Abstract
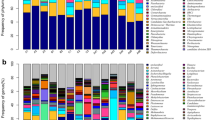
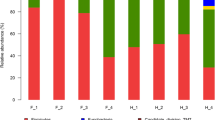
The black-necked crane (Grus nigricollis) is a vulnerable species, breeding exclusively on the high-altitude wetlands of the Qinghai–Tibet Plateau. Bird species harbor diverse communities of microorganisms within their gastrointestinal tracts, which have important roles in the health, nutrition, and physiology of birds. Hitherto, virtually nothing was known about the gut microbial communities associated with wild black-necked cranes. For the first time, this study characterized the gut microbial community compositions, diversity, and functions of black-necked cranes from six wintering areas in China using the Illumina Miseq platform. The taxonomic results revealed that Firmicutes, Proteobacteria, Actinobacteria, and Bacteroidetes were the four most abundant phyla in the gut of black-necked cranes. At the genus level, 11 genera including Lactobacillus, Pseudomonas, Carnobacterium, Pantoea, Enterococcus, Erwinia, Turicibacter, Bacillus, Phenylobacterium, Sanguibacter, and Psychrobacter were dominant. The differences in the gut microbial community alpha and the beta diversities of black-necked cranes among the six wintering areas were investigated. Furthermore, the representative microbial taxa and their predicted functions in each wintering location were also determined. These data represent the first analysis of the gut microbiome of black-necked cranes, providing a baseline for further microbiological studies and a foundation for the conservation of this bird.






Similar content being viewed by others
References
Astudillo-Garcia C, Bell JJ, Webster NS, Glasl B, Jompa J, Montoya JM, Taylor MW (2017) Evaluating the core microbiota in complex communities: a systematic investigation. Environ Microbiol 19:1450–1462
Barbosa A, Balague V, Valera F, Martínez A, Benzal J, Motas M, Diaz JI, Mira A, Pedrós-Alio C (2016) Age-related differences in the gastrointestinal microbiota of chinstrap penguins (Pygoscelis antarctica). PLoS ONE 11:e0153215
Barelli C, Albanese D, Donati C, Pindo M, Dallago C, Rovero F, Cavalieri D, Tuohy KM, Hauffe HC, De Filippo C (2015) Habitat fragmentation is associated with gut microbiota diversity of an endangered primate: implications for conservation. Sci Rep 5:14862
Barnes EM (1979) The intestinal microflora of poultry and game birds during life and after storage. J Appl Bacteriol 46:407–419
Bernardeau M, Guguen M, Vernoux JP (2006) Beneficial lactobacilli in food and feed: long-term use, biodiversity and proposals for specific and realistic safety assessments. FEMS Microbiol Rev 30:487–513
Bishop MA, Drolma T (2007) Tibet autonomous region January 2007 survey for black-necked crane, common crane, and bar-headed goose. China Crane News 11:24–25
Blanco G (2014) Influence of diet on the gastrointestinal flora of wintering red kites. Eur J Wildl Res 60:695–698
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Clarke SF, Murphy EF, Nilaweera K, Ross PR, Shanahan F, O’Toole PW, Cotter PD (2012) The gut microbiota and its relationship to diet and obesity: new insights. Gut Microbes 3:186–202
Dong HY, Lu GY, Zhong XY, Yang XJ (2016) Winter diet and food selection of the black-necked crane Grus nigricollis in Dashanbao, Yunnan, China. PeerJ 4:e1968
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Han XS, Guo YM, Mi CR, Huettmann F, Wen LJ (2017) Machine learning model analysis of breeding habitats for the black necked crane in central Asian uplands under anthropogenic pressures. Sci Rep 7:6114
Harris J, Mirande C (2013) A global overview of cranes: status, threats and conservation priorities. Chin Birds 4:189–209
