Abstract
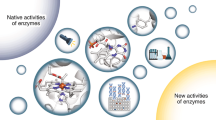
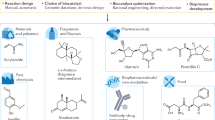
Biocatalysts (enzymes) have many advantages as catalysts for the production of useful compounds as compared to chemical catalysts. The stereoselectivity of the enzymes is one advantage, and thus the stereoselective production of chiral compounds using enzymes is a promising approach. Importantly, industrial application of the enzymes for chiral compound production requires the discovery of a novel useful enzyme or enzyme function; furthermore, improving the enzyme properties through protein engineering and directed evolution approaches is significant. In this review, the significance of several enzymes showing stereoselectivity (quinuclidinone reductase, aminoalcohol dehydrogenase, old yellow enzyme, and threonine aldolase) in chiral compound production is described, and the improvement of these enzymes using protein engineering and directed evolution approaches for further usability is discussed. Currently, enzymes are widely used as catalysts for the production of chiral compounds; however, for further use of enzymes in chiral compound production, improvement of enzymes should be more essential, as well as discovery of novel enzymes and enzyme functions.






Similar content being viewed by others
References
Baik SH, Yoshioka H, Yukawa H, Harayama S (2007) Synthesis of L-threo-3,4-dihydroxyphenylserine (L-threo-DOPS) with thermostabilized low-specific L-threonine aldolase from Streptomyces coelicolor A3(2). J Microbiol Biotechnol 17:721–727
Brown BJ, Deng Z, Karplus PA, Massey V (1998) On the active site of old yellow enzyme. Role of histidine 191 and asparagine 194. J Biol Chem 273:32753–32762
Dadashipur M, Ishida Y, Yamamoto K, Asano Y (2015) Discovery and molecular and biocatalytic properties of hydroxynitrile lyase from an invasive millipede, Chamberlinius hualienensis. Proc Natl Acad Sci U S A 112:10605–10610
di Salvo ML, Remesh SG, Vivoli M, Ghatge MS, Paiardini A, D’Aguanno S, Safo MK, Contestabile R (2014) On the catalytic mechanism and stereospecificity of Escherichia coli L-threonine aldolase. FEBS J. 281:129–145
Dücker N, Baer K, Simon S, Gröger H, Hummel W (2010) Threonine aldolases—screening, properties and applications in the synthesis of non-proteinogenic β-hydroxy-α-amino acids. Appl Microbiol Biotechnol 88:409–424
Farina V, Reeves JT, Senanayake CH, Song JJ (2006) Asymmetric synthesis of active pharmaceutical ingredients. Chem Rev 106:2734–2793
Guan L-J, Ohtsukal J, Okai M, Miyakawa T, Mase T, Zhi Y, Hou F, Ito N, Iwasaki A, Yasohara Y, Tanokura M (2015) A new target region for changing the substrate specificity of amine transaminases. Sci Rep 5:10753
Gwon HJ, Baik SH (2010) Diasteroselective synthesis of L-threo-3,4-dihydroxyphenylserine by low-specific L-threonine aldolase mutants. Biotechnol Lett 32:143–149
Hall M, Stueckler C, Ehammer H, Pointner E, Oberdorfer G, Gruber K, Hauer B, Stuermer R, Kroutil W, Macheroux P, Faber K (2008a) Asymmetric bioreduction of C=C bonds using enoate reductases POR1, OPR3 and YqjM: enzyme-based stereocontrol. Adv Synth Catal 350:411–418
Hall M, Stueckler C, Hauer B, Stuermer R, Friedrich T, Breuer M, Kroutil W, Faber K (2008b) Asymmetric bioreduction of activated C=C bonds using Zymomonas mobilis NCR enoate reductase and old yellow enzymes OYE 1-3 from yeasts. Eur J Org Chem 2008:1511–1516
Hibi M, Ogawa J (2014) Characteristics and biotechnology applications of aliphatic amino acid hydroxylases belonging to the Fe(II)/α-ketoglutarate-dependent dioxygenase superfamily. Appl Microbiol Biotechnol 98:3869–3876
