L, Prestwood L, Kay S, Kane L, Dorman MJ, Martincorena I, Puethe C, Keatley J-P, Tonkin-Hill G, Smith C, Jamrozy D, Beale MA, Patel M, Ariani C, Spencer-Chapman M, Drury E, Lo S, Rajatileka S, Scott C, James K, Buddenborg SK, Berger DJ, Patel G, Garcia-Casado MV, Dibling T, McGuigan S, Rogers HA, Hunter AD, Souster E, Neaverson AS, Goodfellow I, Loman NJ, Pybus OG, Robertson DL, Thomson EC, Rambaut A, Connor TR (2021) Evaluating the effects of SARS-CoV-2 spike mutation D614G on transmissibility and pathogenicity. Cell 184(1):64-75.e11. https://doi.org/10.1016/j.cell.2020.11.020
Wan KS, Tok PSK, Ratnam KKY, Aziz N, Isahak M, Zaki RA, Farid NDN, Hairi NN, Rampal S, Ng CW, Samsudin MF, Venugopal V, Asyraf M, Damanhuri NH, Doraimuthu S, Arumugam CT, Marthammuthu T, Nawawi FA, Baharudin F, Chong DWQ, Jayaraj VJ, Magarita V, Ponnampalavanar S, Hasnan N, Kamarulzaman A, Said MA (2021) Implementation of a COVID-19 surveillance programme for healthcare workers in a teaching hospital in an upper-middle-income country. PLoS ONE 16:1–15. https://doi.org/10.1371/journal.pone.0249394
Wang Y, Xu C, Li J, Hsing IM, Chan M (2005) A CMOS active pixel sensor based DNA micro-array with nano-metallic particles detection protocol. Solid State Electron 49(12):1933–1936. https://doi.org/10.1016/j.sse.2005.09.015
Wang Q, Zhang Y, Wu L, Niu S, Song C, Zhang Z, Lu G, Qiao C, Hu Y, Yuen K-Y, Wang Q, Zhou H, Yan J, Qi J (2020a) Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 181(4):894-904.e9. https://doi.org/10.1016/j.cell.2020.03.045
Wang R, Vemulapati S, Westblade LF, Glesby MJ, Mehta S, Erickson D (2020b) cAST: capillary-based platform for real-time phenotypic antimicrobial susceptibility testing. Anal Chem 92(3):2731–2738. https://doi.org/10.1021/acs.analchem.9b04991
Watanabe Y, Berndsen ZT, Raghwani J, Seabright GE, Allen JD, Pybus OG, McLellan JS, Wilson IA, Bowden TA, Ward AB, Crispin M (2020) Vulnerabilities in coronavirus glycan shields despite extensive glycosylation. Nat Commun 11(1):2688. https://doi.org/10.1038/s41467-020-16567-0
Wei Q, Qi H, Luo W, Tseng D, Ki SJ, Wan Z, Göröcs Z, Bentolila LA, Wu T-T, Sun R, Ozcan A (2013) Fluorescent imaging of single nanoparticles and viruses on a smart phone. ACS Nano 7(10):9147–9155. https://doi.org/10.1021/nn4037706
Weissman D, Alameh M-G, de Silva T, Collini P, Hornsby H, Brown R, LaBranche CC, Edwards RJ, Sutherland L, Santra S, Mansouri K, Gobeil S, McDanal C, Pardi N, Hengartner N, Lin PJC, Tam Y, Shaw PA, Lewis MG, Boesler C, Şahin U, Acharya P, Haynes BF, Korber B, Montefiori DC (2020) D614G spike mutation increases SARS CoV-2 susceptibility to neutralization. Cell Host Microbe:2020.07.22.20159905. https://doi.org/10.1101/2020.07.22.20159905
Wu MC, Gao D-W, Sievers RE, Lee RJ, Hasegawa BH, Dae MW (2003) Pinhole single-photon emission computed tomography for myocardial perfusion imaging of mice. J Am Coll Cardiol 42(3):576–582. https://doi.org/10.1016/S0735-1097(03)00716-2
Xin H, Cheng C, Li B (2013) Trapping and delivery of Escherichia coli in a microfluidic channel using an optical nanofiber. Nanoscale 5(15):6720–6724. https://doi.org/10.1039/C3NR02088F
Xin H, Li Y, Liu Y-C, Zhang Y, Xiao Y-F, Li B (2020) Optical forces: from fundamental to biological applications. Adv Mater 32(37):2001994. https://doi.org/10.1002/adma.202001994
Xu H, Huang S, Qiu C, Liu S, Deng J, Jiao B, Tan X, Ai L, Xiao Y, Belliato M (2020) Monitoring and management of home-quarantined patients with COVID-19 using a WeChat-based telemedicine system: retrospective cohort study. J Med Internet Res 22(7):e19514
Yang X, Shu W, Wang Y, Gong Y, Gong C, Chen Q, Tan X, Peng G-D, Fan X, Rao Y-J (2019) Turbidimetric inhibition immunoassay revisited to enhance its sensitivity via an optofluidic laser. Biosens Bioelectron 131:60–66. https://doi.org/10.1016/j.bios.2019.02.013
Yang T-J, Yu P-Y, Chang Y-C, Hsu S-TD (2021) D614G mutation in the SARS-CoV-2 spike protein enhances viral fitness by desensitizing it to temperature-dependent denaturation. J Biol Chem 297(4):101238. https://doi.org/10.1016/j.jbc.2021.101238
