Abstract
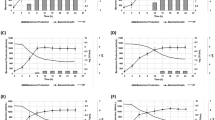
Elastase ofVibrio cholerae caused the lysis of freshly grown cells of Gram-negative (Pseudomonas aeruginosa, Proteus vulgaris, Salmonella paratyphi A andKlebsiella pneumoniae) bacteria. Gram-positive (Staphylococcus aureus andS. epidermidis) organisms were resistant to this enzyme. Heat killed and lyophilized Gram-positive and-negative bacteria (exceptS. aureus andS. epidermidis) showed higher sensitivity to elastase. Both Gram-negative and-positive bacteria were lyzed maximally by elastase at pH 8.0. At this pH, lytic activity of elastase was maximum in Tris-HCl and glycine-NaOH buffers followed by Tris-maleate and cacodylate buffers.
Similar content being viewed by others
References
Janoff A., Blondin J.: The effect of human granulocyte elastase on bacterial suspension.Lab. Invest.29, 454–457 (1973).
Janoff A., Blondin J., Sandhus R.A., Mosser A., Malemud C.: Human neutrophil elastase:In vitro effects on natural substrates suggest important physiological and pathological actions, pp. 603–620 inProtease and Biological Control, Vol. I (E. Reich, D.B. Rifkin, E. Shaw, Eds.), Cold Spring Harbor Laboratory, New York 1975.
Kaur M., Tripathi K.K., Gupta M., Jain P.K., Bansal M.R., Gupta K.G.: Production and partial characterization of elastase ofB. subtilis isolated from cervices of human females.Can. J. Microbiol.34, 855–859 (1988).
Morihara K., Tsuzuki H., Oka T., Inowe M., Ebata M.:Pseudomonas aeruginosa elastase: isolation, crystallisation and preliminary characterization.J. Biol. Chem.240, 3295–3304 (1965).
Sachar L.A., Winter K.K., Sicher N., Frankel S.: Photometric method for estimation of elastase activity.Proc. Soc. Exp. Biol. Med.90, 323–326 (1955).
Thomas R.C., Blout E.R.: Dependence of kinetic parameters for elastase catalysed amide hydrolysis in the length of peptide substrate.Biochemistry12, 57–65 (1973).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kaur, M., Gupta, M., Tripathi, K.K. et al. Lytic effect ofVibrio cholerae elastase on Gram-positive and-negative bacteria. Folia Microbiol 35, 183–189 (1990). https://doi.org/10.1007/BF02820483
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF02820483