Abstract
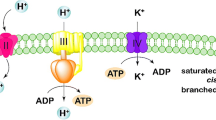
Zygosaccharomyces rouxii plays important roles in the brewing process of fermented foods such as soy sauce, where salt stress is a frequently encountered condition. In this study, effect of heat preadaptation on salt tolerance of Z. rouxii and the protective mechanisms underlying heat preadaptation were investigated based on physiological and transcriptomic analyses. Results showed that cells subjected to heat preadaptation (37 °C, 90 min) prior to salt stress aroused many physiological responses, including maintaining cell surface smooth and intracellular pH level, increasing Na+/K+-ATPase activity. Cells subjected to heat preadaptation increased the amounts of unsaturated fatty acids (palmitoleic C16:1, oleic C18:1, linoleic C18:2) and decreased the amounts of saturated fatty acids (palmitic C16:0, stearic C18:0) which caused the unsaturation degree (unsaturated/saturated = U/S ratio) increased by 2.4 times when compared with cells without preadaptation under salt stress. Besides, salt stress led to increase in contents of 5 amino acids (valine, proline, threonine, glycine, and tyrosine) and decrease of 2 amino acids (serine and lysine). When comparing the cells pre-exposed to heat preadaptation followed by challenged with salt stress and the cells without preadaptation under salt stress, the serine, threonine, and lysine contents increased significantly. RNA sequencing revealed that the metabolic level of glycolysis by Z. rouxii was weakened, while the metabolic levels of the pentose phosphate pathway and the riboflavin were enhanced in cells during heat preadaptation. Results presented in this study may contribute to understand the bases of adaptive responses in Z. rouxii and rationalize its exploitation in industrial processes.
Key points
• Heat preadaptation can improve high salinity tolerance of Z. rouxii.
• Combined physiological and transcriptomic analyses of heat preadaptation mechanisms.
• Provide theoretical support for the application of Z. rouxii.









Similar content being viewed by others
References
Becerra M, Lombardía LJ, González-Siso MI, Rodríguez-Belmonte E, Hauser NC, Cerdán ME (2003) Genome-wide analysis of the yeast transcriptome upon heat and cold shock. Comp Funct Genom 4(4):366–375. https://doi.org/10.1002/cfg.301
Berry DB, Gasch AP (2008) Stress-activated genomic expression changes serve a preparative role for impending stress in yeast. Mol Biol Cell 19(11):4580–4587. https://doi.org/10.1091/mbc.E07-07-0680
Bracey D, Holyoak CD, Nebe-von Caron G, Coote PJ (1998) Determination of the intracellular pH (pHi) of growing cells of Saccharomyces cerevisiae: the effect of reduced-expression of the membrane H+-ATPase. J Microbiol Methods 31(3):113–125. https://doi.org/10.1016/S0167-7012(97)00095-X
Brown A, Fernández IS, Gordiyenko Y, Ramakrishnan V (2016) Ribosome-dependent activation of stringent control. Nature 534(7606):277–280. https://doi.org/10.1038/nature17675
Capaldi AP, Kaplan T, Liu Y, Habib N, Regev A, Friedman N, Regev A, Friedman N, O'Shea EK (2008) Structure and function of a transcriptional network activated by the MAPK Hog1. Nat Genet 40(11):1300–1306. https://doi.org/10.1017/S003118200999117X
Cheng Z, Chi M, Li G, Chen H, Sui Y, Sun H, Wisniewski M, Liu Y, Liu J (2016) Heat shock improves stress tolerance and biocontrol performance of Rhodotorula mucilaginosa. Biol Control 95:49–56. https://doi.org/10.1016/j.biocontrol.2016.01.001
