Abstract
Heparinoids are the starting material for sulodexide production, a drug used as intravenous anti-coagulant, as an alternative to heparin. The origin determination in the starting material for sulodexide, heparin, and derivatives is crucial for safety (including the impact related to bovine spongiform encephalopathy) and efficacy of the final products. Therefore, European countries have decided to approve the production of heparin only from porcine intestinal mucosa. PCR (polymerase chain reaction) methods are available to evaluate the origin species of crude heparin, during heparin production process, while they lack for the same analysis in heparinoids during sulodexide manufacturing processes. Notably, two main critical issues occur during the origin determination by using PCR for heparinoid analysis: first, heparin has been known to inhibit DNA polymerase activity and, second, the DNA amounts are very low in these samples. To overcome these critical issues, our proposed method is based on two fundamental steps, the DNA concentration by glycogen treatment and DNA purification, which occur before and after DNA extraction, respectively. Finally, by applying real-time PCR, we amplify three specific DNA sequences of ruminant species (bovine, ovine, and caprine), to assess possible contamination, and one from swine, to confirm the origin species. To date, such a method is the only one that determines origin species by PCR for heparinoids that guarantee quality, safety, and traceability of heparin-derived pharmaceutical products. In conclusion, our proposed method is an alternative to nuclear magnetic resonance and ELISA methods, because real-time PCR offers significant advantages in sensitivity, specificity, and robustness.

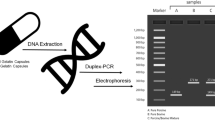
Graphical Abstract






Similar content being viewed by others
References
Auguste C, Dereux S, Rousset M, Anger P. Validation of quantitative polymerase chain reaction methodology for monitoring DNA as a surrogate marker for species material contamination in porcine heparin. Anal Bioanal Chem. 2012;404(1):43–50. https://doi.org/10.1007/s00216-012-6085-5.
Authority of the United States Pharmacopeial Convention. Heparin sodium. In: US Pharmacopeia National Formulary. Rockville; 2019. p. 2148–2153; Volume 1.
U.S. Department of Health and Human Services, Food and Drug Administration, Center for Drug Evaluation and Research, Center for Veterinary Medicine, Center for Devices and Radiological Health. Guidance for Industry. Heparin for drug and medical device use: monitoring crude heparin for quality. 2013. https://www.fda.gov/media/82924/download. Accessed May 2019.
European Directorate for the Quality of Medicine & HealthCare. Heparin sodium. In: European Pharmacopoeia 9.0. Strasbourg; 2017. p. 2644–2646; Volume II.
Bouchard O, Hoppensteadt D, Maia P, de Castro AS, Kumar E, Guler N, et al. Porcine and ovine mucosal heparins and their depolymerized derivatives are comparable in contrast to their bovine equivalents. Blood. 2016;128(22):5027.
Carroll BJ, Piazza G, Goldhaber SZ. Sulodexide in venous disease. J Thromb Haemost. 2019;17(1):31–8. https://doi.org/10.1111/jth.14324.
Veraldi N, Guerrini M, Urso E, Risi G, Bertini S, Bensi D, et al. Fine structural characterization of sulodexide. J Pharm Biomed Anal. 2018;156:67–79. https://doi.org/10.1016/j.jpba.2018.04.012.
Coccheri S, Mannello F. Development and use of sulodexide in vascular diseases: implications for treatment. Drug Des Devel Ther. 2013;8:49–65. https://doi.org/10.2147/DDDT.S6762.
Concannon SP, Wimberley PB, Workman WE. A quantitative PCR method to quantify ruminant DNA in porcine crude heparin. Anal Bioanal Chem. 2011;399(2):757–62. https://doi.org/10.1007/s00216-010-4362-8.
Tanabe S, Hase M, Yano T, Sato M, Fujimura T, Akiyama H. A real-time quantitative PCR detection method for pork, chicken, beef, mutton, and horseflesh in foods. Biosci Biotechnol Biochem. 2007;71(12):3131–5. https://doi.org/10.1271/bbb.70683.
