Abstract
Introduction and hypothesis
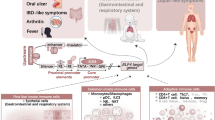
The development of recurrent urinary tract infections (rUTIs) is not completely understood. This review is aimed at investigating the connection between genetics and rUTIs and summarizing the results of studies that have documented variations in gene expression among individuals with rUTIs compared with healthy individuals.
Methods
A systematic search was conducted in Cochrane, Ovid, and PubMed, limiting the results to articles published between 1 January 2000, and 5 July 2022. Only studies comparing the difference in gene expression between individuals with rUTI and healthy individuals utilizing molecular techniques to measure gene expression in blood or urine samples were included in this systematic review. Gene network and pathways analyses were performed using Cytoscape software, with input data obtained from our systematic review of differentially expressed genes in rUTIs.
Results
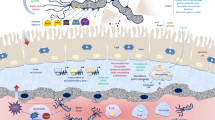
Six studies met our criteria for inclusion. The selected studies used molecular biology methods to quantify gene expression data from blood specimens. The analysis revealed that gene expressions of CXCR1 and TLR4 decreased, whereas CXCR2, TRIF, and SIGIRR increased in patients with rUTI compared with healthy controls. The analysis demonstrated that the most significant pathways were associated with TLR receptor signaling and tolerance, I-kappa B kinase/NF-kappa B signaling, and MyD88-independent TLR signaling.
Conclusions
This systematic review uncovered gene expression variations in several candidate genes and identified a number of underlying biological pathways associated with rUTIs. These findings could shift the treatment and prevention strategies for rUTIs.









Similar content being viewed by others
References
Flores-Mireles AL, Walker JN, Caparon M, Hultgren SJ. Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat Rev Microbiol. 2015;13(5):269–84.
Medina M, Castillo-Pino E. An introduction to the epidemiology and burden of urinary tract infections. Ther Adv Urol. 2019;11:1756287219832172.
Al-Badr A, Al-Shaikh G. Recurrent urinary tract infections management in women: a review. Sultan Qaboos Univ Med J. 2013;13(3):359–67.
Anger J, Lee U, Ackerman AL, et al. Recurrent uncomplicated urinary tract infections in women: AUA/CUA/SUFU guideline. J Urol. 2019;202(2):282–9.
Foxman B. Epidemiology of urinary tract infections: incidence, morbidity, and economic costs. Am J Med. 2002;113(Suppl 1A):5s–13s.
Hopkins WJ, Heisey DM, Lorentzen DF, Uehling DT. A comparative study of major histocompatibility complex and red blood cell antigen phenotypes as risk factors for recurrent urinary tract infections in women. J Infect Dis. 1998;177(5):1296–301.
Grönberg-Hernández J, Sundén F, Connolly J, Svanborg C, Wullt B. Genetic control of the variable innate immune response to asymptomatic bacteriuria. PLoS One. 2011;6(11):e28289.
Ching C, Schwartz L, Spencer JD, Becknell B. Innate immunity and urinary tract infection. Pediatr Nephrol. 2020;35(7):1183–92.
Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. BMJ. 2009;339:b2535.
Kellermeyer L, Harnke B, Knight S. Covidence and Rayyan. J Med Libr Assoc. 2018;106(4):580–3. https://doi.org/10.5195/jmla.2018.513.
Lo CK-L, Mertz D, Loeb M. Newcastle-Ottawa Scale: comparing reviewers’ to authors’ assessments. BMC Med Res Methodol. 2014;14(1):45.
Santos WMD, Secoli SR, Püschel VAA. The Joanna Briggs Institute approach for systematic reviews. Rev Lat Am Enfermagem. 2018;26:e3074.
Warde-Farley D, Donaldson SL, Comes O, et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010;38(Suppl_2):W214–20.
Isali I, Mahran A, Khalifa AO, et al. Gene expression in stress urinary incontinence: a systematic review. Int Urogynecol J. 2020;31(1):1–14.
