Abstract
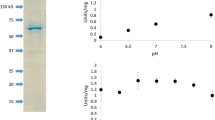
A gene from the thermophilic Gram-negative bacterium Rhodothermus marinus JCM9785, encoding a dye-linked d-amino acid dehydrogenase homologue, was overexpressed in Escherichia coli, and its product was purified and characterized. The expressed enzyme was a highly thermostable dye-linked d-amino acid dehydrogenase that retained more than 80 % of its activity after incubation for 10 min at up to 70 °C. When enzyme-catalyzed dehydrogenation of several d-amino acids was carried out using 2,6-dichloroindophenol as the electron acceptor, d-phenylalanine was the most preferable substrate among the d-amino acids tested. Immediately upstream of the dye-linked d-amino acid dehydrogenase gene (dadh) was a gene encoding a 4-hydroxyproline 2-epimerase homologue (hypE). That gene was successfully expressed in E. coli, and the gene product exhibited strong 4-hydroxyproline 2-epimerase activity. Reverse transcription PCR and quantitative real-time PCR showed that the six genes containing the dadh and hypE genes were arranged in an operon and were required for catabolism of trans-4-hydroxy-l-proline in R. marinus. This is the first description of a dye-linked d-amino acid dehydrogenase (Dye-DADH) with broad substrate specificity involved in trans-4-hydroxy-l-proline catabolism.






Similar content being viewed by others

References
Boulette ML, Baynham PJ, Jorth PA, Kukavica-Ibrulj I (2009) Characterization of alanine catabolism in Pseudomonas aeruginosa and its importance for proliferation in vivo. J Bacteriol 191:6329–6334
Cherkin A, Davis JL, Garman MW (1978) D-Proline: stereospecificity and sodium chloride dependence of lethal convulsant activity in the chick. Pharmacol Biochem Behav 8:623–625
Feng Z, Cáceres NE, Sarath G, Barletta RG (2002) Mycobacterium smegmatis L-alanine dehydrogenase (Ald) is required for proficient utilization of alanine as a sole nitrogen source and sustained anaerobic growth. J Bacteriol 184:5001–5010
Freese E, Oosterwyk J (1963) The induction of alanine dehydrogenase. Biochemistry 2:1212–1216
Frew JE, Hill HA (1987) Electrochemical biosensors. Anal Chem 59:933A–944A
Hamase K, Inoue T, Morikawa A, Konno R, Zaitsu K (2001) Determination of free D-proline and D-leucine in the brains of mutant mice lacking D-amino acid oxidase activity. Anal Biochem 298:253–258
He W, Li C, Lu CD (2011) Regulation and characterization of the dadRAX locus for D-amino acid catabolism in Pseudomonas aeruginosa PAO1. J Bacteriol 193:2107–2115
Jones H, Venables WA (1983) Effects of solubilisation on some properties of the membrane-bound respiratory enzyme D-amino acid dehydrogenase of Escherichia coli. FEBS Lett 151:189–192
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lobocka M, Henning J, Wild J, Klopotowski T (1994) Organization and expression of the Escherichia coli K-12 dad operon encoding the smaller subunit of D-amino acid dehydrogenase and the catabolic alanine racemase. J Bacteriol 176:1500–1510
Mathew E, Zhi J, Freundlich M (1996) Lrp is a direct repressor of the dad operon in Escherichia coli. J Bacteriol 178:7234–7240
Nagata Y, Higashi M, Ishii Y, Sano H, Tanigawa M, Nagata K, Noguchi K, Urade M (2006) The presence of high concentrations of free D-amino acids in human saliva. Life Sci 78:1677–1681
Ohshima T, Tanaka S (1993) Dye-linked L-malate dehydrogenase from thermophilic Bacillus species DSM 465. Purification and characterization. Eur J Biochem 214:37–42
Olsiewski JP, Kaczorowski GJ, Walsh C (1980) Purification and properties of D-amino acid dehydrogenase, an inducible membrane-bound iron-sulfur flavoenzyme from Escherichia coli B. J Biol Chem 255:4487–4494
Sakuraba H, Satomura T, Kawakami R, Yamamoto S, Kawarabayasi Y, Kikuchi H, Ohshima T (2002) L-aspartate oxidase is present in the anaerobic hyperthermophilic archaeon Pyrococcus horikoshii OT-3: characteristics and role in the de novo biosynthesis of nicotinamide adenine dinucleotide proposed by genome sequencing. Extremophiles 6:275–281
Satomura T, Kawakami R, Sakuraba H, Ohshima T (2002) Dye-linked D-proline dehydrogenase from hyperthermophilic archaeon Pyrobaculum islandicum is a novel FAD-dependent amino acid dehydrogenase. J Biol Chem 277:12861–12867
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tsukada K (1966) D-Amino acid dehydrogenases of Pseudomonas fluorescens. J Biol Chem 241:4522–4528
Watanabe S, Morimoto D, Fukumori F, Shinomiya H, Nishiwaki H, Kawano-Kawada M, Sasai Y, Watanabe Y, Tozawa Y (2012) Identification and characterization of D-hydroxyproline dehydrogenase and ∆1-pyrroline-4-hydroxy-2-carboxylate deaminase involved in novel L-hydroxyproline metabolism of bacteria: metabolic convergent evolution. J Biol Chem 287:32674–33288
Wild J, Walczak W, Krajewska-Grynkiewicz K, Klopotowski T (1974) D-amino acid dehydrogenase: the enzyme of the first step of D-histidine and D-methionine racemization in Salmonella typhimurium. Mol Gen Genet 128:131–146
Acknowledgments
This work was supported by a Grant-in-Aid for Young Scientific Research (B) to Takenori Satomura from the Japan Society for the Promotion of Science and a Grant-in-Aid for Scientific Research (A) to Toshihisa Ohshima.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Satomura, T., Ishikura, M., Koyanagi, T. et al. Dye-linked d-amino acid dehydrogenase from the thermophilic bacterium Rhodothermus marinus JCM9785: characteristics and role in trans-4-hydroxy-l-proline catabolism. Appl Microbiol Biotechnol 99, 4265–4275 (2015). https://doi.org/10.1007/s00253-014-6263-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6263-9