Horinouchi N, Ogawa J, Kawano T, Sakai T, Saito K, Matsumoto S, Sasaki M, Mikami Y, Shimizu S (2006a) One-pot microbial synthesis of 2′-deoxyribonucleoside from glucose, acetaldehyde, and a nucleobase. Biotechnol Lett 28:877–881
Horinouchi N, Ogawa J, Kawano T, Sakai T, Saito K, Matsumoto S, Sasaki M, Mikami Y, Shimizu S (2006b) Efficient production of 2-deoxyribose 5-phosphate from glucose and acetoaldehyde by coupling of the alcoholic fermentation system of baker’s yeast and deoxyriboaldolase-expressing Escherichia coli. Biosci Biotechnol Biochem 70:1371–1378
Horinouchi N, Sakai T, Kawano T, Matsumoto S, Sasaki M, Hibi M, Shima J, Shimizu S, Ogawa J (2012) Construction of microbial platform for an energy-requiring bioprocess: practical 2′-deoxyribonucleoside production involving a C-C coupling reaction with high energy substrates. Microb Cell Factories 11:82
Horita S, Kataoka M, Kitamura N, Nakagawa T, Miyakawa T, Ohtsuka J, Nagata K, Shimizu S, Tanokura M (2015) An engineered old yellow enzyme that enables efficient synthesis of (4R,6R)-actinol in a one-pot reduction system. Chembiochem 16:440–445
Hou F, Miyakawa T, Kataoka M, Takeshita D, Kumashiro S, Uzura A, Urano N, Nagata K, Shimizu S, Tanokura M (2014) Structural basis for high substrate-binding affinity and enantioselectivity of 3-quinuclidinone reductase AtQR. Biochem Biophys Res Commun 446:911–915
Isotani K, Kurokawa J, Itoh N (2012) Production of (R)-3-quinuclidinol by E. coli biocatalysts possessing NADH-dependent 3-quinuclidinone reductase (QNR and bacC) from Microbacterium luteolum and Leifsonia alcohol dehydrogenase (LSADH). Int J Mol Sci 13:13542–13552
Isotani K, Kurokawa J, Suzuki F, Nomoto S, Negishi T, Matsuda M, Itoh N (2013) Gene cloning and characterization of two NADH-dependent 3-quinuclidinone reductases from Microbacterium luteolum JCM 9174. Appl Environ Microbiol 79:1378–1384
Jennewein S, Schürmann M, Wolberg M, Hilker I, Luiten R, Wubbolts M, Mink D (2006) Directed evolution of an industrial biocatalyst: 2-deoxy-D-ribose 5-phosphate aldolase. Biotechnol J 1:537–548
Jörnvall H, Persson B, Krrok M, Atrian S, Gonzàlez-Duarte R, Jefery J, Ghosh D (1995) Short-chain dehydrogenase/reductase (SDR). Biochemistry 34:6003–6013
Kataoka M, Shimizu K, Sakamoto K, Yamada H, Shimizu S (1995a) Optical resolution of racemic pantolactone with a novel fungal enzyme, lactonohydrolase. Appl Microbiol Biotechnol 43:974–977
Kataoka M, Shimizu K, Sakamoto K, Yamada H, Shimizu S (1995b) Lactonohydrolase-catalyzed optical resolution of pantoyl lactone: selection of a potent enzyme producer and optimization of culture and reaction conditions for practical resolution. Appl Microbiol Biotechnol 44:333–338
Kataoka M, Wada M, Nishi K, Yamada H, Shimizu S (1997a) Purification and characterization of L-allo-threonine aldolase from Aeromonas jandaei DK-39. FEMS Microbiol Lett 151:245–248
Kataoka M, Ikemi M, Morikawa T, Miyoshi T, Nishi K, Wada M, Yamada H, Shimizu S (1997b) Isolation and characterization of D-threonine aldolase, a pyridoxal-5′-phosphate-dependent enzyme from Arthrobacter sp. DK-38. Eur J Biochem 248:385–393
Kataoka M, Kotaka A, Hasegawa A, Wada M, Yoshizumi A, Nakamori S, Shimizu S (2002) Old yellow enzyme from Candida macedoniensis catalyzes stereospecific reduction of C=C bond of ketoisophorone. Biosci Biotechnol Biochem 66:2651–2657
Kataoka M, Kita K, Wada M, Yasohara Y, Hasegawa J, Shimizu S (2003) Novel bioreduction system for the production of chiral alcohols. Appl Microbiol Biotechnol 62:437–445
Kataoka M, Kotaka A, Thiwthong R, Wada M, Nakamori S, Shimizu S (2004) Cloning and overexpression of the old yellow enzyme gene of Candida macedoniensis, and its application to the production of a chiral compound. J Biotechnol 114:1–9
