Abstract
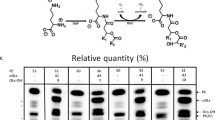
Bacterial cytochrome P450 enzymes in cytochrome P450 (CYP)153 family were recently reported as fatty acid ω-hydroxylase. Among them, CYP153As from Marinobacter aquaeolei VT8 (CYP153A33), Alcanivorax borkumensis SK2 (CYP153A13), and Gordonia alkanivorans (CYP153A35) were selected, and their specific activities and product yields of ω-hydroxy palmitic acid based on whole cell reactions toward palmitic acid were compared. Using CamAB as redox partner, CYP153A35 and CYP153A13 showed the highest product yields of ω-hydroxy palmitic acid in whole cell and in vitro reactions, respectively. Artificial self-sufficient CYP153A35-BMR was constructed by fusing it to the reductase domain of CYP102A1 (i.e., BM3) from Bacillus megaterium, and its catalytic activity was compared with CYP153A35 and CamAB systems. Unexpectedly, the system with CamAB resulted in a 1.5-fold higher yield of ω-hydroxy palmitic acid than that using A35-BMR in whole cell reactions, whereas the electron coupling efficiency of CYP153A35-BM3 reductase was 4-fold higher than that of CYP153A35 and CamAB system. Furthermore, various CamAB expression systems according to gene arrangements of the three proteins and promoter strength in their gene expression were compared in terms of product yields and productivities. Tricistronic expression of the three proteins in the order of putidaredoxin (CamB), CYP153A35, and putidaredoxin reductase (CamA), i.e., A35-AB2, showed the highest product yield from 5 mM palmitic acid for 9 h in batch reaction owing to the concentration of CamB, which is the rate-limiting factor for the activity of CYP153A35. However, in fed-batch reaction, A35-AB1, which expressed the three proteins individually using three T7 promoters, resulted with the highest product yield of 17.0 mM (4.6 g/L) ω-hydroxy palmitic acid from 20 mM (5.1 g/L) palmitic acid for 30 h.





Similar content being viewed by others
References
Asperger O, Naumann A, Kleber H-P (1981) Occurrence of cytochrome P-450 in Acinetobacter strains after growth on n-hexadecane. FEMS Microbiol Lett 11(4):309–312. doi:10.1111/j.1574-6968.1981.tb06986.x
Bae JH, Park BG, Jung E, Lee PG, Kim BG (2014) fadD deletion and fadL overexpression in Escherichia coli increase hydroxy long-chain fatty acid productivity. Appl Microbiol Biotechnol 98(21):8917–8925. doi:10.1007/s00253-014-5974-2
Bell SG, Harford-Cross CF, Wong LL (2001) Engineering the CYP101 system for in vivo oxidation of unnatural substrates. Protein Eng 14(10):797–802
Bell SG, Dale A, Rees NH, Wong LL (2010a) A cytochrome P450 class I electron transfer system from Novosphingobium aromaticivorans. Appl Microbiol Biotechnol 86(1):163–175. doi:10.1007/s00253-009-2234-y
Bell SG, Xu F, Johnson EO, Forward IM, Bartlam M, Rao Z, Wong LL (2010b) Protein recognition in ferredoxin-P450 electron transfer in the class I CYP199A2 system from Rhodopseudomonas palustris. J Biol Inorg Chem 15(3):315–328. doi:10.1007/s00775-009-0604-7
Benveniste I, Tijet N, Adas F, Philipps G, Salaun JP, Durst F (1998) CYP86A1 from Arabidopsis thaliana encodes a cytochrome P450-dependent fatty acid omega-hydroxylase. Biochem Biophys Res Commun 243(3):688–693. doi:10.1006/bbrc.1998.8156
Benveniste I, Saito T, Wang Y, Kandel S, Huang HW, Pinot F, Kahn RA, Salaun JP, Shimoji M (2006) Evolutionary relationship and substrate specificity of Arabidopsis thaliana fatty acid omega-hydroxylase. Plant Sci 170(2):326–338. doi:10.1016/j.plantsci.2005.08.028
Bernhardt R, Urlacher VB (2014) Cytochromes P450 as promising catalysts for biotechnological application: chances and limitations. Appl Microbiol Biotechnol 98(14):6185–6203. doi:10.1007/s00253-014-5767-7
Bordeaux M, de Girval D, Rullaud R, Subileau M, Dubreucq E, Drone J (2014) High-cell-density cultivation of recombinant Escherichia coli, purification and characterization of a self-sufficient biosynthetic octane omega-hydroxylase. Appl Microbiol Biotechnol 98(14):6275–6283. doi:10.1007/s00253-014-5671-1
Choi KY, Kim TJ, Koh SK, Roh CH, Pandey BP, Lee N, Kim BG (2009) A-ring ortho-specific monohydroxylation of daidzein by cytochrome P450s of Nocardia farcinica IFM10152. Biotechnol J 4(11):1586–1595. doi:10.1002/biot.200900157