Chuang MY, Tsai WC, Kuo TY, Chen HM, Chen WJ (2016) Comparative proteome analysis reveals proteins involved in salt adaptation in Photobacterium damselae subsp. piscicida. J Basic Microbiol 56(11):1234–1243. https://doi.org/10.1002/jobm.201600091
Cui R, Zheng J, Wu C, Zhou R (2014) Effect of different halophilic microbial fermentation patterns on the volatile compound profiles and sensory properties of soy sauce moromi. Eur Food Res Technol 239(2):321–331. https://doi.org/10.1007/s00217-014-2225-9
Dakal TC, Solieri L, Giudici P (2014) Adaptive response and tolerance to sugar and salt stress in the food yeast Zygosaccharomyces rouxii. Int J Food Microbiol 185:140–157. https://doi.org/10.1016/j.ijfoodmicro.2014.05.015
Devanthi PVP, Linforth R, Kadri HE, Gkatzionis K (2018) Water-in-oil-in-water double emulsion for the delivery of starter cultures in reduced-salt moromi fermentation of soy sauce. Food Chem 257:243–251. https://doi.org/10.1016/j.foodchem.2018.03.022
Dhar R, Sägesser R, Weikert C, Wagner A (2012) Yeast adapts to a changing stressful environment by evolving cross-protection and anticipatory gene regulation. Mol Biol Evol 30(3):573–588. https://doi.org/10.1093/molbev/mss253
Gandhi A, Shah NP (2016) Effect of salt stress on morphology and membrane composition of Lactobacillus acidophilus, Lactobacillus casei, and Bifidobacterium bifidum, and their adhesion to human intestinal epithelial-like Caco-2 cells. J Diary Sci 99(4):2594–2605. https://doi.org/10.3168/jds.2015-10718
Gong X, Yu H, Chen J, Han B (2012) Cell surface properties of Lactobacillus salivarius under osmotic stress. Eur Food Res Technol 234(4):671–678. https://doi.org/10.1007/s00217-012-1677-z
Greenacre EJ, Brocklehurst TF (2006) The acetic acid tolerance response induces cross-protection to salt stress in Salmonella typhimurium. Int J Food Microbiol 112(1):62–65. https://doi.org/10.1016/j.ijfoodmicro.2006.05.012
Guan Q, Haroon S, Bravo DG, Will JL, Gasch AP (2012) Cellular memory of acquired stress resistance in Saccharomyces cerevisiae. Genetics 192(2):495–505. https://doi.org/10.1534/genetics.112.143016
He G, Deng J, Wu C, Huang J (2017a) A partial proteome reference map of Tetragenococcus halophilus and comparative proteomic and physiological analysis under salt stress. RSC Adv 7(21):12753–12763. https://doi.org/10.1039/c6ra22521g
He G, Wu C, Huang J, Zhou R (2017b) Metabolic response of Tetragenococcus halophilus under salt stress. Biotechnol Bioproc E 22(4):366–375. https://doi.org/10.1007/s12257-017-0015-5
Hou L, Yu Y, Wang C, Wang C (2014) Research on salt-tolerant gene GPD1 in Zygosaccharomyces rouxii. Process of the 2012 international Conference on Applied Biotechnology (ICAB 2012). Lect Notes Electr Eng 250:1157–1163. https://doi.org/10.1007/978-3-642-37922-2_123
Jansen M, Veurink JH, Euverink GJ, Dijkhuizen L (2003) Growth of the salt-tolerant yeast Zygosaccharomyces rouxii in microtiter plates: effects of NaCl, pH and temperature on growth and fusel alcohol production from branched-chain amino acids. FEMS Yeast Res 3(3):313–318. https://doi.org/10.1016/S1567-1356(02)00162-9
Komisarczuk AZ, Kongshaug H, Nilsen F (2018) Gene silencing reveals multiple functions of Na+/K+-ATPase in the salmon louse ( Lepeophtheirus salmonis ). Exp Parasitol 185:79–91. https://doi.org/10.1016/j.exppara.2018.01.005
Long X, Tian J, Liao X, Tian Y (2018) Adaptations of Bacillus shacheensis HNA-14 required for long-term survival under osmotic challenge: a multi-omics perspective. RSC Adv 8(48):27525–27536. https://doi.org/10.1039/C8RA05472J
