Abstract
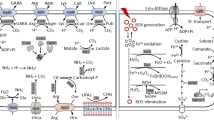
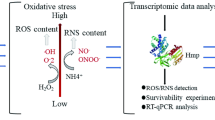
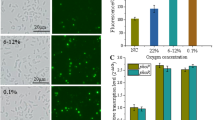
The industrial organism Corynebacterium glutamicum is often subjected to acid stress during large-scale fermentation for the production of bio-based chemicals. The capacity of the cells to thrive in acidic environments is a prerequisite for achieving high product yields. In this study, we obtained an acid-adapted strain using an adaptive laboratory evolution strategy. Physiological characterizations revealed that the adapted strain achieved improved cell viability after acid-stress challenge, with a higher cytoplasmic pHin level, a lower intracellular reactive oxygen species (ROS), and an enhanced morphological integrity of the cells, when compared to those of the original control strain. Transcriptome analysis indicated that several important cellular processes were altered in the adapted strain, including sulfur metabolism, iron transport, and central metabolic pathways. Further research displayed that KatA and Dps cooperatively mediated intracellular ROS scavenging, which was required for resistance to low-pH stress in C. glutamicum. Furthermore, the repression of sulfur assimilation by the McbR regulator also contributed to the improvement of acid-stress tolerance. Moreover, two copper chaperone genes cg1328 and cg3292 were found to be involved in promoting cell survival under acid-stress conditions. Finally, a new recombinant C. glutamicum strain with enhanced acid tolerance was generated by the combined overexpression of katA, dps, mcbR, and cg1328, showing 18.4 ± 2.5% higher biomass yields than the wild-type strain under acid-stress conditions. These findings will provide new insights into the understanding and genetic improvement of acid tolerance in C. glutamicum.






Similar content being viewed by others
References
Andrews SC, Robinson AK, Rodríguez-Quiñones F (2003) Bacterial iron homeostasis. FEMS Microbiol Rev 27(2–3):215–237. https://doi.org/10.1016/S0168-6445(03)00055-X
Beales N (2004) Adaptation of microorganisms to cold temperatures, weak acid preservatives, low pH, and osmotic stress: a review. Compr Rev Food Sci F 3(1):1–20. https://doi.org/10.1111/j.1541-4337.2004.tb00057.x
Bellapadrona G, Ardini M, Ceci P, Stefanini S, Chiancone E (2010) Dps proteins prevent Fenton-mediated oxidative damage by trapping hydroxyl radicals within the protein shell. Free Radic Biol Med 48(2):292–297. https://doi.org/10.1016/j.freeradbiomed.2009.10.053
Brune I, Werner H, Huser AT, Kalinowski J, Puhler A, Tauch A (2006) The DtxR protein acting as dual transcriptional regulator directs a global regulatory network involved in iron metabolism of Corynebacterium glutamicum. BMC Genomics 7(21):21. https://doi.org/10.1186/1471-2164-7-21
Cecchini G (2003) Function and structure of complex II of the respiratory chain. Annu Rev Biochem 72:77–109. https://doi.org/10.1146/annurev.biochem.72.121801.161700
Chung HJ, Bang W, Drake MA (2006) Stress response of Escherichia coli. Compr Rev Food Sci Food Saf 5(3):52–64. https://doi.org/10.1111/j.1541-4337.2006.00002.x
Djoko KY, Phan MD, Peters KM, Walker MJ, Schembri MA, McEwan AG (2017) Interplay between tolerance mechanisms to copper and acid stress in Escherichia coli. Proc Natl Acad Sci U S A 114(26):6818–6823. https://doi.org/10.1073/pnas.1620232114