Choi KY, Park HY, Kim BG (2010) ) characterization of bi-functional CYP154 from Nocardia farcinica IFM10152 in the O-dealkylation and ortho-hydroxylation of formononetin. Enzym Microb Technol 47(7):327–334. doi:10.1016/j.enzmictec.2010.08.006
Choi KY, Jung E, Jung DH, An BR, Pandey BP, Yun H, Sung C, Park HY, Kim BG (2012) Engineering of daidzein 3′-hydroxylase P450 enzyme into catalytically self-sufficient cytochrome P450. Microb Cell Factories 11:81. doi:10.1186/1475-2859-11-81
Choi KY, Jung E, Yun H, Yang YH, Kim BG (2014) Engineering class I cytochrome P450 by gene fusion with NADPH-dependent reductase and S. avermitilis host development for daidzein biotransformation. Appl Microbiol Biotechnol 98(19):8191–8200. doi:10.1007/s00253-014-5706-7
Diaper DGM, Mitchell DL (1960) An improved preparation of omega-Hydroxy aliphatic acids and their esters. Canadian Journal of Chemistry-Revue Canadienne De Chimie 38(10):1976–1982. doi:10.1139/V60-266
Dodhia VR, Fantuzzi A, Gilardi G (2006) Engineering human cytochrome P450 enzymes into catalytically self-sufficient chimeras using molecular Lego. J Biol Inorg Chem 11(7):903–916. doi:10.1007/s00775-006-0144-3
Fairhead M, Giannini S, Gillam EMJ, Gilardi G (2005) Functional characterisation of an engineered multidomain human P450 2E1 by molecular Lego. J Biol Inorg Chem 10(8):842–853. doi:10.1007/s00775-005-0033-1
Funhoff EG, Salzmann J, Bauer U, Witholt B, van Beilen JB (2007) Hydroxylation and epoxidation reactions catalyzed by CYP153 enzymes. Enzym Microb Technol 40(4):806–812. doi:10.1016/j.enzmictec.2006.06.014
Gillam EM, Baba T, Kim BR, Ohmori S, Guengerich FP (1993) Expression of modified human cytochrome P450 3 A4 in Escherichia coli and purification and reconstitution of the enzyme. Arch Biochem Biophys 305(1):123–131. doi:10.1006/abbi.1993.1401
Girhard M, Klaus T, Khatri Y, Bernhardt R, Urlacher VB (2010) Characterization of the versatile monooxygenase CYP109B1 from Bacillus subtilis. Appl Microbiol Biotechnol 87(2):595–607. doi:10.1007/s00253-010-2472-z
Hannemann F, Bichet A, Ewen KM, Bernhardt R (2007) Cytochrome P450 systems--biological variations of electron transport chains. Biochim Biophys Acta 1770(3):330–344. doi:10.1016/j.bbagen.2006.07.017
Hardwick JP (2008) Cytochrome P450 omega hydroxylase (CYP4) function in fatty acid metabolism and metabolic diseases. Biochem Pharmacol 75(12):2263–2275. doi:10.1016/j.bcp.2008.03.004
Harpaz Y, Gerstein M, Chothia C (1994) Volume changes on protein folding. Structure 2(7):641–649
Honda Malca S, Scheps D, Kuhnel L, Venegas-Venegas E, Seifert A, Nestl BM, Hauer B (2012) Bacterial CYP153A monooxygenases for the synthesis of omega-hydroxylated fatty acids. Chem Commun (CamB) 48(42):5115–5117. doi:10.1039/c2cc18103g
Johnston JB, Ouellet H, Podust LM, Ortiz de Montellano PR (2011) Structural control of cytochrome P450-catalyzed omega-hydroxylation. Arch Biochem Biophys 507(1):86–94. doi:10.1016/j.abb.2010.08.011
Kim KR, Oh DK (2013) Production of hydroxy fatty acids by microbial fatty acid-hydroxylation enzymes. Biotechnol Adv 31(8):1473–1485. doi:10.1016/j.biotechadv.2013.07.004
Kim D, Cryle MJ, De Voss JJ, Ortiz de Montellano PR (2007) Functional expression and characterization of cytochrome P450 52 A21 from Candida albicans. Arch Biochem Biophys 464(2):213–220. doi:10.1016/j.abb.2007.02.032
Lu WH, Ness JE, Xie WC, Zhang XY, Minshull J, Gross RA (2010) Biosynthesis of monomers for plastics from renewable oils. J Am Chem Soc 132(43):15451–15455. doi:10.1021/Ja107707v
Metzger JO, Bornscheuer U (2006) Lipids as renewable resources: current state of chemical and biotechnological conversion and diversification. Appl Microbiol Biotechnol 71(1):13–22. doi:10.1007/s00253-006-0335-4
Nelson DR (1998) Cytochrome P450 nomenclature. Methods Mol Biol 107:15–24. doi:10.1385/0-89603-519-0:15
Omura T, Sato R (1964) The carbon monoxide-binding pigment of liver Microsomes. I evidence for its hemoprotein. Nature J Biol Chem 239:2370–2378