Kataoka M, Nakamura Y, Urano N, Ishige T, Shi G, Kita S, Sakamoto K, Shimizu S (2006) A novel NADP+-dependent L-1-amino-2-propanol dehydrogenase from Rhodococcus erythropolis MAK154: a promising enzyme for the production of double chiral aminoalcohols. Lett Appl Microbiol 43:430–435
Kataoka M, Ishige T, Urano N, Nakamura Y, Sakuradani E, Fukui S, Kita S, Sakamoto K, Shimizu S (2008) Cloning and expression of the L-1-amino-2-propanol dehydrogenase gene from Rhodococcus erythropolis, and its application to double chiral compound production. Appl Microbiol Biotechnol 80:597–604
Kielkopf CL, Burley SK (2002) X-ray structures of threonine aldolase complexes: structural basis of substrate recognition. Biochemistry 41:11711–11720
Kohli RM, Massey V (1998) The oxidative half-reaction of old yellow enzyme. The role of tyrosine 196. J Biol Chem 273:32763–32770
Kolet SP, Jadhav DD, Priyadarshini B, Swarge BN, Thulasiram HV (2014) Fungi mediated production and practical purification of (R)-3-quinuclidinol. Tetrahedron Lett 55:5911–5914
Liu JQ, Nagata S, Dairi T, Misono H, Shimizu S, Yamada H (1997a) The GLY1 gene of Saccharomyces cerevisiae encodes a low-specific L-threonine aldolase that catalyzes cleavage of L-allo-threonine and L-threonine to glycine: expression of the gene in Escherichia coli and purification and characterization of the enzyme. Eur J Biochem 245:289–293
Liu JQ, Dairi T, Kataoka M, Shimizu S, Yamada H (1997b) L-allo-threonine aldolase from Aeromonas jandaei DK-39: Gene cloning, nucleotide sequencing, and identification of the pyridoxal 5′-phosphate-binding lysine residue by site-directed mutagenesis. J Bacteriol 179:3555–3560
Liu JQ, Dairi T, Ito N, Kataoka M, Shimizu S, Yamada H (1998a) Gene cloning, biochemical characterization and physiological role of a thermostable low-specificity L-threonine aldolase from Escherichia coli. Eur J Biochem 255:220–226
Liu JQ, Ito S, Dairi T, Ito N, Kataoka M, Shimizu S, Yamada H (1998b) Gene cloning, nucleotide sequencing, and purification and characterization of the low-specificity L-threonine aldolase from Pseudomonas sp. strain NCIMB 10558. Appl Environ Microbiol 64:549–554
Liu JQ, Dairi T, Ito N, Kataoka M, Shimizu S, Yamada H (1998c) A novel metal-activated pyridoxal enzyme with a unique primary structure, low specificity D-threonine aldolase from Arthrobacter sp. strain DK-38. J Biol Chem 273:16678–16685
Liu JQ, Ito S, Dairi T, Itoh N, Shimizu S, Yamada H (1998d) Low-specificity L-threonine aldolase of Pseudomonas sp. NCIMB 10558: purification, characterization and its application to β-hydroxy-α-amino acid synthesis. Appl Microbiol Biotechnol 49:702–708
Liu JQ, Odani M, Dairi T, Itoh N, Shimizu S, Yamada H (1999) A new route to L-threo-3-[4-(methylthio)phenylserine], a key intermediate for the synthesis of antibiotics: recombinant low-specificity D-threonine aldolase-catalyzed stereospecific resolution. Appl Microbiol Biotechnol 51:586–591
Liu JQ, Odani M, Yasuoka M, Dairi T, Itoh N, Kataoka M, Shimizu S, Yamada H (2000) Gene cloning and overproduction of low-specificity D-threonine aldolase from Alcaligenes xylosoxidans and its application for production of a key intermediate for parkinsonism drug. Appl Microbiol Biotechnol 54:44–51
Liu JQ, Dairi T, Itoh N, Kataoka M, Shimizu S (2003) A novel enzyme, D-3-hydroxyaspartate aldolase from Paracoccus denitrificans IFO 13301: purification, characterization, and gene cloning. Appl Microbiol Biotechnol 62:53–60
Mikami M, Korenaga T, Ohkuma T, Noyori R (2000) Asymmetric activation/deactivation of racemic Ru catalysis for highly enantioselective hydrogenation of ketonic substrates. Angew Chem Int Ed Eng 39:3707–3710
