Abstract
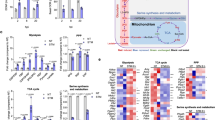
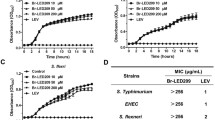
Salmonella enterica serovar Typhimurium (S. Typhimurium) is a zoonotic pathogen that causes severe gastroenteritis. The 5′-nucleotidases of pathogens can dephosphorylate adenosine phosphates, boost adenosine levels and suppress the pro-inflammatory immune response. In our previous study, an extracellular nuclease, 5′-nucleotidase, was identified in the extracellular proteins of S. Typhimurium. However, the nuclease activity and the function of the 5′-nucleotidase of S. Typhimurium have not been explored. In the present study, deletion of the 5′-nucleotidase gene is dispensable for S. Typhimurium growth, even under environmental stress. Fluorescence microscopy revealed that the 5′-nucleotidase mutant induced more macrophage extracellular traps (METs) than the wild type did. Furthermore, recombinant 5′-nucleotidase protein (r5Nuc) could degrade λDNA, and the nuclease activity of r5Nuc was optimum at 37 °C and pH 6.0–7.0. The Mg2+ enhanced the nuclease activity of r5Nuc, whereas Zn2+ inhibited it. Meanwhile, deletion of the 5′-nucleotidase gene increased the bactericidal activity of METs, and r5Nuc could degrade METs and inhibit the bactericidal activity of METs. In conclusion, S. Typhimurium growth was independent of 5′-nucleotidase, but the nuclease activity of 5′-nucleotidase assisted S. Typhimurium to evade macrophage-mediated extracellular killing through degrading METs.








Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Afonso M, Mestre AR, Silva G et al (2021) Candida extracellular nucleotide metabolism promotes neutrophils extracellular traps escape. Front Cell Infect Microbiol 11:678568. https://doi.org/10.3389/fcimb.2021.6785
Brinkmann V, Reichard U, Goosmann C et al (2004) Neutrophil extracellular traps kill bacteria. Science 303(5663):1532–1535. https://doi.org/10.1126/science.1092385
Buhr ND, Neumann A, Jerjomiceva N et al (2014) Streptococcus suis DNase Ssna contributes to degradation of neutrophil extracellular traps (NETs) and evasion of NET-mediated antimicrobial activity. Microbiology (reading) 160:385–395. https://doi.org/10.1099/mic.0.072199-0
Chang A, Khemlani A, Kang H et al (2011) Functional analysis of Streptococcus pyogenes nuclease A (Spna), a novel group A streptococcal virulence factor. Mol Microbiol 79(6):1629–1642. https://doi.org/10.1111/j.1365-2958.2011.07550.x
Dangel ML, Dettmann JC, Haßelbarth S et al (2019) The 5′-nucleotidase S5na is dispensable for evasion of phagocytosis and biofilm formation in Streptococcus pyogenes. PLoS ONE 14(1):e0211074. https://doi.org/10.1111/j.1365-2958.2011.07550.x
Derré-Bobillot A, Cortes-Perez NG, Yamamoto Y et al (2013) Nuclease A (Gbs0661), an extracellular nuclease of Streptococcus agalactiae, attacks the neutrophil extracellular traps and is needed for full virulence. Mol Microbiol 89(3):518–531. https://doi.org/10.1111/mmi.12295
Dos Santos AMP, Ferrari RG, Conte-Junior CA (2019) Virulence factors in Salmonella Typhimurium: the sagacity of a bacterium. Curr Microbiol 76(6):762–773. https://doi.org/10.1007/s00284-018-1510-4
Fan J, Zhang Y, Chuang-Smith ON et al (2012) Ecto-5′-nucleotidase: a candidate virulence factor in Streptococcus sanguinis experimental endocarditis. PLoS ONE 7(6):e38059. https://doi.org/10.1371/journal.pone.0038059
Galán JE (2021) Salmonella Typhimurium and inflammation: a pathogen-centric affair. Nat Rev Microbiol 19(11):716–725. https://doi.org/10.1038/s41579-021-00561-4
Halder LD, Abdelfatah MA, Jo EA et al (2017) Factor H binds to extracellular DNA traps released from human blood monocytes in response to Candida albicans. Front Immunol 7:671. https://doi.org/10.3389/fimmu.2016.00671
Heuts DPH, Weissenborn MJ, Olkhov RV et al (2012) Crystal structure of a soluble form of human CD73 with ecto-5′-nucleotidase activity. ChemBioChem 13(16):2384–2391. https://doi.org/10.1002/cbic.201200426
Innes D, Beacham IR, Burns DM (2001) The role of the intracellular inhibitor of periplasmic UDP-sugar hydrolase (5′-nucleotidase) in Escherichia coli: cytoplasmic localisation of 5′-nucleotidase is conditionally lethal. J Basic Microbiol 41(6):329–337. https://doi.org/10.1002/1521-4028(200112)41:6%3c329::AID-JOBM329%3e3.0.CO;2-4
King PT, Sharma R, O′Sullivan K et al (2015) Nontypeable Haemophilus Influenzae induces sustained lung oxidative stress and protease expression. PLoS ONE 10(3):e0120371. https://doi.org/10.1371/journal.pone.0120371
Kurtz JR, Goggins JA, McLachlan JB (2017) Salmonella infection: interplay between the bacteria and host immune system. Immunol Lett 190:42–50. https://doi.org/10.1016/2Fj.imlet.2017.07.006
