Abstract
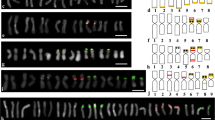
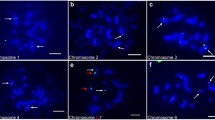
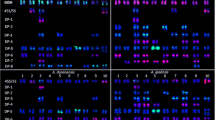
A molecular cytogenetic map of Chinese cabbage (Brassica rapa ssp. pekinensis, 2n=20) was constructed based on the 4′-6-diamino-2-phenylindole dihydrochloride-stained mitotic metaphase and pachytene chromosomes and multicolor fluorescence in situ hybridization (McFISH), using three repetitive DNA sequences, 5S rDNA, 45S rDNA, and C11-350H. The lengths of mitotic metaphase chromosomes ranged from 1.46 μm to 3.30 μm. Five 45S and three 5S rDNA loci identified were assigned to different chromosomes. The C11-350H loci were located on all the mitotic metaphase chromosomes, except chromosomes 2 and 4. The pachytene karyotype consisted of two metacentric (chromosomes 1 and 6), five submetacentric (chromosomes 3, 4, 5, 9 and 10), two subtelocentric (chromosomes 7 and 8), and one acrocentric (chromosome 2) chromosome(s). The mean lengths of ten pachytene chromosomes ranged from 23.7 μm to 51.3 μm, with a total of 385.3 μm, which is 17.5-fold longer than that of the mitotic metaphase chromosomes. In the proposed pachytene karyotype, all the chromosomes of B. rapa ssp. pekinensis can be identified on the basis of chromosome length, centromere position, heterochromatin pattern, and the location of the three repetitive sequences. Moreover, the precise locations of the earlier reported loci of 5S rDNA, 45S rDNA, and Chinese cabbage tandem DNA repeat C11-350H were established using McFISH analysis. We also identified a 5S rDNA locus on the long arm of pachytene bivalent 7, which could not be detected in the mitotic metaphase chromosomes in the present and earlier studies. The deduced karyotype will be useful for structural and functional genomic studies in B. rapa.



Similar content being viewed by others
References
Armstrong SJ, Fransz P, Marshall DF, Jones GH (1998) Physical mapping of DNA repetitive sequences to mitotic and meiotic chromosomes of Brassica oleracea var. alboglabra by fluorescence in situ hybridization. Heredity 81:666–667
Cheng BK, Heneen WK, Chen BY (1995) Mitotic karyotypes of Brassica campestris and Brassica alboglabra and identification of the B. alboglabra chromosome in an addition line. Genome 38:313–319
Cheng Z, Buell CR, Wing RA, Gu M, Jiang J (2001) Toward a cytological characterization of the rice genome. Genome Res 11:2133–2141
Cheng Z, Buell CR, Wing RA, Jiang J (2002) Resolution of fluorescence in situ hybridization mapping on rice mitotic prometaphase chromosomes, meiotic pachytene chromosomes and extended DNA fibers. Chromosome Res 10:379–387
de Jong JH, Fransz P, Zabel P (1999) High resolution FISH in plants—techniques and applications. Trends Plant Sci 4:258–263
Fransz P, Armstrong S, Alonso-Blanco C, Fischer TC, Torres-Ruiz RA, Jones G (1998) Cytogenetics for the model system Arabidopsis thaliana. Plant J 13:867–876
Fransz P, Armstrong S, de Jong JH, Parnell LD (2000) Integrated cytogenetic map of chromosome arm 4S of A. thaliana: structural organization of heterochromatic knob and centromere region. Cell 100:367–376
Fransz P, Soppe W, Schubert I (2003) Heterochromatin in interphase nuclei of Arabidopsis thaliana. Chromosome Res 11:227–240
Fukui K, Nakayama S, Ohmido N, Yoshiaki H, Yamabe M (1998) Quantitative karyotyping of three diploid Brassica species by imaging methods and localization of 45S rDNA loci on the identified chromosomes. Theor Appl Genet 96:325–330
Gerlach OL, Bedrock JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885
