Abstract
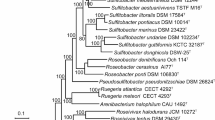
Filamentous iron oxides accumulating bacteria Sphaerotilus natans subsp. natans and S. natans subsp. sulfidivorans were described as subspecies based on 99.7% identity of their 16S rRNA sequences, in spite of important physiological difference. The ANI between their genomes was 94.7%, which indicate their assignment to different species. S. natans subsp. sulfidivorans and S. montanus possess genes for a complete SOX system, while S. natans subsp. natans encode only SoxYZ. There are genes for the Calvin cycle in the genomes of S. hippei DSM 566T, S. natans subsp. sulfidivorans D-501T, and S. montanus HST. Lithoautotrophy on reduced sulfur compounds is probably possible for S. natans subsp. sulfidivorans and S. montanus, but not for S. natans subsp. natans. Considering significant differences in the genome characteristics and metabolic potential of S. natans subsp. sulfidivorans and S. natans subsp. natans, we propose their classification as different species, S. natans and S. sulfidivorans sp. nov.

Similar content being viewed by others
References
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Belousova EV, Chernousova EYu, Dubinina GA, Tourova TP, Grabovich MYu (2013) Detection and analysis of sulfur metabolism genes in Sphaerotilus natans subsp. sulfidivorans representatives. Microbiology 82:586–593. https://doi.org/10.1134/S0026261713050020
Dondero NC (1961) Sphaerotilus, its nature and economic significance. Adv Appl Microbiol 3:77–107. https://doi.org/10.1016/S0065-2164(08)70507-0
Emerson D, Ghiorse WC (1993) Ultrastructure and chemical composition of the sheath of Leptothrix discophora SP-6. J Bacteriol 175:7808–7818. https://doi.org/10.1128/jb.175.24.7808-7818.1993
Fleming EJ, Woyke T, Donatello AR, Kuypers MMM, Sczyrba A, Littmann S, Emerson D (2018) Insights into the fundamental physiology of the uncultured Fe-oxidizing bacterium Leptothrix ochracea. Appl Environ Microb. https://doi.org/10.1128/AEM.02239-17
Fomenkov A, Grabovich M, Belousova E, Smolyakov D, Dubinina G, Roberts RJ (2019) Complete genome sequence and methylome analysis of Sphaerotilus natans subsp. sulfidivorans D-507. Microbiol Resour Announc. https://doi.org/10.1128/MRA.01194-19
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P (2007) DNA–DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Gridneva E, Chernousova E, Dubinina G, Akimov V, Kuever J, Detkova E, Grabovich M (2011) Sphaerotilus montanus sp. nov., Sphaerotilus hippei sp. nov., Sphaerotilus natans subsp. natans subsp. nov. and Sphaerotilus natans subsp. sulfidivorans subsp. nov., and an emended description of the genus Sphaerotilus. Int J Syst Evol Microbiol 61:916–925. https://doi.org/10.1099/ijs.0.023887-0
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O, Notes A (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Ishihara H, Hashimoto H, Taketa E, Suzuki T, Mandai K, Kunoh H, Takada J (2014) Silicon-rich, iron oxide microtubular sheath produced by an iron-oxidizing bacterium, Leptothrix sp. strain OUMS1, in culture. Minerals 4:565–577. https://doi.org/10.3390/min4030565
Kämpfer P (1998) Some Chemotaxonomic and Physiological Properties of the Genus Sphaerotilus. Syst Appl Microbiol 21(2):245–250
Konstantinidis KT, Rosselló-Móra R, Amann R (2017) Uncultivated microbes in need of their own taxonomy. ISME J 11:2399–2406. https://doi.org/10.1038/ismej.2017.113
Kützing FT (1833) Beitrag zur Kenntnis über die Entstehung und Metamorphose der niederen vegetalischen Organismen, nebst einer systematischen Zusammensetzung der hierher gehörigen niederen Algenformen. Linnaea 8:335–387
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinf. https://doi.org/10.1186/1471-2105-14-60
Mulder EG, van Veen WL (1963) Investigations on the Sphaerotilus-Leptothrix group. Antonie Van Leeuwenhoek 29:121–153. https://doi.org/10.1007/BF02046045
Nelson YM, Lion LW, Shuler ML, Ghiorse WC (1999) Lead binding to metal oxide and organic phases of natural aquatic biofilms. Limnol Oceanogr 44:1715–1729. https://doi.org/10.4319/lo.1999.44.7.1715
Olm MR, Crits-Christoph A, Diamond S, Lavy A, Matheus Carnevali PB, Banfield JF (2019) Consistent metagenome-derived metrics verify and delineate bacterial species boundaries. mSystems 5:e00731-e819. https://doi.org/10.1128/mSystems.00731-19
Rodriguez RLM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. Peer J Preprints 4:e1900v1. https://doi.org/10.7287/peerj.preprints.1900v1
Solisio C, Lodi A, Converti A, Borghi MD (2000) The effect of acid pre-treatment on the biosorption of chromium (III) by Sphaerotilus natans from industrial wastewater. Water Res 34:3171–3178. https://doi.org/10.1016/S0043-1354(00)00059-2
Spring S (2006) The genera Leptothrix and Sphaerotilus. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The Prokaryotes, 3rd edn. Springer Verlag, New York, pp 758–777. https://doi.org/10.1007/0-387-30745-1_35
Takeda M, Kondo K, Yamada M, Koizumi J, Mashima T, Matsugami A, Katahira M (2010) Solubilization and structural determination of a glycoconjugate which is assembled into the sheath of Leptothrix cholodnii. Int J Biol Macromol 46:206–211. https://doi.org/10.1016/j.ijbiomac.2009.12.006
Takeda M, Umezu T, Kawasaki Y, Shimura S, Kondo K, Koizumi J (2012) A spatial relationship between sheath elongation and cell proliferation in Sphaerotilus natans. Biosci Biotechnol Biochem 76:2357–2359
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Van Veen WL, Mulder EG, Deinema MH (1978) The Sphaerotilus-Leptothrix group of bacteria. Microbiol Rev 42:329–356
Yoon SH, Ha SM, Lim J, Kwon S, Chun J (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Funding
This work was partly supported by the Russian Science Foundation [project 20-14-00137 to MYG].
Author information
Authors and Affiliations
Contributions
Conceptualization, validation, and project administration: MYG and NVR; methodology, investigation, and writing—original draft preparation: DDS; methodology, investigation: NDM, AVB, and AVM; writing—review and editing: MVG, TSR, and NVR.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Grabovich, M.Y., Smolyakov, D.D., Beletsky, A.V. et al. Reclassification of Sphaerotilus natans subsp. sulfidivorans Gridneva et al. 2011 as Sphaerotilus sulfidivorans sp. nov. and comparative genome analysis of the genus Sphaerotilus. Arch Microbiol 203, 1595–1599 (2021). https://doi.org/10.1007/s00203-020-02158-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02158-6