Liao C, Zhang M, Cheng X et al (2020) Identification and characterization of the nuclease activity of the extracellular proteins from Salmonella enterica serovar Typhimurium. Curr Microbiol 77(11):3651–3660. https://doi.org/10.1007/s00284-020-02201-1
Liu P, Wu X, Liao C et al (2014) Escherichia coli and Candida albicans induced macrophage extracellular trap-like structures with limited microbicidal activity. PLoS ONE 9(2):e90042. https://doi.org/10.1371/journal.pone.0090042
Loureiro A, Pais C, Sampaio P (2019) Relevance of macrophage extracellular traps in C. albicans killing. Front Immunol 10:2767. https://doi.org/10.3389/fimmu.2019.02767
Ma F, Guo X, Fan H (2016) Extracellular nucleases of Streptococcus equi subsp. zooepidemicus degrade neutrophil extracellular traps and impair macrophage activity of the host. Appl Environ Microbiol 83(2):e02468-e2516. https://doi.org/10.1128/AEM.02468-16
Mitiku F, Hartley CA, Sansom FM et al (2018) The major membrane nuclease Mnua degrades neutrophil extracellular traps induced by Mycoplasma bovis. Vet Microbiol 218:13–19. https://doi.org/10.1016/j.vetmic.2018.03.002
Mónaco A, Canales-Huerta N, Jara-Wilde J et al (2021) Salmonella Typhimurium triggers extracellular traps release in murine macrophages. Front Cell Infect Microbiol 11:639768. https://doi.org/10.3389/fcimb.2021.639768
Morita C, Sumioka R, Nakata M et al (2014) Cell wall-anchored nuclease of Streptococcus sanguinis contributes to escape from neutrophil extracellular trap-mediated bacteriocidal activity. PLoS ONE 9(8):e103125. https://doi.org/10.1371/journal.pone.0103125
Patra SD, Mohakud NK, Panda RK et al (2021) Prevalence and multidrug resistance in Salmonella enterica Typhimurium: an overview in south east Asia. World J Microbiol Biotechnol 37(11):185. https://doi.org/10.1007/s11274-021-03146-8
Rittmann D, Sorger-Herrmann U, Wendisch VF (2005) Phosphate starvation-inducible gene ushA encodes a 5′ nucleotidase required for growth of Corynebacterium glutamicum on medium with nucleotides as the phosphorus source. Appl Environ Microbiol 71(8):4339–4344. https://doi.org/10.1128/AEM.71.8.4339-4344.2005
Rungelrath V, Kobayashi SD, DeLeo FR (2020) Neutrophils in innate immunity and systems biology-level approaches. Wiley Interdiscip Rev Syst Biol Med 12(1):e1458. https://doi.org/10.1002/wsbm.1458
Sharma P, Garg N, Sharma A et al (2019) Nucleases of bacterial pathogens as virulence factors, therapeutic targets and diagnostic markers. Int J Med Microbiol 309(8):151354. https://doi.org/10.1016/j.ijmm.2019.151354
Soh KY, Loh JMS, Proft T (2020) Cell wall-anchored 5′-nucleotidases in gram-positive cocci. Mol Microbiol 113(4):691–698. https://doi.org/10.1111/mmi.14442
Thammavongsa V, Kern JW, Missiakas DM et al (2009) Staphylococcus aureus synthesizes adenosine to escape host immune responses. J Exp Med 206(11):2417–2427. https://doi.org/10.1084/jem.20090097
Thammavongsa V, Schneewind O, Missiakas DM (2011) Enzymatic properties of Staphylococcus aureus adenosine synthase (AdsA). BMC Biochem 12:56. https://doi.org/10.1186/1471-2091-12-56
Zakataeva NP (2021) Microbial 5′-nucleotidases: their characteristics, roles in cellular metabolism, and possible practical applications. Appl Microbiol Biotechnol 105(20):7661–7681. https://doi.org/10.1007/s00253-021-11547-w
Zheng L, Khemlani A, Lorenz N et al (2015) Streptococcal 5′-nucleotidase A (S5na), a novel Streptococcus pyogenes virulence factor that facilitates immune evasion. J Biol Chem 290(52):31126–31137. https://doi.org/10.1074/jbc.M115.677443
Zimmermann H (1992) 5′-Nucleotidase: molecular structure and functional aspects. Biochem J 285:345–365. https://doi.org/10.1042/2Fbj2850345
Acknowledgements
This work was supported by the youth backbone teachers training program of Henan University of Science and Technology (13450009). We thank Ningbo freescience Information Technology Co., Ltd. for editing the manuscript.
Funding
The youth backbone teachers training program of Henan University of Science and Technology, 13450009.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Man Qian, Ke Xu, Mengke Zhang, Junhui Niu, Tianxiang Zhao. The first draft of the manuscript was written by Man Qian and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Consent for publication
All the authors read and consent to publish this manuscript.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qian, M., Xu, K., Zhang, M. et al. 5′-Nucleotidase is dispensable for the growth of Salmonella Typhimurium but inhibits the bactericidal activity of macrophage extracellular traps. Arch Microbiol 205, 20 (2023). https://doi.org/10.1007/s00203-022-03353-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-03353-3