Misono H, Maeda H, Tuda K, Ueshima S, Miyazaki N, Nagata S (2005) Characterization of an iducible phenylserine aldolase from Pseudomonas putida 24-1. Appl Environ Microbiol 71:4602–4609
Mitsukura K, Suzuki M, Tada K, Yoshida T, Nagasawa T (2010) Asymmetric synthesis of chiral cyclic amine from cyclic imine by bacterial whole-cell catalyst of enantioselective imine reductase. Org Biomol Chem 8:4533–4535
Mitsukura K, Suzuki M, Shinoda S, Kuramot T, Yoshida T, Nagasawa T (2011) Purification and characterization of a novel (R)-imine reductase from Streptomyces sp. GF3587. Biosci Biotechnol Biochem 75:1778–1782
Nomoto F, Hirayama Y, Ikunaka M, Inoue T, Otsuka K (2003) A practical chemoenzymatic process to access (R)-quinuclidin-3-ol on scale. Tetrahedron Asymmetry 14:1871–1877
Noyori R, Ohkuma T (2001) Asymmetric catalysis by architectural and functional molecular engineering: practical chemo- and stereoselective hydrogenation of ketones. Angew Chem Int Ed Eng 40:40–73
Ogawa J, Shimizu S (1997) Diversity and versatility of microbial hydantoin-transforming enzymes. J Mol Catal B Enzym 2:163–176
Padhi SK, Bougioukou DJ, Stewart JD (2009) Site-saturation mutagenesis of tryptophan 116 of Saccharomyces pastorianus old yellow enzyme uncovers stereocomplementary variants. J Am Chem Soc 131:3271–3280
Persson B, Krrok M, Jörnvall H (1991) Characteristics of short-chain alcohol dehydrogenases and related enzymes. Eur J Biochem 200:537–543
Qin HM, Imai FL, Miyakawa T, Kataoka M, Kitamura N, Urano N, Mori K, Kawabata H, Okai M, Ohtsuka J, Hou F, Nagata K, Shimizu S, Tanokura M (2014) L-allo-threonine aldolase with an H128Y/S292R mutation from Aeromonas jandaei DK-39 reveals the structural basis of changes in substrate stereoselectivity. Acta Crystallogr D Biol Crystallogr 70:1695–1703
Reß T, Hummel W, Hanlon SP, Iding H, Gröger H (2015) The organic-synthetic potential of recombinant ene reductases: substrate-scope evaluation and process optimization. ChemCatChem 7:1302–1311
Richter N, Gröger H, Hummel W (2011) Asymmetric reduction of activated alkenes using an enoate reductase from Gluconobacter oxydans. Appl Microbiol Biotechnol 89:79–89
Rosche B, Sandford V, Breuer M, Hauer B, Rogers PL (2002) Enhanced production of R-phenylacetylcarbinol (R-PAC) through enzymatic biotransformation. J Mol Catal B Enzym 19-20:109–115
Rustler S, Motejadded H, Altenbuchner J, Stolz A (2008) Simultaneous expression of an arylactonitrilase from Pseudomonas fluorescens and a (S)-oxynitrilase from Manihot esculenta in Pichia pastoris for the synthesis od (S)-mandelic acid. Appl Microbiol Biotechnol 80:87–97
Sakamoto K, Honda K, Wada K, Kita S, Tsuzaki K, Nose H, Kataoka M, Shimizu S (2005) Practical resolution system for DL-pantoyl lactone using the lactonase from Fusarium oxysporum. J Biotechnol 118:99–106
Savile CK, Janey JM, Mundorff EC, Moore JC, Tam S, Jarvis WR, Colbeck JC, Krebber A, Fleitz FJ, Brands J, Devine PN, Huisman GW, Hughes GJ (2010) Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science 329:305–309
Shimada Y, Yasuda S, Takahashi M, Hayashi T, Miyazawa N, Sato I, Abiru Y, Uchiyama S, Hishigaki H (2010) Cloning and expression of a novel NAD(P)H-dependent daidzein reductase, an enzyme involved in the metabolism of daidzein, from equol-producing Lactococcus strain 20-92. Appl Environ Microbiol 76:5892–5901
Steinreiber J, Fesko K, Reisinger C, Schürmann M, van Assema F, Wolberg M, Mink D, Griengl H (2007) Threonine aldolases—an emerging tool for organic synthesis. Tetrahedron 63:918–926
Sternbach LH, Kaiser S (1952) Antispasmodics. I. Bicyclic basic alcohols. J Am Chem Soc 74:2215–2218