Machielsen R, van Alen-Boerrigter IJ, Koole LA, Bongers RS, Kleerebezem M, Van Hylckama Vlieg JET (2010) Indigenous and environmental modulation of frequencies of mutation in Lactobacillus plantarum. Appl Environ Microbiol 76(5):1587–1595. https://doi.org/10.1128/AEM.02595-09
Marin K, Stirnberg M, Eisenhut M, Krämer R, Hagemann M (2006) Osmotic stress in Synechocystis sp. PCC 6803: low tolerance towards nonionic osmotic stress results from lacking activation of glucosylglycerol accumulation. Microbiol 152(7):2023–2030. https://doi.org/10.1099/mic.0.28771-0
Martínez-Pastor MT, Marchler G, Schüller C, Marchler-Bauer A, Ruis H, Estruch F (1996) The Saccharomyces cerevisiae zinc finger proteins Msn2p and Msn4p are required for transcriptional induction through the stress response element (STRE). EMBO J 15(9):2227–2235. https://doi.org/10.1002/j.1460-2075.1996.tb00576.x
Meng Q, Hatakeyama M, Sugawara E (2014) Formation by yeast of 2-furanmethanethiol and ethyl 2-mercaptopropionate aroma compounds in Japanese soy sauce. Biosci Biotechnol Biochem 78(1):109–114. https://doi.org/10.1080/09168451.2014.877820
Mitchell A, Romano GH, Groisman B, Yona A, Dekel E, Kupiec M, Dahan O, Pilpel Y (2009) Adaptive prediction of environmental changes by microorganisms. Nature 7252(460):220–225. https://doi.org/10.1038/nature08112
Pittman JR, Buntyn JO, Posadas G, Nanduri B, Pendarvis K, Donaldson JR (2015) Proteomic analysis of cross protection provided between cold and osmotic stress in Listeria monocytogenes. J Protemoe Res 13(4):1896–1904. https://doi.org/10.1021/pr401004a
Shi X, Ren J, Qin Y, Zhou S, Wang X (2017) Overexpression of SDH confers tolerance to salt and osmotic stress, but decreases ABA sensitivity in Arabidopsis. Plant Biol 20(2):327–337. https://doi.org/10.1111/plb.12664
Święciło A (2016) Cross-stress resistance in Saccharomyces cerevisiae yeast-new insight into an old phenomenon. Cell Stress Chaperones 21(2):187–200. https://doi.org/10.1007/s12192-016-0667-7
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatcis 25(9):1105–1111. https://doi.org/10.1093/bioinformatics/btp120
Tripathy S, Sen R, Padhi SK, Mohanty S, Maiti NK (2014) Upregulation of transcripts for metabolism in diverse environments is a shared response associated with survival and adaptation of Klebsiella pneumoniaein response to temperature extremes. Funct Integr Genomic 14(3):591–601. https://doi.org/10.1007/s10142-014-0382-3
Tutar Y, Okan S (2012) Heat shock protein 70 purification and characterization from Cyprinion macrastomus macrastomus and Garra rufa obtusa. J Therm Biol 37(1):95–99. https://doi.org/10.1016/j.jtherbio.2011.11.002
Wang D, Hao Z, Zhao J, Jin Y, Huang J, Zhou R, Wu C (2019a) Comparative physiological and transcriptomic analyses reveal salt tolerance mechanisms of Zygosaccharomyces rouxii. Process Biochem 82:59–67. https://doi.org/10.1016/j.procbio.2019.04.009
Wang D, Zhang M, Huang J, Zhou R, Jin Y, Wu C (2019b) Zygosaccharomyces rouxii combats salt stress by maintaining cell membrane structure and functionality. J Microbiol Biotechnol 30(1):62–70. https://doi.org/10.4014/jmb.1904.04006
Watanabe Y, Hirasaki M, Tohnai N, Yagi K, Abe S, Tamai Y (2003) Salt shock enhances the expression of ZrATP2, the gene for the mitochondrial ATPase β subunit of Zygosaccharomyces rouxii. J Biosci Bioeng 96(2):193–195. https://doi.org/10.1016/S1389-1723(03)90125-3
