Abstract
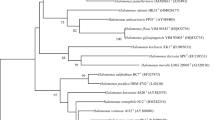
A novel extremely halophilic archaeon, designated WN019T, was isolated from the natural saline-alkali wetland soil of Binhai new district, Tianjin, China. Cells of WN019T were aerobic, motile, and pleomorphic rod-shaped, 0.5–0.8 µm in width and 2.0–2.5 µm in length, and the growth occurred optimally at 33–37 °C, pH 7.5–8.0, and in the presence of 15.0–20.0% (w/v) NaCl. Phylogenetic analyses based on 16S rRNA gene sequence comparison showed that the isolate belonged to the genus Halorubrum and exhibited moderate sequence similarity of 97.8% to Halorubrum saccharovorum JCM 8865T. The major respiratory quinones of strain WN019T were MK-8 and MK-8 (H2), and the major polar lipids were glycolipid (GL), phospholipid (PL), phosphatidylglycerol-sulphate (PGS), phosphatidylglycerol (PG) and phosphatidylglycerol-phosphate-methyl ester (Me-PGP). The DNA G + C content of the strain was 67.4 mol%. The average nucleotide identity (ANI) and digital DNA–DNA hybridization (dDDH) value based on whole genome sequences of strain WN019T and Halorubrum saccharovorum JCM 8865T were 87.5% and 35.4%, respectively. Phenotypic, chemotaxonomic, phylogenetic, and genomic analyses suggested that strain WN019T represents a novel species of the genus Halorubrum, for which the name Halorubrum salipaludis sp. nov. is proposed. The type strain is WN019T (= KCTC 4269T = ACCC 19977T).


Similar content being viewed by others
References
Chaumeil P, Mussig AJ, Hugenholtz P, Parks DH (2020) GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36(6):1925–1927. https://doi.org/10.1093/bioinformatics/btz848
Chen S, He J, Zhang J, Xu Y, Huang J, Ke L (2017a) Halorubrum salsamenti sp. nov., a novel halophilic archaeon isolated from a brine of salt mine. Curr Microbiol 74(11):1358–1364. https://doi.org/10.1007/s00284-017-1325-8
Chen S, Xu Y, Ke L (2017b) Halorubrum trueperi sp. nov., a halophilic archaeon isolated from a salt mine. Int J Syst Evol Micr 67(5):1564–1570. https://doi.org/10.1099/ijsem.0.001762
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu XW, De Meyer S, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68(1):461–466. https://doi.org/10.1099/ijsem.0.002516
Corral P, de la Haba RR, Sanchez-Porro C, Amoozegar MA, Papke RT, Ventosa A (2015) Halorubrum persicum sp. nov., an extremely halophilic archaeon isolated from sediment of a hypersaline lake. Int J Syst Evol Micr 65(6):1770–1778. https://doi.org/10.1099/ijs.0.000175
de la Haba RR, Corral P, Sanchez-Porro C, Infante-Dominguez C, Makkay AM, Amoozegar MA, Ventosa A, Papke RT (2018) Genotypic and lipid analyses of strains from the archaeal genus Halorubrum reveal insights into their taxonomy, divergence, and population structure. Front Microbiol. https://doi.org/10.3389/fmicb.2018.00512
Delcher AL, Bratke KA, Powers EC, Salzberg SL (2007) Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23(6):673–679. https://doi.org/10.1093/bioinformatics/btm009
Dyall-Smith M (2009) The Halohandbook: Protocols for haloarchaeal genetics, version 7.2, p. 118. http://www.haloarchaea.com/resources/halohandbook/
Feng J, Zhou PJ, Liu SJ (2004) Halorubrum xinjiangense sp. nov., a novel halophile isolated from saline lakes in China. Int J Syst Evol Micr 54(5):1789–1791. https://doi.org/10.1099/ijs.0.63209-0
Fraser SL, Jorgensen JH (1997) Reappraisal of the antimicrobial susceptibilities of Chryseobacterium and Flavobacterium species and methods for reliable susceptibility testing. Antimicrob Agents Chemother 41(12):2738–2741. https://doi.org/10.1128/AAC.41.12.2738