Follmann M, Ochrombel I, Krämer R, Trötschel C, Poetsch A, Rückert C, Hüser A, Persicke M, Seiferling D, Kalinowski J, Marin K (2009) Functional genomics of pH homeostasis in Corynebacterium glutamicum revealed novel links between pH response, oxidative stress, iron homeostasis and methionine synthesis. BMC Genomics 10:621. https://doi.org/10.1186/1471-2164-10-621
Foster JW (2004) Escherichia coli acid resistance: tales of an amateur acidophile. Nat Rev Microbiol 2(11):898–907. https://doi.org/10.1038/nrmicro1021
Genevaux P, Georgopoulos C, Kelley WL (2007) The Hsp70 chaperone machines of Escherichia coli: a paradigm for the repartition of chaperone functions. Mol Microbiol 66(4):840–857. https://doi.org/10.1111/j.1365-2958.2007.05961.x
Giuffre A, Borisov VB, Arese M, Sarti P, Forte E (2014) Cytochrome bd oxidase and bacterial tolerance to oxidative and nitrosative stress. Biochim Biophys Acta 1837(7):1178–1187. https://doi.org/10.1016/j.bbabio.2014.01.016
Harrison MD, Jones CE, Dameron CT (1999) Copper chaperones: function, structure and copper-binding properties. J Biol Inorg Chem 4(2):145–153. https://doi.org/10.1007/s007750050297
Heydari H, Siow CC, Tan MF, Jakubovics NS, Wee WY, Mutha NV, Wong GJ, Ang MY, Yazdi AH, Choo SW (2014) CoryneBase: Corynebacterium genomic resources and analysis tools at your fingertips. PLoS One 9(1):e86318. https://doi.org/10.1371/journal.pone.0086318
Jakoby M, Ngouoto-Nkili CE, Burkovski A (1999) Construction and application of new Corynebacterium glutamicum vectors. Biotechnol Tech 13(6):437–441. https://doi.org/10.1023/A:1008968419217
Kakinuma Y (1998) Inorganic cation transport and energy transduction in Enterococcus hirae and other streptococci. Microbiol Mol Biol Rev 62(4):1021–1045
Kanjee U, Houry WA (2013) Mechanisms of acid resistance in Escherichia coli. Annu Rev Microbiol 67:65–81. https://doi.org/10.1146/annurev-micro-092412-155708
Keilhauer C, Eggeling L, Sahm H (1993) Isoleucine synthesis in Corynebacterium glutamicum: molecular analysis of the ilvB-ilvN-ilvC operon. J Bacteriol 175(17):5595–5603. https://doi.org/10.1128/jb.175.17.5595-5603.1993
Kobayashi H, Saito H, Kakegawa T (2000) Bacterial strategies to inhabit acidic environments. J Gen Appl Microbiol 46(5):235–243. https://doi.org/10.2323/Jgam.46.235
Krulwich TA, Sachs G, Padan E (2011) Molecular aspects of bacterial pH sensing and homeostasis. Nat Rev Microbiol 9(5):330–343. https://doi.org/10.1038/nrmicro2549
Lee JY, Na YA, Kim E, Lee HS, Kim P (2016) The actinobacterium Corynebacterium glutamicum, an industrial workhorse. J Microbiol Biotechnol 26(5):807–822. https://doi.org/10.4014/jmb.1601.01053
Lee JY, Seo J, Kim ES, Lee HS, Kim P (2013) Adaptive evolution of Corynebacterium glutamicum resistant to oxidative stress and its global gene expression profiling. Biotechnol Lett 35(5):709–717. https://doi.org/10.1007/s10529-012-1135-9
Letek M, Fiuza M, Ordonez E, Villadangos AF, Ramos A, Mateos LM, Gil JA (2008) Cell growth and cell division in the rod-shaped actinomycete Corynebacterium glutamicum. Antonie Van Leeuwenhoek 94(1):99–109. https://doi.org/10.1007/s10482-008-9224-4
Liu Y, Yang X, Yin Y, Lin J, Chen C, Pan J, Si M, Shen X (2016) Mycothiol protects Corynebacterium glutamicum against acid stress via maintaining intracellular pH homeostasis, scavenging ROS, and S-mycothiolating MetE. J Gen Appl Microbiol 62(3):144–153. https://doi.org/10.2323/jgam.2016.02.001