Stuermer R, Hauer B, Hall M, Faber K (2007) Asymmetric bioreduction of activated C=C bonds using enoate reductases from the old yellow enzyme family. Curr Opin Chem Biol 11:203–213
Takeshita D, Kataoka M, Miyakawa T, Miyazono K, Kumashiro S, Nagai T, Urano N, Uzura A, Nagata K, Shimizu S, Tanokura M (2014) Structural basis of stereospecific reduction by quinuclidinone reductase. AMB Express 4:6
Toogood HS, Gardiner JM, Scrutton NS (2010) Biocatalytic reductions and chemical versatility of the old yellow enzyme family of flavoprotein oxidoreductases. ChemCatChem 2:892–914
Tsuji N, Honda K, Wada M, Okano K, Ohtake H (2014) Isolation and characterization of a thermotolerant ene reductase from Geobacillus sp. 30 and its heterologous expression in Rhodococcus opacus. Appl Microbiol Biotechnol 98:5925–5935
Uhl MK, Oberdorfer G, Steinkellner G, Riegler-Berket L, Mink D, van Assema F, Schürmann M, Gruber K (2015) The crystal structure of D-threonine aldolase from Alcaligenes xylosoxidans provides insight into a metal ion assisted PLP-dependent mechanism. PLoS One 10:e0124056
Urano N, Kataoka M, Ishige T, Kita S, Sakamoto K, Shimizu S (2011a) Genetic analysis around aminoalcohol dehydrogenase gene of Rhodococcus erythropolis MAK154: a putative GntR transcription factor in transcriptional regulation. Appl Microbiol Biotechnol 89:739–746
Urano N, Fukui S, Kumashiro S, Ishige T, Kita S, Sakamoto K, Kataoka M, Shimizu S (2011b) Directed evolution of an aminoalcohol dehydrogenase for efficient production of double chiral aminoalcohols. J Biosci Bioeng 111:266–271
Uzura A, Nomoto F, Sakoda A, Nishimoto Y, Kataoka M, Shimizu S (2009) Stereoselective synthesis of (R)-3-quinuclidinol through asymmetric reduction of 3-quinuclidinone with 3-quinuclidinone reductase of Rhodotorula rubra. Appl Microbiol Biotechnol 83:617–626
Wada M, Yoshizumi A, Noda Y, Kataoka M, Shimizu S, Takagi H, Nakamori S (2003) Production of a doubly chiral compound, (4R,6R)-4-hydroxy-2,2,6-trimethylcyclohexanone, by two-step enzymatic asymmetric reduction. Appl Environ Microbiol 69:933–937
Wang Y, Li J, Wu Q, Zhu D (2013) Microbial stereospecific reduction of 3-quinuclidinone with newly isolated Nocardia sp. and Rhodococcus erythropolis. J Mol Catal B Enzym 88:14–19
Winkler CK, Tasnádi G, Clay D, Hall M, Faber K (2012) Asymmetric bioreduction of activated alkenes to industrially relevant optically active compounds. J Biotechnol 162:381–389
Yamada H, Shimizu S, Shimada H, Tani Y, Takahashi S, Ohashi T (1980) Production of D-phenylglycine-related amino acids by immobilized microbial cells. Biochemie 62:395–399
Yoshizumi A, Wada M, Takagi H, Shimizu S, Nakamori S (2001) Cloning, sequence analysis, and expression in Escherichia coli of the gene encoding monovalent cation-activated levodione reductase from Corynebacterium aquaticum M-13. Biosci Biotechnol Biochem 65:830–836
Zhang WX, Xu GC, Huang L, Pan J, Yu HL, Xu JH (2013) Highly efficient synthesis of (R)-3-quinuclidinol in a space-time yield of 916 g L−1 d−1 using a new bacterial reductase ArQR. Org Lett 15:4917–4919
Acknowledgments
This review included the results obtained by the Targeted Proteins Research Program (TPRP) of the Ministry of Education, Culture, Sports, Science, and Technology of Japan (MEXT).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
This article does not contain studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Kataoka, M., Miyakawa, T., Shimizu, S. et al. Enzymes useful for chiral compound synthesis: structural biology, directed evolution, and protein engineering for industrial use. Appl Microbiol Biotechnol 100, 5747–5757 (2016). https://doi.org/10.1007/s00253-016-7603-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7603-8