Harrison E, Heslop-Harrison JS (1995) Centromeric repetitive DNA sequences in the genus Brassica. Theor Appl Genet 90:157–165
Hasterok R, Jenkins G, Langdon T, Jones RN, Maluszynska J (2001) Ribosomal DNA is an effective marker of Brassica chromosomes. Theor Appl Genet 103:486–490
Iwabuchi M, Itoh K, Shimamoto K (1991) Molecular and cytological characterization of repetitive DNA sequences in Brassica. Theor Appl Genet 81:349–355
Iwano M, Sakamoto K, Suzuki G, Watanabe M, Takayama S, Fukui K, Hinata K, Isogai A (1998) Visualization of a self-incompatibility gene in Brassica campestris L. by multicolor FISH. Theor Appl Genet 96:751–757
Kim SY, Lim YP, Bang JW (1998) Cytogenetic analysis of Brassica campestris var. pekinensis using C-banding and FISH. Korean J Genet 20:285–294
Koo DH, Hur YK, Jin DC, Bang JW (2002) Karyotype analysis of a cucumber cultivar (Cucumis sativus L. cv. Winter Long) using C-banding and bicolor FISH. Mol Cell 13:413–418
Kulikova O, Gualtieri G, Kim DJ, Cook D, Huguet T, Bisseling T (2001) Integration of the FISH pachytene and genetic maps of Medicago truncatula. Plant J 27:49–58
Levan A, Frekga K, Sandberg A (1964) Nomenclature for centromeric position in chromosomes. Hereditas 52:201–220
Nishibayasahi S (1992) Banding in mitotic chromosomes of Brassica campestris var. pekinensis with a trypsin-Giemsa method. Genome 35:899–901
Olin-Fatih M (1994) A new method for differential staining of Brassica metaphase chromosomes and karyotypes of B. campestris, B. oleracea and B. napus. Hereditas 120:253–259
Olin-Fatih M, Heneen WK (1992) C-banded karyotypes of Brassica campestris, B. oleracea, and B. napus. Genome 35:583–589
Plader W, Hoshi Y, Malepszy S (1998) Sequential fluorescence staining with CMA and DAPI for somatic chromosome identification in cucumber (Cucumis sativus L.). J Appl Genet 39:249–258
Sadder MT, Weber G (2001) Karyotype of maize (Zea mays L.) mitotic metaphase chromosomes as revealed by fluorescence in situ hybridization (FISH) with cytogenetic DNA markers. Plant Mol Biol Rep 19:117–123
Snowdon RJ, Köhler W, Köhler A (1997) Chromosomal localization of rDNA loci in the Brassica A and C genomes. Genome 40:582–587
Snowdon RJ, Friedrich T, Friedt T, Köhler W (2002) Identifying the chromosomes of the A- and C-genome diploid Brassica species B. rapa (syn. campestris) and B. oleracea in their amphidiploid B. napus. Theor Appl Genet 104:533–538
Takamine N (1916) Über die ruhenden und die präsynaptischen Phasen der Reduktionsteilung. Bot Mag 30:293–303
U N (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Zhong XB, de Jong JH, Zabel P (1996) Preparation of tomato meiotic pachytene and mitotic metaphase chromosomes suitable for fluorescence in situ hybridization (FISH). Chromosome Res 4:24–28
Ziolkowski PA, Sadowski J (2002) FISH-mapping of rDNAs and Arabidopsis BACs on pachytene complements of selected Brassicas. Genome 45:189–197
Acknowledgements
This work was partially supported by grants from the Korean Science and Engineering Foundation (98-0402-0601-5) and the Rural Development Administration (BioGreen 21 Program), Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Friebe
Rights and permissions
About this article
Cite this article
Koo, DH., Plaha, P., Lim, Y.P. et al. A high-resolution karyotype of Brassica rapa ssp. pekinensis revealed by pachytene analysis and multicolor fluorescence in situ hybridization. Theor Appl Genet 109, 1346–1352 (2004). https://doi.org/10.1007/s00122-004-1771-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-004-1771-0