Mousavian Z, Khodabandeh M, Sharifi-Zarchi A, Nadafian A, Mahmoudi A. StrongestPath: a Cytoscape application for protein-protein interaction analysis. BMC Bioinformatics. 2021;22(1):352.
Frendéus B, Godaly G, Hang L, Karpman D, Svanborg C. Interleukin-8 receptor deficiency confers susceptibility to acute pyelonephritis. J Infect Dis. 2001;183(Suppl 1):S56–60.
Frendéus B, Godaly G, Hang L, Karpman D, Lundstedt AC, Svanborg C. Interleukin 8 receptor deficiency confers susceptibility to acute experimental pyelonephritis and may have a human counterpart. J Exp Med. 2000;192(6):881–90.
Lundstedt AC, McCarthy S, Gustafsson MC, et al. A genetic basis of susceptibility to acute pyelonephritis. PLoS One. 2007;2(9):e825.
Lundstedt AC, Leijonhufvud I, Ragnarsdottir B, Karpman D, Andersson B, Svanborg C. Inherited susceptibility to acute pyelonephritis: a family study of urinary tract infection. J Infect Dis. 2007;195(8):1227–34.
Smithson A, Sarrias MR, Barcelo J, et al. Expression of interleukin-8 receptors (CXCR1 and CXCR2) in premenopausal women with recurrent urinary tract infections. Clin Diagn Lab Immunol. 2005;12(12):1358–63.
Ragnarsdóttir B, Samuelsson M, Gustafsson MC, Leijonhufvud I, Karpman D, Svanborg C. Reduced toll-like receptor 4 expression in children with asymptomatic bacteriuria. J Infect Dis. 2007;196(3):475–84.
Raghuwanshi SK, Su Y, Singh V, Haynes K, Richmond A, Richardson RM. The chemokine receptors CXCR1 and CXCR2 couple to distinct G protein-coupled receptor kinases to mediate and regulate leukocyte functions. J Immunol. 2012;189(6):2824–32.
Morris SW, Nelson N, Valentine MB, et al. Assignment of the genes encoding human interleukin-8 receptor types 1 and 2 and an interleukin-8 receptor pseudogene to chromosome 2q35. Genomics. 1992;14(3):685–91.
Bertini R, Barcelos LS, Beccari AR, et al. Receptor binding mode and pharmacological characterization of a potent and selective dual CXCR1/CXCR2 non-competitive allosteric inhibitor. Br J Pharmacol. 2012;165(2):436–54.
Godaly G, Hang L, Frendéus B, Svanborg C. Transepithelial neutrophil migration is CXCR1 dependent in vitro and is defective in IL-8 receptor knockout mice. J Immunol. 2000;165(9):5287–94.
Janssens S, Beyaert R. Role of Toll-like receptors in pathogen recognition. Clin Microbiol Rev. 2003;16(4):637–46.
Takeda K, Akira S. Toll-like receptors in innate immunity. Int Immunol. 2005;17(1):1–14.
Nahid MA, Satoh M, Chan EK. MicroRNA in TLR signaling and endotoxin tolerance. Cell Mol Immunol. 2011;8(5):388–403.
Butcher SK, O'Carroll CE, Wells CA, Carmody RJ. Toll-like like receptors drive specific patterns of tolerance and training on restimulation of macrophages. Front Immunol. 2018;9:933.
Ciesielska A, Matyjek M, Kwiatkowska K. TLR4 and CD14 trafficking and its influence on LPS-induced pro-inflammatory signaling. Cell Mol Life Sci. 2021;78(4):1233–61.
Yamamoto M, Sato S, Hemmi H, et al. Role of adaptor TRIF in the MyD88-independent toll-like receptor signaling pathway. Science. 2003;301(5633):640–3.
Nilsen NJ, Vladimer GI, Stenvik J, et al. A role for the adaptor proteins TRAM and TRIF in toll-like receptor 2 signaling. J Biol Chem. 2015;290(6):3209–22.