Auguste C, Dereux S, Martinez C, Anger P. New developments in quantitative polymerase chain reaction applied to control the quality of heparins. Anal Bioanal Chem. 2011;399(2):747–55. https://doi.org/10.1007/s00216-010-4232-4.
Sakai Y, Kotoura S, Yano T, Kurihara T, Uchida K, Miake K, et al. Quantification of pork, chicken and beef by using a novel reference molecule. Biosci Biotechnol Biochem. 2011;75(9):1639–43. https://doi.org/10.1271/bbb.110024.
Ding M, Bullotta A, Caruso L, Gupta P, Rinaldo CR, Chen Y. An optimized sensitive method for quantitation of DNA/RNA viruses in heparinized and cryopreserved plasma. J Virol Methods. 2011;176(1-2):1–8. https://doi.org/10.1016/j.jviromet.2011.05.012.
Huang Q, Yao CY, Chen B, Wang F, Huang JF, Zhang X, et al. Species-specific identification by inhibitor-controlled PCR of ruminant components contaminating industrial crude porcine heparin. Mol Cell Probes. 2006;20(3-4):250–8. https://doi.org/10.1016/j.mcp.2006.01.005.
Yokota M, Tatsumi N, Nathalang O, Yamada T, Tsuda I. Effects of heparin on polymerase chain reaction for blood white cells. J Clin Lab Anal. 1999;13(3):133–40.
European Directorate for the Quality of Medicine & HealthCare. Nucleic acid amplification techniques. In: European Pharmacopoeia 9.0. Strasbourg; 2017. p. 214–219; Volume I.
Houiste C, Auguste C, Macrez C, Dereux S, Derouet A, Anger P. Quantitative PCR and disaccharide profiling to characterize the animal origin of low-molecular-weight heparins. Clin Appl Thromb Hemost. 2009;15(1):50–8. https://doi.org/10.1177/1076029608320831.
Tovar AM, Santos GR, Capille NV, Piquet AA, Glauser BF, Pereira MS, et al. Structural and haemostatic features of pharmaceutical heparins from different animal sources: challenges to define thresholds separating distinct drugs. Sci Rep. 2016;6:35619. https://doi.org/10.1038/srep35619.
Beutler E, Gelbart T, Kuhl W. Interference of heparin with the polymerase chain reaction. Biotechniques. 1990;9(2):166.
Helms C, Graham MY, Dutchik JE, Olson MV. A new method for purifying lambda DNA from phage lysates. DNA. 1985;4(1):39–49. https://doi.org/10.1089/dna.1985.4.39.
Hengen PN. Carriers for precipitating nucleic acids. Trends Biochem Sci. 1996;21(6):224–5.
Tracy S. Improved rapid methodology for the isolation of nucleic acids from agarose gels. Prep Biochem. 1981;11(3):251–68. https://doi.org/10.1080/00327488108061767.
Ekins J, Peters SM, Jones YL, Swaim H, Ha T, La Neve F, et al. Development of a multiplex real-time PCR assay for the detection of ruminant DNA. J Food Prot. 2012;75(6):1107–12. https://doi.org/10.4315/0362-028X.JFP-11-415.
Acknowledgements
The research was performed in the Quality Control and R&D laboratories of Biofer S.p.A., Medolla, Italy. The authors would like to thank Senior Management of the company, and particularly, Dr. Alessandro Lapini Sacchetti and Dr. Gianmaria Ristori for their support, endorsement, and extensive knowledge in the field of this class of active substances.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 745 kb)
Rights and permissions
About this article
Cite this article
Pecorini, S., Camurri, G., Torrini, L. et al. Highly sensitive real-time PCR method to identify species origin in heparinoids. Anal Bioanal Chem 412, 289–298 (2020). https://doi.org/10.1007/s00216-019-02235-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-02235-w