Hubalek Z (2004) An annotated checklist of pathogenic microorganisms associated with migratory birds. J Wildl Dis 40:639–659
IUCN (2018) The IUCN red list of threatened species (version 2018-2). https://www.iucnredlist.org. Accessed 26 Jun 2019
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40:D109–D114
Knutie SA, Gotanda KM (2018) A non-invasive method to collect fecal samples from wild birds for microbiome studies. Microb Ecol 76:851–855
Kohl KD (2012) Diversity and function of the avian gut microbiota. J Comp Physiol B 182:591–602
Kong DJ, Luo WX, Liu Q, Li ZQ, Huan GY, Zhang JJ, Yang XJ (2018) Habitat use, preference, and utilization distribution of two crane species (Genus: Grus) in Huize National Nature Reserve, Yunnan–Guizhou Plateau, China. PeerJ 6:e5105
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821
Ley R, Turnbaugh P, Klein SI, Gordon J (2006) Microbial ecology: human gut microbes associated with obesity. Nature 444:1022–1023
Li FS (2014) IUCN black-necked crane (Grus nigricollis) conservation plan. Zool Res 35:3–9
Li L, Zhao X (2015) Comparative analyses of fecal microbiota in Tibetan and Chinese Han living at low or high altitude by barcoded 454 pyrosequencing. Sci Rep 5:14682
Li H, Qu J, Li T, Wirth S, Zhang Y, Zhao X, Li X (2018) Diet simplification selects for high gut microbial diversity and strong fermenting ability in high-altitude pikas. Appl Microbiol Biotechnol 102:6739–6751
Michel AJ, Ward LM, Goffredi SK, Dawson KS, Baldassarre DT, Brenner A, Gotanda KM, McCormack JE, Mullin SW, O’Neill A et al (2018) The gut of the finch: uniqueness of the gut microbiome of the Galápagos vampire finch. Microbiome 6:167
Morrison M, Pope PB, Denman SE, McSweeney CS (2009) Plant biomass degradation by gut microbiomes: more of the same or something new? Curr Opin Biotechnol 20:358–363
Price JT, Paladino FV, Lamont MM, Witherington BE, Bates ST, Soule T (2017) Characterization of the juvenile green turtle (Chelonia mydas) microbiome throughout an ontogenetic shift from pelagic to neritic habitats. PLoS ONE 12:e0177642
Qian FW, Wu HQ, Gao LB, Zhang HG, Li FS, Zhong XY, Zheng GM (2009) Migration routes and stopover sites of black-necked cranes determined by satellite tracking. J Field Ornithol 80:19–26
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glockner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596
Rimoldi S, Terova G, Ascione C, Giannico R, Brambilla F (2018) Next generation sequencing for gut microbiome characterization in rainbow trout (Oncorhynchus mykiss) fed animal by-product meals as an alternative to fishmeal protein sources. PLoS ONE 13:e0193652
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60
Sharpton TJ (2018) Role of the gut microbiome in vertebrate evolution. MSystems 3:e00174-17
Sommer F, Backhed F (2013) The gut microbiota-masters of host development and physiology. Nat Rev Microbiol 11:227–238
Song HT, Zhang YS, Gao HF, Guo YH, Li SN (2014) Plateau wetlands, an indispensable habitat for the black-necked crane (Grus nigricollis)—a review. Wetlands 34:629–639
Spergser J, Loncaric I, Tichy A, Fritz J, Scope A (2018) The cultivable autochthonous microbiota of the critically endangered Northern bald ibis (Geronticus eremita). PLoS ONE 13:e0195255
Trevelline BK, Fontaine SS, Hartup BK, Kohl KD (2019) Conservation biology needs a microbial renaissance: a call for the consideration of host-associated microbiota in wildlife management practices. Proc R Soc B 286:20182448
Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE, Sogin ML, Jones WJ, Roe BA, Affourtit JP et al (2009) A core gut microbiome in obese and lean twins. Nature 457:480–484
