Abstract
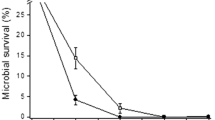
The Great Salt Plains (GSP), an unvegetated, barren salt flat that is part of the Salt Plains National Wildlife Refuge near Cherokee, Oklahoma, is the site of the Salt Plains Microbial Observatory. At the GSP the briny remains of an ancient sea rise to the surface, evaporate under dry conditions, and leave crusts of white salt. Adaptation to this environment requires development of coping mechanisms providing tolerance to desiccating conditions due to the high salinity, extreme temperatures, alkaline pH, unrelenting exposure to solar UV radiation, and prevailing winds. Several lines of evidence suggest that the same DNA repair mechanisms that are usually associated with UV light or chemically induced DNA damage are also important in protecting microbes from desiccation. Because little is known about the DNA repair capacity of microorganisms from hypersaline terrestrial environments, we explored the DNA repair capacity of microbial isolates from the GSP. We used survival following exposure to UV light as a convenient tool to assess DNA repair capacity. Two species of Halomonas (H. salina and H. venusta) that have been isolated repeatedly from the GSP were chosen for analysis. The survival profiles were compared to those of Escherichia coli, Pseudomonas aeruginosa, and Halomonas spp. from aquatic saline environments. Survival of GSP organisms exceeded that of the freshwater organism P. aeruginosa, although they survived no better than E. coli. The GSP isolates were much more resistance to killing by UV than were the aquatic species of Halomonas reported in the literature [Martin et al. (2000) Can J Microbiol 46:180−187]. Unlike E. coli, the GSP isolates did not appear to have an inducible, error-prone repair mechanism. However, they demonstrated high levels of spontaneous mutation.



Similar content being viewed by others
References
SF Altschul W Gish W Miller EW Myers DJ Lipman (1990) ArticleTitleBasic local alignment search tool. J Mol Biol 215 403–410 Occurrence Handle10.1006/jmbi.1990.9999 Occurrence Handle1:CAS:528:DyaK3MXitVGmsA%3D%3D Occurrence Handle2231712
HL Alder EB Roessler (1964) Introduction to Probability and Statistics, 3rd ed. W H Freeman San Francisco
JR Battista (1997) ArticleTitleAgainst all odds: the survival strategies of Deinococcus radiodurans. Ann Rev Microbiol 51 203–224 Occurrence Handle1:CAS:528:DyaK2sXms1Shsbg%3D
MG Booth WH Jeffrey RV Miller (2001) ArticleTitleRecA expression in response to solar UVR in the marine bacterium Vibrio natriegens. Microb Ecol 42 531–539 Occurrence Handle10.1007/s00248-001-1009-5 Occurrence Handle1:CAS:528:DC%2BD38XntFWqsw%3D%3D Occurrence Handle12024236
MG Booth L Hutchinson M Brumsted P Aas RB Coffin RC Downer Jr CA Kelley MM Lyons JD Pakuski SL Holder-Sandvik WH Jeffrey RV Miller (2001) ArticleTitleQuantification recA gene expression as an indicator of repair potential in marine bacterioplankton communities of Antarctica. Aquat. Microb Ecol 24 51–59
MA Buchheim JA Buchheim T Carlson P Kugrens (2002) ArticleTitlePhylogeny of Lobocharacium (Chlorophyceae) and allies: A study of 18S and 26S rDNA data. J Phycol 38 376–383 Occurrence Handle10.1046/j.1529-8817.2002.01145.x Occurrence Handle1:CAS:528:DC%2BD38XjvFChu7w%3D
TM Caton LR Witte HD Ngyuen JA Buchhein MA Buchheim MA Schneegurt (2004) ArticleTitleHalotolerant aerobic heterotrophic bacteria from the Great Salt Plains of Oklahoma. Microb Ecol 48(4) in press
JR Cole B Chai TL Marsh RJ Farris Q Wang SA Kulam S Chandra DM McGarrell TM Schmidt GM Garrity JM Tiedje (2003) ArticleTitleThe Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res 131 442–443 Occurrence Handle10.1093/nar/gkg039
U Edwards H Rogall H Blöcker M Emde EC Böttger (1989) ArticleTitleIsolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17 7843–7853 Occurrence Handle1:CAS:528:DyaK3cXjtFKksg%3D%3D Occurrence Handle2798131
SJ Elledge GC Walker (1983) ArticleTitleThe muc genes of pKM101 are induced by DNA damage. J Bacterial 155 1306–1315 Occurrence Handle1:CAS:528:DyaL3sXlsFejtr8%3D
WH Jeffrey RJ Pledger P Aas S Hager RB Coffin R Von Haven DL Mitchell (1996) ArticleTitleDiel and depth profiles of DNA photodamage in bacterioplankton exposed to ambient solar ultraviolet radiation. Mar Ecol Prog Ser 137 283–291 Occurrence Handle1:CAS:528:DyaK28XlsVSmsro%3D
WK Joklik HP Willett DB Amos CM Wilfert (1992) Zinsser Microbiology, 20th ed. Appleton & Mange Norwalk, CT
SP Kidambi MG Booth TA Kokjohn RV Miller (1996) ArticleTitlerecA-dependence of the response of Pseudomonas aeruginosa to UVA and UVB irradiation. Microbiology 142 1033–1040 Occurrence Handle1:CAS:528:DyaK28Xis1GltLw%3D Occurrence Handle8936329
TA Kokjohn RV Miller (1985) ArticleTitleMolecular cloning and characterization of the recA gene of Pseudomonas aeruginosa. J Bacteriol l63 568–572