Riva F, Bonavita E, Barbati E, Muzio M, Mantovani A, Garlanda C. TIR8/SIGIRR is an interleukin-1 receptor/Toll like receptor family member with regulatory functions in inflammation and immunity. Front Immunol. 2012;3:322.
Li D, Zhang X, Chen B. SIGIRR participates in negative regulation of LPS response and tolerance in human bladder epithelial cells. BMC Immunol. 2015;16(1):73.
Yu W, Haque I, Venkatraman A, Menden HL, Mabry SM, Roy BC, et al. SIGIRR mutation in human necrotizing enterocolitis (NEC) disrupts STAT3-dependent microRNA expression in neonatal gut. Cell Mol Gastroenterol Hepatol. 2022;13(2):425–40.
Terlizzi ME, Gribaudo G, Maffei ME. UroPathogenic Escherichia coli (UPEC) infections: virulence factors, bladder responses, antibiotic, and non-antibiotic antimicrobial strategies. Front Microbiol. 2017;8:1566.
Bien J, Sokolova O, Bozko P. Role of uropathogenic Escherichia coli virulence factors in development of urinary tract infection and kidney damage. Int J Nephrol. 2012;2012:681473.
Yadav M, Zhang J, Fischer H, et al. Inhibition of TIR domain signaling by TcpC: MyD88-dependent and independent effects on Escherichia coli virulence. PLoS Pathog. 2010;6(9):e1001120.
Dobrindt U, Wullt B, Svanborg C. Asymptomatic bacteriuria as a model to study the coevolution of hosts and bacteria. Pathogens. 2016;5(1):21.
Spencer JD, Schwaderer AL, Becknell B, Watson J, Hains DS. The innate immune response during urinary tract infection and pyelonephritis. Pediatr Nephrol. 2014;29(7):1139–49.
Ambite I, Lutay N, Stork C, Dobrindt U, Wullt B, Svanborg C. Bacterial suppression of RNA polymerase II-dependent host gene expression. Pathogens. 2016;5(3):49
Zanoni I, Granucci F. Role of CD14 in host protection against infections and in metabolism regulation. Front Cell Infect Microbiol. 2013;3:32.
Schreiber HL t, Conover MS, et al. Bacterial virulence phenotypes of Escherichia coli and host susceptibility determine risk for urinary tract infections. Sci Transl Med. 2017;9(382):eaaf1283.
Zaffanello M, Malerba G, Cataldi L, et al. Genetic risk for recurrent urinary tract infections in humans: a systematic review. J Biomed Biotechnol. 2010;2010:321082.
Sorić Hosman I, Cvitković Roić A, Lamot L. A systematic review of the (un)known host immune response biomarkers for predicting recurrence of urinary tract infection. Front Med (Lausanne). 2022;9:931717.
Zaffanello M, Tardivo S, Cataldi L, Fanos V, Biban P, Malerba G. Genetic susceptibility to renal scar formation after urinary tract infection: a systematic review and meta-analysis of candidate gene polymorphisms. Pediatr Nephrol. 2011;26(7):1017–29.
Funding
None.
Author information
Authors and Affiliations
Contributions
I.I.: manuscript writing, project development, analysis; T.R.W.: manuscript writing, abstract/paper review/analysis; A.F.B.: manuscript writing, abstract/paper review; C.H.W.W.: analysis, manuscript writing; F.R.S.: manuscript writing, analysis; R.P.: manuscript writing; A.H.: manuscript writing; D.S.: project development, manuscript writing, project oversight.
Corresponding author
Ethics declarations
Conflicts of interest
D.S.: Renalis (research support), Caldera (consulting fee); A.H.: Collamedix (ownership stake); R.P.: Johnson & Johnson, MaternaMD (consulting fee), ; none of the other authors report any conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Isali, I., Wong, T.R., Batur, A.F. et al. Recurrent urinary tract infection genetic risk: a systematic review and gene network analysis. Int Urogynecol J 35, 259–271 (2024). https://doi.org/10.1007/s00192-023-05671-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00192-023-05671-6