Ringle M, Khatri Y, Zapp J, Hannemann F, Bernhardt R (2013) Application of a new versatile electron transfer system for cytochrome P450-based Escherichia coli whole-cell bioconversions. Appl Microbiol Biotechnol 97(17):7741–7754. doi:10.1007/s00253-012-4612-0
Scheller U, Zimmer T, Becher D, Schauer F, Schunck WH (1998) Oxygenation cascade in conversion of n-alkanes to alpha,omega-dioic acids catalyzed by cytochrome P450 52 A3. J Biol Chem 273(49):32528–32534
Scheps D, Malca SH, Hoffmann H, Nestl BM, Hauer B (2011) Regioselective omega-hydroxylation of medium-chain n-alkanes and primary alcohols by CYP153 enzymes from Mycobacterium marinum and Polaromonas sp. strain JS666. Org Biomol Chem 9(19):6727–6733. doi:10.1039/c1ob05565h
Scheps D, Honda Malca S, Richter SM, Marisch K, Nestl BM, Hauer B (2013) Synthesis of omega-hydroxy dodecanoic acid based on an engineered CYP153A fusion construct. Microb Biotechnol 6(6):694–707. doi:10.1111/1751-7915.12073
Steen EJ, Kang Y, Bokinsky G, Hu Z, Schirmer A, McClure A, Del Cardayre SB, Keasling JD (2010) Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature 463(7280):559–562. doi:10.1038/nature08721
Sung C, Jung E, Choi KY, Bae JH, Kim M, Kim J, Kim EJ, Kim PI, Kim BG (2015) The production of omega-hydroxy palmitic acid using fatty acid metabolism and cofactor optimization in Escherichia coli. Appl Microbiol Biotechnol. doi:10.1007/s00253-015-6630-1
Tijet N, Helvig C, Pinot F, Le Bouquin R, Lesot A, Durst F, Salaun JP, Benveniste I (1998) Functional expression in yeast and characterization of a clofibrate-inducible plant cytochrome P-450 (CYP94A1) involved in cutin monomers synthesis. Biochem J 332(Pt 2):583–589
van Beilen JB, Funhoff EG, van Loon A, Just A, Kaysser L, Bouza M, Holtackers R, Rothlisberger M, Li Z, Witholt B (2006) Cytochrome P450 alkane hydroxylases of the CYP153 family are common in alkane-degrading eubacteria lacking integral membrane alkane hydroxylases. Appl Environ Microbiol 72(1):59–65. doi:10.1128/AEM.72.1.59-65.2006
Vandamme EJ, Soetaert W (2002) Bioflavours and fragrances via fermentation and biocatalysis. J Chem Technol Biotechnol 77(12):1323–1332. doi:10.1002/Jctb.722
Wang X, Jin D, Zhou L, Wu L, An W, Zhao L (2014) Draft genome sequence of Gordonia alkanivorans strain CGMCC6845, a halotolerant hydrocarbon-degrading bacterium. Genome Announc 2(1). doi:10.1128/genomeA.01274-13
Woodley JM (2006) Microbial biocatalytic processes and their development. Adv Appl Microbiol 60:1–15. doi:10.1016/S0065-2164(06)60001-4
Yang W, Bell SG, Wang H, Zhou W, Hoskins N, Dale A, Bartlam M, Wong LL, Rao Z (2010) Molecular characterization of a class I P450 electron transfer system from Novosphingobium aromaticivorans DSM12444. J Biol Chem 285(35):27372–27384. doi:10.1074/jbc.M110.118349
Yokota TCONMCL, Okino HCONMCL, Takagi MCONMCL (1988) Process for producing omega-hydroxy fatty acids. Google Patents,
Young CC, Lin TC, Yeh MS, Shen FT, Chang JS (2005) Identification and kinetic characteristics of an indigenous diesel-degrading Gordonia alkanivorans strain. World J Microbiol Biotechnol 21(8–9):1409–1414. doi:10.1007/s11274-005-5742-7
Zimmer T, Ohkuma M, Ohta A, Takagi M, Schunck WH (1996) The CYP52 multigene family of Candida maltosa encodes functionally diverse n-alkane-inducible cytochromes P450. Biochem Biophys Res Commun 224(3):784–789. doi:10.1006/bbrc.1996.1100
Acknowledgments
This paper is dedicated to Romas Kazlauskas on the occasion of his 60th birthday. This study was funded by a grant of the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea. (HN12C0055) and Seoul R&BD Program (PA140024).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This paper does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that there is no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 305 kb)
Rights and permissions
About this article
Cite this article
Jung, E., Park, B.G., Ahsan, M.M. et al. Production of ω-hydroxy palmitic acid using CYP153A35 and comparison of cytochrome P450 electron transfer system in vivo. Appl Microbiol Biotechnol 100, 10375–10384 (2016). https://doi.org/10.1007/s00253-016-7675-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7675-5