TA Kokjohn RV Miller (1994) ArticleTitleIncN plasmids mediate UV resistance and error-prone repair in Pseudomonas aeruginosa PAO. Microbiology 140 43–48 Occurrence Handle1:CAS:528:DyaK2cXivFWnsrY%3D Occurrence Handle8162189
DR Maddison WP Maddison (2000) MacClade4: Analysis of Phylogeny and Character Evolution Sinauer Associates Sunderland, MA
EL Martin RL Reinhardt LL Baum MR Becker JJ Shaffer TA Kokjohn (2000) ArticleTitleThe effects of ultraviolet radiation on the moderate halophile Halomonas elongata and the extreme halophile Halobacterium salinarum. Can J Microbiol 46 180–187 Occurrence Handle10.1139/cjm-46-2-180 Occurrence Handle1:CAS:528:DC%2BD3cXhtlSqsLs%3D Occurrence Handle10721487
RV Miller (2000) recA: The gene and its protein product. S Luria (Eds) Encyclopedia of Microbiology, 2nd ed. Academic Press San Diego 43–54
RV Miller JM Decker (1978) ArticleTitlePeptidases of Pseudomonas aeruginosa: evidence for membrane association. J Bacteriol 133 165–171 Occurrence Handle1:CAS:528:DyaE1cXosFKltw%3D%3D Occurrence Handle412832
RV Miller TA Kokjohn (1990) ArticleTitleGeneral Microbiology of recA: Environmental and evolutionary significance. Annl Rev Microbiol 44 365–394 Occurrence Handle1:CAS:528:DyaK3MXnvVagug%3D%3D
RV Miller C-MC Ku (1978) ArticleTitleCharacterization of Pseudomonas aeruginosa mutants deficient in the establishment of lysogeny. J Bacteriol 134 875–883 Occurrence Handle1:STN:280:CSeC1MvgtVA%3D Occurrence Handle96103
RV Miller W Jeffrey D Mitchell M Elasri (1999) ArticleTitleBacterial response to solar ultraviolet light. ASM News 65 535–541
FJ Mondello RV Miller (1984) ArticleTitleIdentification of Pseudomonas plasmids able to suppress the lysogeny-establishment-deficiency (Les−) phenotype. Plasmid 11 185–187 Occurrence Handle1:CAS:528:DyaL2cXktFyht70%3D Occurrence Handle6427800
D Posada KA Crandall (1998) ArticleTitleMODELTEST: testing the model of DNA substitution. Bioinformatics 14 817–818 Occurrence Handle10.1093/bioinformatics/14.9.817 Occurrence Handle1:CAS:528:DyaK1MXktlCltw%3D%3D Occurrence Handle9918953
E Quesada A Ventosa F Rodriguez-Valera A Ramos-Cormenzana (1983) ArticleTitleNumerical taxonomy of moderately halophilic Gram-negative bacteria from hypersaline soils. J Gen Microbiol 129 2649–2657
AJ Rattray JN Strathern (2003) ArticleTitleError-prone DNA polymerases: when making a mistake is the only way to get ahead. Ann Rev Genet 37 31–66 Occurrence Handle10.1146/annurev.genet.37.042203.132748 Occurrence Handle1:CAS:528:DC%2BD2cXhsFGltg%3D%3D Occurrence Handle14616055
F Rodriquez-Valera R Riuz-Berraquero A Ramos-Cormenzana (1980) ArticleTitleIsolation of extremely halophilic bacteria able to grow in defined organic media with single carbon sources. J Gen Microbiol 119 535–538
SG Sedgwick PA Goodwin (1985) ArticleTitleDifferences in mutagenic and recombinational DNA repair in enterobacteria. Proc Natl Acad Sci USA 82 4172–4176
CS Simonson TA Kokjohn RV Miller (1990) ArticleTitleInducible UV repair potential of Pseudomonas aeruginosa PAO. J Gen Microbiol 136 1241–1249 Occurrence Handle1:CAS:528:DyaK3cXltFWlsbs%3D Occurrence Handle2121899
BT Smith GC Walker (1998) ArticleTitleMutagenesis and more: umuDC and the Escherichia coli SOS response. Genetics 148 1599–1610 Occurrence Handle1:CAS:528:DyaK1cXks1eiur4%3D Occurrence Handle9560379
Swofford, DL (2002) PAUP*: Phylogenetic Analysis using Parsimony, Version 4.0b10. Sinauer Associates, Sunderland, MA
GC Walker (1984) ArticleTitleMutagenesis and inducible responses to deoxyribonucleic acid damage in Escherichia coli. Microbiol Rev 48 60–93 Occurrence Handle1:CAS:528:DyaL2cXhslWhsL4%3D Occurrence Handle6371470
Acknowledgment
We thank W. Warde of the Statistics Department at Oklahoma State University for assistance in statistical analysis and W. Henley and A. Kirkwood for insightful discussions. Weather data were provided free of charge from the Oklahoma Mesonet, managed by the University of Oklahoma and Oklahoma State University. Primary support for this work was supported by the Microbial Observatories program of the National Science Foundation (MCB-0132097, MCB-0131659, and MCB-132083). Additional support was provided by grants from the Kansas Biomedical Research Infrastructure Network (KBRIN), the Office of Research Administration at Wichita State University, the Mervin Bovaird Institute for Molecular Biology and Biotechnology at the University of Tulsa, and the Oklahoma EPSCoR program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wilson, C., Caton, T., Buchheim, J. et al. DNA-Repair Potential of Halomonas spp. from the Salt Plains Microbial Observatory of Oklahoma. Microb Ecol 48, 541–549 (2004). https://doi.org/10.1007/s00248-004-0243-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-004-0243-z