Liu YP, Tang HZ, Lin ZL, Xu P (2015) Mechanisms of acid tolerance in bacteria and prospects in biotechnology and bioremediation. Biotechnol Adv 33(7):1484–1492. https://doi.org/10.1016/j.biotechadv.2015.06.001
Lund P, Tramonti A, De Biase D (2014) Coping with low pH: molecular strategies in neutralophilic bacteria. FEMS Microbiol Rev 38(6):1091–1125. https://doi.org/10.1111/1574-6976.12076
Marles-Wright J, Lewis RJ (2007) Stress responses of bacteria. Curr Opin Struct Biol 17(6):755–760. https://doi.org/10.1016/j.sbi.2007.08.004
Milse J, Petri K, Ruckert C, Kalinowski J (2014) Transcriptional response of Corynebacterium glutamicum ATCC 13032 to hydrogen peroxide stress and characterization of the OxyR regulon. J Biotechnol 190:40–54. https://doi.org/10.1016/j.jbiotec.2014.07.452
Mols M, van Kranenburg R, van Melis CC, Moezelaar R, Abee T (2010) Analysis of acid-stressed Bacillus cereus reveals a major oxidative response and inactivation-associated radical formation. Environ Microbiol 12(4):873–885. https://doi.org/10.1111/j.1462-2920.2009.02132.x
Okibe N, Suzuki N, Inui M, Yukawa H (2011) Efficient markerless gene replacement in Corynebacterium glutamicum using a new temperature-sensitive plasmid. J Microbiol Meth 85(2):155–163. https://doi.org/10.1016/j.mimet.2011.02.012
Palumaa P (2013) Copper chaperones. The concept of conformational control in the metabolism of copper. FEBS Lett 587(13):1902–1910. https://doi.org/10.1016/j.febslet.2013.05.019
Papadimitriou K, Alegria A, Bron PA, de Angelis M, Gobbetti M, Kleerebezem M, Lemos JA, Linares DM, Ross P, Stanton C, Turroni F, van Sinderen D, Varmanen P, Ventura M, Zuniga M, Tsakalidou E, Kok J (2016) Stress physiology of lactic acid bacteria. Microbiol Mol Biol Rev 80(3):837–890. https://doi.org/10.1128/MMBR.00076-15
Park S, Imlay JA (2003) High levels of intracellular cysteine promote oxidative DNA damage by driving the Fenton reaction. J Bacteriol 185(6):1942–1950. https://doi.org/10.1128/jb.185.6.1942-1950.2003
Radmacher E, Alderwick LJ, Besra GS, Brown AK, Gibson KJ, Sahm H, Eggeling L (2005) Two functional FAS-I type fatty acid synthases in Corynebacterium glutamicum. Microbiology 151(Pt 7):2421–2427. https://doi.org/10.1099/mic.0.28012-0
Rausch T, Wachter A (2005) Sulfur metabolism: a versatile platform for launching defence operations. Trends Plant Sci 10(10):503–509. https://doi.org/10.1016/j.tplants.2005.08.006
Rey DA, Pühler A, Kalinowski J (2003) The putative transcriptional repressor McbR, member of the TetR-family, is involved in the regulation of the metabolic network directing the synthesis of sulfur containing amino acids in Corynebacterium glutamicum. J Biotechnol 103(1):51–65. https://doi.org/10.1016/S0168-1656(03)00073-7
Rivera M (2017) Bacterioferritin: structure, dynamics, and protein-protein interactions at play in iron storage and mobilization. Acc Chem Res 50(2):331–340. https://doi.org/10.1021/acs.accounts.6b00514
Ruckert C, Milse J, Albersmeier A, Koch DJ, Puhler A, Kalinowski J (2008) The dual transcriptional regulator CysR in Corynebacterium glutamicum ATCC 13032 controls a subset of genes of the McbR regulon in response to the availability of sulphide acceptor molecules. BMC Genomics 9:483. https://doi.org/10.1186/1471-2164-9-483