Wu C, Zhang J, Chen W, Wang M, Du G, Chen J (2012) A combined physiological and proteomic approach to reveal lactic-acid-induced alterations in Lactobacillus casei Zhang and its mutant with enhanced lactic acid tolerance. Appl Microbiol Biotechnol 93(2):707–722. https://doi.org/10.1007/s00253-011-3757-6
Wu Y, Ding W, Jia L, He Q (2015) The rheological properties of tara gum (Caesalpinia spinosa). Food Chem 168(1):366–371. https://doi.org/10.1016/j.foodchem.2014.07.083
Xiao M, Zhu X, Xu H, Tang J, Zhang X (2017) A novel point mutation in RpoB improves osmotolerance and succinic acid production in Escherichia coli. BMC Biotechnol 17(10):1–11. https://doi.org/10.1186/s12896-017-0337-6
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S, Kong L, Gao G, Li CY, Wei L (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:316–322. https://doi.org/10.1093/nar/gkr483
Yan X, Xun M, Li J, Wu L, Zheng J (2016) Activation of Na+/K+-ATPase attenuates high glucose-induced H9c2 cell apoptosis via suppressing ROS accumulation and MAPKs activities by DRm217. Acta Biochim Biophys Sin 48(10):883–893. https://doi.org/10.1093/abbs/gmw079
Yao S, Zhou R, Jin Y, Zhang L, Huang J, Wu C (2019) Effect of co-culture with Tetragenococcus halophilus on the physiological characterization and transcription profiling of Zygosaccharomyces rouxii. Food Res Int 121(7):348–358. https://doi.org/10.1016/j.foodres.2019.03.053
Zhang X, St Leger RJ, Fang W (2018) Stress-induced pyruvate accumulation contributes to cross protection in a fungus. Environ Microbiol 20(3):1158–1169. https://doi.org/10.1111/1462-2920.14058
Zhao S, Zhang Q, Hao G, Liu X, Zhao J, Chen Y, Zhang H, Chen W (2014) The protective role of glycine betaine in Lactobacillus plantarum ST-III against salt stress. Food Control 44:208–213. https://doi.org/10.1016/j.foodcont.2014.04.002
Zheng J, Liang R, Zhang L, Wu C, Zhou R, Liao X (2013) Characterization of microbial communities in strong aromatic liquor fermentation pit muds of different ages assessed by combined DGGE and PLFA analyses. Food Res Int 54(1):660–666. https://doi.org/10.1016/j.foodres.2013.07.058
Zhong X, Ren AM, Guo JF, Liu XT, Feng JK (2012) A theoretical investigation of two typical two-photon pH fluorescent probes. Photochem Photobiol 89(2):300–309. https://doi.org/10.1111/php.12015
Zhou A, Baidoo E, He Z, Mukhopadhyay A, Baumohl JK, Benke P, Joachimiak MP, Xie M, Song R, Arkin AP (2013) Characterization of NaCl tolerance in Desulfovibrio vulgaris Hildenborough through experimental evolution. ISME J 7(9):1790–1802. https://doi.org/10.1038/ismej.2013.60
Acknowledgments
The authors would thank Zhonghui Wang (College of Biomass Science and Engineering, Sichuan University) for her great help in AFM and amino acid analyzer observation.
Funding
This work was financially supported by the National Natural Science Foundation of China (31871787, 31671849), and Open Funding Project of Key Laboratory of Wuliangye-Flavor Liquor Solid-State Fermentation, China National Light Industry (2019JJ004).
Author information
Authors and Affiliations
Contributions
D.W., C.W., J. Z., R.Z., J. H., D.Z., and Y.J. conceived and designed research. D.W. and M.Z. conducted experiments. D.W., M.Z., and C.W. analyzed data and wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, D., Zhang, M., Huang, J. et al. Heat preadaptation improved the ability of Zygosaccharomyces rouxii to salt stress: a combined physiological and transcriptomic analysis. Appl Microbiol Biotechnol 105, 259–270 (2021). https://doi.org/10.1007/s00253-020-11005-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-11005-z