Abstract
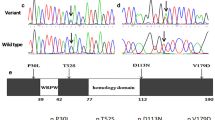
Conotruncal heart defects (CTDs) represent a group of severe and complicated congenital cardiovascular malformations and require opportune clinical interventions once diagnosed. Occurrence of CTD is related to the functional abnormality of the second heart field (SHF), and variants of genes which regulate the development of the second heart field have been recognized as the main genetic factors leading to CTDs. Previous studies indicated that transcriptional complex SIX1/EYA1 may contribute to SHF development, and SIX1/EYA1 knockout mice exhibited a series of conotruncal malformations. Here, we recruited and sequenced 600 Chinese conotruncal heart defect patients and 300 controls. We screened out one novel SIX1 mutation (SIX1-K134R) and four EYA1 rare mutations (EYA1-A227T, EYA1-R296H, EYA1-Q397R, EYA1-G426S), all variants were present only in the case cohort, and the mutated sites were highly conserved. We then analyzed mutations by software including Sift, PolyPhen-2, PROVEAN, Mutation Taster, HOPE, and SWISS-PdbViewer. The results showed that the mutations had varying degrees of pathogenic risk, protein properties, spatial conformations, and domain functions which might be altered or influenced. Through biological and in silico analyses, our study suggests an association between SIX1/EYA1 mutations and cardiovascular malformations, SIX1/EYA1 mutations might be partially responsible for CTDs.


Similar content being viewed by others
References
De Luca A, Sarkozy A, Ferese R et al (2011) New mutations in ZFPM2/FOG2 gene in tetralogy of Fallot and double outlet right ventricle. Clin Genet 80:184–190
Bentham J, Bhattacharya S (2008) Genetic mechanisms controlling cardiovascular development. Ann N Y Acad Sci 1123:10–19
Cheyette BN, Green PJ, Martin K, Garren H, Hartenstein V, Zipursky SL (1994) The Drosophila sine oculis locus encodes a homeodomain-containing protein required for the development of the entire visual system. Neuron 12:977–996
Li X, Oghi KA, Zhang J et al (2003) Eya protein phosphatase activity regulates Six1-Dach-Eya transcriptional effects in mammalian organogenesis. Nature 426:247–254
Rayapureddi JP, Kattamuri C, Steinmetz BD et al (2003) Eyes absent represents a class of protein tyrosine phosphatases. Nature 426:295–298
Vincent C, Kalatzis V, Abdelhak S et al (1997) BOR and BO syndromes are allelic defects of EYA1. Eur J Hum Genet 5:242–246
Wang SH, Wu CC, Lu YC et al (2012) Mutation screening of the EYA1, SIX1, and SIX5 genes in an East Asian cohort with branchio-oto-renal syndrome. Laryngoscope 122:1130–1136
Guo C, Sun Y, Zhou B et al (2011) A Tbx1-Six1/Eya1-Fgf8 genetic pathway controls mammalian cardiovascular and craniofacial morphogenesis. J Clin Investig 121:1585–1595
Shimasaki N, Watanabe K, Hara M, Kosaki K (2004) EYA1 mutation in a newborn female presenting with cardiofacial syndrome. Pediatr Cardiol 25:411–413
Blue GM, Humphreys D, Szot J et al (2017) The promises and challenges of exome sequencing in familial, non-syndromic congenital heart disease. Int J Cardiol 230:155–163
Solayman M, Saleh MA, Paul S, Khalil MI, Gan SH (2017) In silico analysis of nonsynonymous single nucleotide polymorphisms of the human adiponectin receptor 2 (ADIPOR2) gene. Comput Biol Chem 68:175–185
Venselaar H, Te Beek TA, Kuipers RK, Hekkelman ML, Vriend G (2010) Protein structure analysis of mutations causing inheritable diseases. An e-Science approach with life scientist friendly interfaces. BMC Bioinform 11:548
Li B, Pu T, Liu Y, Xu Y, Xu R (2017) CITED2 mutations in conserved regions contribute to conotruncal heart defects in Chinese children. DNA Cell Biol. doi:10.1089/dna.2017.3701
Hoffman JI, Kaplan S (2002) The incidence of congenital heart disease. J Am Coll Cardiol 39:1890–1900
Nakajima Y (2010) Second lineage of heart forming region provides new understanding of conotruncal heart defects. Congenit Anom 50:8–14
Ilagan R, Abu-Issa R, Brown D et al (2006) Fgf8 is required for anterior heart field development. Development 133:2435–2445
Pacheco-Leyva I, Matias AC, Oliveira DV et al (2016) CITED2 cooperates with ISL1 and promotes cardiac differentiation of mouse embryonic stem cells. Stem Cell Rep 7:1037–1049
Baldini A, Fulcoli FG, Illingworth E (2017) Tbx1: transcriptional and developmental functions. Curr Top Dev Biol 122:223–243
Xu PX, Woo I, Her H, Beier DR, Maas RL (1997) Mouse Eya homologues of the Drosophila eyes absent gene require Pax6 for expression in lens and nasal placode. Development 124:219–231
Chen B, Kim EH, Xu PX (2009) Initiation of olfactory placode development and neurogenesis is blocked in mice lacking both Six1 and Six4. Dev Biol 326:75–85
Rayapureddi JP, Hegde RS (2006) Branchio-oto-renal syndrome associated mutations in eyes absent 1 result in loss of phosphatase activity. FEBS Lett 580:3853–3859
Acknowledgements
The study was supported by the National Natural Science Foundation of China (81270233) and the Three Years’ Action of Shanghai Health Bureau (GWTV-23).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declared that they have no conflicts of interest.
Ethical Approval
This research conformed to the ethical guidelines of the 2013 Declaration of Helsinki and was approved by local ethics committee of Xinhua Hospital.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Li, B., Xu, L., Hong, N. et al. In Silico Analyses Reveal the Relationship Between SIX1/EYA1 Mutations and Conotruncal Heart Defects. Pediatr Cardiol 39, 176–182 (2018). https://doi.org/10.1007/s00246-017-1744-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00246-017-1744-0