Si M, Zhao C, Burkinshaw B, Zhang B, Wei D, Wang Y, Dong TG, Shen X (2017) Manganese scavenging and oxidative stress response mediated by type VI secretion system in Burkholderia thailandensis. Proc Natl Acad Sci U S A 114(11):E2233–E2242. https://doi.org/10.1073/pnas.1614902114
Silhavy TJ, Kahne D, Walker S (2010) The bacterial cell envelope. Cold Spring Harb Perspect Biol 2(5):a000414. https://doi.org/10.1101/cshperspect.a000414
Teramoto H, Inui M, Yukawa H (2013) OxyR acts as a transcriptional repressor of hydrogen peroxide-inducible antioxidant genes in Corynebacterium glutamicum R. FEBS J 280(14):3298–3312. https://doi.org/10.1111/febs.12312
Wang T, Gao F, Kang Y, Zhao C, Su T, Li M, Si M, Shen X (2016) Mycothiol peroxidase MPx protects Corynebacterium glutamicum against acid stress by scavenging ROS. Biotechnol Lett 38(7):1221–1228. https://doi.org/10.1007/s10529-016-2099-y
Wennerhold J, Krug A, Bott M (2005) The AraC-type regulator RipA represses aconitase and other iron proteins from Corynebacterium under iron limitation and is itself repressed by DtxR. J Biol Chem 280(49):40500–40508. https://doi.org/10.1074/jbc.M508693200
Wieschalka S, Blombach B, Bott M, Eikmanns BJ (2013) Bio-based production of organic acids with Corynebacterium glutamicum. Microb Biotechnol 6(2):87–102. https://doi.org/10.1111/1751-7915.12013
Winterbourn CC (1995) Toxicity of iron and hydrogen peroxide: the Fenton reaction. Toxico Lett 82-83:969–974. https://doi.org/10.1016/0378-4274(95)03532-X
Wu C, He G, Zhang J (2014) Physiological and proteomic analysis of Lactobacillus casei in response to acid adaptation. J Ind Microbiol Biotechnol 41(10):1533–1540. https://doi.org/10.1007/s10295-014-1487-3
Wu C, Zhang J, Wang M, Du G, Chen J (2012) Lactobacillus casei combats acid stress by maintaining cell membrane functionality. J Ind Microbiol Biotechnol 39(7):1031–1039. https://doi.org/10.1007/s10295-012-1104-2
Xu N, Wei L, Liu J (2017) Biotechnological advances and perspectives of γ-aminobutyric acid production. World J Microbiol Biot 33(3):64. https://doi.org/10.1007/s11274-017-2234-5
Xu N, Zheng YY, Wang XC, Krulwich TA, Ma YH, Liu J (2018) The lysine 299 residue endows the multisubunit Mrp1 antiporter with dominant roles in Na+ resistance and pH homeostasis in Corynebacterium glutamicum. Appl Environ Microbiol 84(10):e00110–e00118. https://doi.org/10.1128/AEM.00110-18
Zhang J, Fu RY, Hugenholtz J, Li Y, Chen J (2007) Glutathione protects Lactococcus lactis against acid stress. Appl Environ Microbiol 73(16):5268–5275. https://doi.org/10.1128/AEM.02787-06
Acknowledgements
We are grateful to Prof. Masayuki Inui (Research Institute of Innovative Technology for the Earth, Japan) for generously providing plasmids.
Funding
This study was supported by the National Natural Science Foundation of China (No. 31500044), the Natural Science Foundation of Tianjin (No. 17JCQNJC09600, No. 17JCYBJC24000), the Tianjin Science and Technology Project (15PTCYSY00020), the Foundation of Hebei Educational Committee (ZD2017047) and the “Hundred Talents Program” of the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 429 kb)
Rights and permissions
About this article
Cite this article
Xu, N., Lv, H., Wei, L. et al. Impaired oxidative stress and sulfur assimilation contribute to acid tolerance of Corynebacterium glutamicum. Appl Microbiol Biotechnol 103, 1877–1891 (2019). https://doi.org/10.1007/s00253-018-09585-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-09585-y