Abstract
Summary
A quantitative trait locus (QTL) for BMD maps to chromosome 1p36. We have analyzed a high density SNP panel from this region for linkage and association to BMD in 39 osteoporosis pedigrees. Our results support the presence of genes controlling BMD on 1p36 and indicate new candidates for further analyses.
Introduction
Low BMD is one of the major risk factors for osteoporosis. Following a genome scan in a sample of Caucasian families recruited through probands with low BMD, a region on 1p36 near marker D1S214 received support as a QTL for BMD from linkage (maximum lod-score = 2.87) and linkage disequilibrium (LD) analysis (p < 0.01).
Methods
To better characterize the genetic risk factors for low BMD located in this genomic region, we have genotyped the same group of families for 1095 SNPs located across 11 Mb on 1p36. Linkage and LD analyses have been performed using the variance component approach.
Results
Multivariate linkage analysis indicated two QTLs for femoral neck BMD, lumbar spine BMD and trochanter BMD simultaneously on 1p36, with maximum lod-scores of 4.37 at 12 cM and 3.59 at 22 cM. LD analysis identified several SNPs potentially associated with BMD, including the RERE gene SNP rs11121179 (p = 0.000005 for lumbar spine BMD). Other candidate genes include G1P2, SSU72 and CCDC27 (each containing 1 SNP with p < 0.001 for at least one BMD trait).
Conclusions
This study supports the presence in 1p36 of QTLs affecting BMD at multiple skeletal sites. Replication of our results in other independent cohorts is warranted.

Similar content being viewed by others
References
Ralston SH (2007) Genetics of osteoporosis. Proc Nutr Soc 66:158–165
Devoto M, Shimoya K, Caminis J et al (1998) A first-stage autosomal genome screen in extended pedigrees suggests genes predisposing to low bone mineral density are on chromosome 1p, 2p, and 4q. Eur J Hum Genet 6:151–157
Devoto M, Specchia C, Li HH et al (2001) Variance component linkage analysis indicates a QTL for femoral neck bone mineral density on chromosome 1p36. Hum Mol Genet 10:2447–2452
Wilson SG, Reed PW, Bansal A et al (2003) Comparison of genome screens for two independent cohorts provides replication of suggestive linkage of bone mineral density to 3p21 and 1p36. Am J Hum Genet 72:144–155
Wynne F, Drummond FJ, Daly M et al (2003) Suggestive linkage of 2p22–25 and 11q12–13 with low bone mineral density at the lumbar spine in the Irish population. Calcif Tissue Int 72:651–658
Karasik D, Cupples LA, Hannan MT et al (2004) Genome screen for a combined bone phenotype using principal component analysis: the Framingham study. Bone 34:547–556
Streeten EA, McBride DJ, Pollin TI et al (2006) Quantitative trait loci for BMD identified by autosome-wide linkage scan to chromosomes 7q and 21q in men from the Amish Family Osteoporosis Study. J Bone Miner Res 21:1433–1442
Xiao P, Shen H, Guo YF et al (2006) Genomic regions identified for BMD in a large sample including epistatic interactions and gender-specific effects. J Bone Miner Res 21:1536–1544
Lee YH, Rho YH, Choi SJ et al (2006) Meta-analysis of genome-wide linkage studies for bone mineral density. J Hum Genet 51:480–486
Ioannidis JP, Ng MY, Sham PC et al (2007) Meta-analysis of genome-wide scans provides evidence for sex- and site-specific regulation of bone mass. J Bone Miner Res 22:173–183
Devoto M, Spotila LD, Stabley DL et al (2005) Univariate and bivariate variance component linkage analysis of a whole-genome scan for loci contributing to bone mineral density. Eur J Hum Genet 13:781–788
O’Connell JR, Weeks DE (1998) PedCheck: A program for identifying genotype incompatibilities in linkage analysis. Am J Hum Genet 63:259–266
Barrett JC, Fry B, Maller J et al (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Wigginton JE, Cutler DJ, Abecasis GR (2005) A note on exact tests of Hardy–Weinberg equilibrium. Am J Hum Genet 76:887–893
Almasy L, Blangero J (1998) Multipoint quantitative trait linkage analysis in general pedigrees. Am J Hum Genet 62:1198–1211
Abecasis GR, Cardon LR, Cookson WO (2000) A general test of association for quantitative traits in nuclear families. Am J Hum Genet 66:279–292
Havill LM, Dyer TD, Richardson DK et al (2005) The quantitative trait linkage disequilibrium test: a more powerful alternative to the quantitative transmission disequilibrium test for use in the absence of population stratification. BMC Genet 6:S91–95
Abecasis GR, Wigginton JE (2005) Handling marker-marker linkage disequilibrium: pedigree analysis with clustered markers. Am J Hum Genet 77:754–767
Abecasis GR, Cherny SS, Cookson WO et al (2002) Merlin-rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 30:97–101