Tzeng TD, Pao YY, Chen PC, Weng FCH, Jean WD, Wang D (2015) Effects of host phylogeny and habitats on gut microbiomes of oriental river prawn (Macrobrachium nipponense). PLoS ONE 10:e0132860
Waite DW, Taylor MW (2015) Exploring the avian gut microbiota: current trends and future directions. Front Microbiol 6:673
Waite DW, Eason DK, Taylor MW (2014) Influence of hand rearing and bird age on the fecal microbiota of the critically endangered kakapo. Appl Environ Microbiol 80:4650–4658
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang W, Zheng SS, Sharshov K, Cao J, Sun H, Yang F, Wang XL, Li LX (2016) Distinctive gut microbial community structure in both the wild and farmed swan goose (Anser cygnoides). J Basic Microbiol 56:1299–1307
Wang W, Zheng SS, Sharshov K, Sun H, Yang F, Wang X, Li LX, Xiao ZX (2017) Metagenomic profiling of gut microbial communities in both wild and artificially reared Bar-headed goose (Anser indicus). Microbiologyopen 6:e00429
Wu HQ, Zha K, Zhang M, Yang XJ (2009) Nest site selection by black-necked crane Grus nigricollis in the Ruoergai wetland, China. Bird Conserv Int 19:277–286
Wu YN, Yang YZ, Cao L, Yin HQ, Xu MY, Wang ZJ, Liu YY, Wang X, Deng Y (2018) Habitat environments impacted the gut microbiome of long-distance migratory swan geese but central species conserved. Sci Rep 8:13314
Xie Y, Xia P, Wang H, Yu H, Giesy JP, Zhang Y, Mora MA, Zhang X (2016) Effects of captivity and artificial breeding on microbiota in feces of the red-crowned crane (Grus japonensis). Sci Rep 6:33350
Zhang Z, Xu D, Wang L, Hao J, Wang J, Zhou X, Wang W, Qiu Q, Huang X, Zhou J et al (2016) Convergent evolution of rumen microbiomes in high-altitude mammals. Curr Biol 26:1873–1879
Zhao N, Wang SP, Li HY, Liu SL, Li M, Luo J, Su W, He HX (2018) Influence of novel highly pathogenic avian influenza A (H5N1) virus infection on migrating whooper swans fecal microbiota. Front Cell Infect Microbiol 8:46
Zhou J, Shen J, Zhang R, Tang X, Li J, Xu B, Ding J, Gao Y, Xu D, Huang Z (2015) Molecular and biochemical characterization of a novel multidomain xylanase from Arthrobacter sp. GN16 isolated from the feces of Grus nigricollis. Appl Biochem Biotechnol 175:573–588
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant no. 31960277), the Natural Science Foundation of Qinghai Province of China (Grant no. 2018-ZJ-932Q), the Project of Qinghai Science and Technology Department (Grant no. 2016-ZJ-Y01), the Open Project of State Key Laboratory of Plateau Ecology and Agriculture, Qinghai University (Grant no. 2017-ZZ-21), and the Project of Tao He Yuan National Wetland Park in Qinghai Province (Grant no. 2018-THY-03). Dr. Wen Wang was supported by “1000 Talent” programs of Qinghai Province. The authors would like to express their sincere thanks to Doctor Ian Mather for his kind help in revising the English in this manuscript.
Author information
Authors and Affiliations
Contributions
WW and LXL designed experiments; WW, LXL, SYW, and YTS collected samples; FW, AZW, SYW, and YTS conducted experiments; WW, FW, and ZMLC conducted data analysis; WW, KS, and AD drafted the manuscripts; AZW and LXL revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest. All the authors read and approved the final manuscript.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, W., Wang, F., Li, L. et al. Characterization of the gut microbiome of black-necked cranes (Grus nigricollis) in six wintering areas in China. Arch Microbiol 202, 983–993 (2020). https://doi.org/10.1007/s00203-019-01802-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-019-01802-0