Yu H, Tan Y, Cunningham BT (2014) Smartphone fluorescence spectroscopy. Anal Chem 86(17):8805–8813. https://doi.org/10.1021/ac502080t
Yu X-C, Zhi Y, Tang S-J, Li B-B, Gong Q, Qiu C-W, Xiao Y-F (2018) Optically sizing single atmospheric particulates with a 10-nm resolution using a strong evanescent field. Light Sci Appl 7(4):18003–18003. https://doi.org/10.1038/lsa.2018.3
Yurkovetskiy L, Wang X, Pascal KE, Tomkins-Tinch C, Nyalile TP, Wang Y, Baum A, Diehl WE, Dauphin A, Carbone C, Veinotte K, Egri SB, Schaffner SF, Lemieux JE, Munro JB, Rafique A, Barve A, Sabeti PC, Kyratsous CA, Dudkina NV, Shen K, Luban J (2020) Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183(3):739-751.e8. https://doi.org/10.1016/j.cell.2020.09.032
Zandi M, Soltani S, Sanami S, Rasooli A (2021) Spike protein mutations and the effects on SARS-CoV-2 pathogenesis. JCMA 6(2):148–153. https://doi.org/10.22037/jcma.v6i2.33800
Zhang Y, Dou X, Dai Y, Wang X, Min C, Yuan X (2018) All-optical manipulation of micrometer-sized metallic particles. Photonics Res 6(2):66–71. https://doi.org/10.1364/PRJ.6.000066
Zhang Q, Yu H, Barbiero M, Wang B, Gu M (2019) Artificial neural networks enabled by nanophotonics. Light Sci Appl 8(1):42. https://doi.org/10.1038/s41377-019-0151-0
Zhang H, Penninger JM, Li Y, Zhong N, Slutsky AS (2020a) Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Intensive Care Med 46(4):586–590. https://doi.org/10.1007/s00134-020-05985-9
Zhang L, Fang X, Liu X, Ou H, Zhang H, Wang J, Li Q, Cheng H, Zhang W, Luo Z (2020b) Discovery of sandwich type COVID-19 nucleocapsid protein DNA aptamers. ChemComm 56(70):10235–10238. https://doi.org/10.1039/D0CC03993D
Zhang Y, Xi H, Juhas M (2021) Biosensing detection of the SARS-CoV-2 D614G mutation. Trends Genet 37(4):299–302. https://doi.org/10.1016/j.tig.2020.12.004
Zhao X, Zhao N, Shi Y, Xin H, Li B (2020) Optical fiber tweezers: a versatile tool for optical trapping and manipulation. Micromachines 11(2):114
Zhou D, Dejnirattisai W, Supasa P, Liu C, Mentzer AJ, Ginn HM, Zhao Y, Duyvesteyn HME, Tuekprakhon A, Nutalai R, Wang B, Paesen GC, Lopez-Camacho C, Slon-Campos J, Hallis B, Coombes N, Bewley K, Charlton S, Walter TS, Skelly D, Lumley SF, Dold C, Levin R, Dong T, Pollard AJ, Knight JC, Crook D, Lambe T, Clutterbuck E, Bibi S, Flaxman A, Bittaye M, Belij-Rammerstorfer S, Gilbert S, James W, Carroll MW, Klenerman P, Barnes E, Dunachie SJ, Fry EE, Mongkolsapaya J, Ren J, Stuart DI, Screaton GR (2021) Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine-induced sera. Cell 184(9):2348–2361.e6. https://doi.org/10.1016/j.cell.2021.02.037
Zou L, Ruan F, Huang M, Liang L, Huang H, Hong Z, Yu J, Kang M, Song Y, Xia J, Guo Q, Song T, He J, Yen H-L, Peiris M, Wu J (2020) SARS-CoV-2 viral load in upper respiratory specimens of infected patients. N Engl J Med 382(12):1177–1179. https://doi.org/10.1056/NEJMc2001737
Funding
The authors gratefully acknowledge the financial support of the Department of Electrical, Electronic and Systems Engineering/Faculty of Engineering and Built Environment/Universiti Kebangsaan Malaysia (UKM) for their encouragement and grant support (TAP-K013436) and GUP-2019–010).
Author information
Authors and Affiliations
Contributions
B.A.T., wrote the initial manuscript draft, Q.A.J., conceptualization and methodology, Y.A.M., investigation, and N.A., supervision and validation with the support of M.S.D. Z. and A.A.A.B. All authors have reviewed and accepted the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
This manuscript has not been previously published and is not now being considered by any journal for publication.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Taha, B.A., Al-Jubouri, Q., Al Mashhadany, Y. et al. Photonics enabled intelligence system to identify SARS-CoV 2 mutations. Appl Microbiol Biotechnol 106, 3321–3336 (2022). https://doi.org/10.1007/s00253-022-11930-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-11930-1