Gutiérrez MC, Castillo AM, Pagaling E et al (2008) Halorubrum kocurii sp. nov., an archaeon isolated from a saline lake. Int J Syst Evol Micr 58(9):2031–2035. https://doi.org/10.1099/ijs.0.65840-0
Klatt CG, Inskeep WP, Herrgard MJ, Jay ZJ, Rusch DB, Tringe SG, Parenteau MN, Ward DM, Boomer SM, Bryant DA et al (2013) Community structure and function of high-temperature chlorophototrophic microbial mats inhabiting diverse geothermal environments. Front Microbiol. https://doi.org/10.3389/fmicb.2013.00106
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Lagesen K, Hallin P, Rodland EA, Staerfeldt H, Rognes T, Ussery DW (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35(9):3100–3108. https://doi.org/10.1093/nar/gkm160
Li R, Li Y, Kristiansen K, Wang J (2008) SOAP: short oligonucleotide alignment program. Bioinformatics 24(5):713–714. https://doi.org/10.1093/bioinformatics/btn025
Li R, Zhu H, Ruan J et al (2010) De novo assembly of human genomes with massively parallel short read sequencing. Genome Res 20(2):265–272. https://doi.org/10.1101/gr.097261.109
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25(5):955–964. https://doi.org/10.1093/nar/25.5.955
McGenity TJ, Grant WD (1995) Transfer of Halobacterium saccharovorum, Halobacterium sodomense, Halobacterium trapanicum NRC-34021 and Halobacterium lacusprofundi to the Genus Halorubrum gen. nov., as Halorubrum saccharovorum comb. nov., Halorubrum sodomense comb. nov., Halorubrum trapanicum comb. nov., and Halorubrum lacusprofundi comb. nov. Syst Appl Microbiol 18(2):237–243. https://doi.org/10.1016/S0723-2020(11)80394-2
Meier-Kolthoff JP, Auch AF, Klenk H, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. https://doi.org/10.1186/1471-2105-14-60
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37(5):1530–1534. https://doi.org/10.1093/molbev/msaa015
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:223–241
Ochsenreiter T, Pfeifer F, Schleper C (2002) Diversity of Archaea in hypersaline environments characterized by molecular-phylogenetic and cultivation studies. Extremophiles 6(4):267–274. https://doi.org/10.1007/s00792-001-0253-4
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25(7):1043–1055. https://doi.org/10.1101/gr.186072.114
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microb 49(1):1–7
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106(45):19126–19131. https://doi.org/10.1073/pnas.0906412106
Richter M, Rosselló-Móra R, Gloeckner FO, Peplies J (2016) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32(6):929–931. https://doi.org/10.1093/bioinformatics/btv681
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 27(6):863–864. https://doi.org/10.1093/bioinformatics/btr026
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882. https://doi.org/10.1093/nar/25.24.4876
Yim KJ, Cha I, Lee H et al (2014) Halorubrum halophilum sp nov, an extremely halophilic archaeon isolated from a salt-fermented seafood. Anton Leeuw Int J G 105(3):603–612. https://doi.org/10.1007/s10482-014-0115-6
Acknowledgements
This work was supported by Science and Technology Partnership Program, Ministry of Science and Technology of China (KY202002003), National Natural Science Foundation of China (NSFC No. 31670113). We would like to thank Prof. Aharon Oren for very valuable help in naming the organism.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gong, Q., Zhao, P., Miao, S. et al. Halorubrum salipaludis sp. nov., isolated from the saline–alkaline soil. Arch Microbiol 204, 103 (2022). https://doi.org/10.1007/s00203-021-02729-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-021-02729-1