John S, Shephard N, Liu G et al (2004) Whole-genome scan, in a complex disease, using 11,245 single-nucleotide polymorphisms: comparison with microsatellites. Am J Hum Genet 75:54–64
Huang Q, Shete S, Amos CI (2004) Ignoring linkage disequilibrium among tightly linked markers induces false-positive evidence of linkage for affected sib pair analysis. Am J Hum Genet 75:1106–1112
Boyles AL, Scott WK, Martin ER et al (2005) Linkage disequilibrium inflates type I error rates in multipoint linkage analysis when parental genotypes are missing. Hum Hered 59:220–227
Almasy L, Dyer TD, Blangero J (1997) Bivariate quantitative trait linkage analysis: pleiotropy versus co-incident linkages. Genet Epidemiol 14:953–958
Kajita M, Ezura Y, Iwasaki H et al (2003) Association of the -381T/C promoter variation of the brain natriuretic peptide gene with low bone-mineral density and rapid postmenopausal bone loss. J Hum Genet 48:77–81
Spotila LD, Rodriguez H, Koch M et al (2003) Association analysis of bone mineral density and single nucleotide polymorphisms in two candidate genes on chromosome 1p36. Calcif Tissue Int 73:140–146
Tasker PN, Macdonald H, Fraser WD et al (2006) Association of PLOD1 polymorphisms with bone mineral density in a population-based study of women from the UK. Osteoporos Int 17:1078–1085
Yamada Y, Ando F, Shimokata H (2007) Association of candidate gene polymorphisms with bone mineral density in community-dwelling Japanese women and men. Int J Mol Med 19:791–801
Spotila LD, Rodriguez H, Koch M et al (2000) Association of a polymorphism in the TNFR2 gene with low bone mineral density. J Bone Miner Res 15:1376–1383
Albagha OM, Tasker PN, McGuigan FE et al (2002) Linkage disequilibrium between polymorphisms in the human TNFRSF1B gene and their association with bone mass in perimenopausal women. Hum Mol Genet 11:2289–2295
Tasker PN, Albagha OM, Masson CB et al (2004) Association between TNFRSF1B polymorphisms and bone mineral density, bone loss and fracture. Osteoporos Int 15:903–908
Huang QY, Shen H, Deng HY et al (2006) CA repeat polymorphism of the TNFR2 gene is not associated with bone mineral density in two independent Caucasian populations. J Bone Miner Metab 24:132–137
Grundberg E, Carling T, Brändström H et al (2004) A deletion polymorphism in the RIZ gene, a female sex steroid hormone receptor coactivator, exhibits decreased response to estrogen in vitro and associates with low bone mineral density in young Swedish women. J Clin Endocrinol Metab 89:6173–6178
Stolk L, van Meurs JB, Arp PP et al (2008) The RIZ Pro704 insertion-deletion polymorphism, bone mineral density and fracture risk: The Rotterdam study. Bone 42:286–293
Acknowledgments
The authors acknowledge the assistance of Dr. Alan Tenenhouse with the pedigree collection. The authors would also like to thank the co-authors of reference 10 for granting permission to cite unpublished results from the collaborative meta-analysis. This work was supported in part by a research grant from the National Osteoporosis Foundation.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table 1S
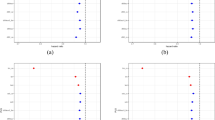
Results of QTLD analysis for selected SNPs (p < 0.01) corresponding to Figure 1 (DOC 133 KB)
Table 2S
Results of QTLD analysis for all SNPs in the RERE gene region (DOC 83.5 KB)
Fig. 1S
Pattern of LD in a region of 11 Mb (NCBI Build 35) on 1p36. The lines indicate mean pair-wise D’ (A) and r 2 (B) in a 500-kb sliding window (solid line: HapMap CEU population, dot line: our sample) (GIF 94.6 KB)
Fig. 2S
High resolution image file (TIF 233 KB)
Fig. 2S
LD test results in part of the RERE gene (DNA size 465.07 kb, mRNA size 8026 bp, 24 exons) and the genomic regions on the 3’ side of the gene on chromosome 1p36 between 8,325,000 and 8,475,000 base pairs (NCBI Build 35). (A) Negative log10 association P-values for FN, LS and TR BMD, respectively. (B) Pairwise r 2 plot from International HapMap CEU population data in the same region, where the intensity of the shading corresponds to different values of the disequilibrium coefficient r 2. The most significant marker, rs11121179, is contained within a large block of LD. Another less significant marker (rs301810, 0.001 < P < 0.01) is located within a smaller adjacent block (GIF 245 KB)
Rights and permissions
About this article
Cite this article
Zhang, H., Sol-Church, K., Rydbeck, H. et al. High resolution linkage and linkage disequilibrium analyses of chromosome 1p36 SNPs identify new positional candidate genes for low bone mineral density. Osteoporos Int 20, 341–346 (2009). https://doi.org/10.1007/s00198-008-0668-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00198-008-0668-1