Summary
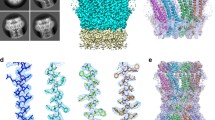
A procedure has been developed to isolate gap junction-enriched subcellular fractions from Drosophila. Crude membranes from larval homogenates were extracted with 1% N-lauroyl sarcosine in 6 M urea and the gap junctions were collected by centrifugation. The major proteins were separated by SDS PAGE and purified by electro-elution. Electron microscopy revealed structurally pleiomorphic gap junctions in the fractions which included (1) conventional, 16–18 nm-wide septalaminar, (2) collapsed, 13–15 nm-wide pentalaminar, (3) split, and (4) aggregated forms. The fractions contained five major proteins with apparent molecular weights of 18, 26, 36, 52 and 54 kD. Evidence based on (1) the degradation and aggregation behavior of the major proteins following electro-elution and reelectrophoresis, (2) immunological cross-reactivities by affinity-purified antibodies against the major proteins on immunoblots, and (3) immunofluorescent staining of presumptive gap junctions in Drosophila imaginal discs at the light-microscopic level and immunogold staining of purified gap junctions at the electron-microscopic level suggests that the major proteins are interrelated and of gap-junction origin.
Similar content being viewed by others
References
Bennett MVL, Spray DC (1985) Gap Junctions. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, pp 1–409
Berdan RC (1987) Intercellular communication in arthropods: biophysical, ultrastructural and biochemical approaches. In Cell-To-Cell Communication, WC DeMello, editor. Plenum Publishing Corporation, New York, pp 299–370
Berdan RC, Gilula NB (1988) The arthropod gap junction and pseudo-gap junction: Isolation and preliminary biochemical analysis. Cell Tissue Res 251:257–274
Bock G, Clark S (1987) Junctional complexes of epithelial cells. Ciba Foundation Symposium 125, John Wiley and Sons, New York, pp 78–167
Caveney S (1985) The role of gap junctions in development. Ann Rev Physiol 47:319–335
Dermietzel R, Leibstein A, Frixen U, Janssen-Timmen U, Traub O, Willecke K (1984) Gap junctions in several tissues share antigenic determinants with liver gap junctions. EMBO J 3:2261–2270
Epstein ML, Gilula NB (1977) A study of communication specificity between cells in culture. J Cell Biol 75:769–787
Evans WH (1988) Gap junctions: Towards a molecular structure. Bio Essays 8:3–6
Finbow ME, Buultjens TEJ, Lane NJ, Shuttleworth J, Pitts JD (1984) Isolation and characterization of arthropod gap junctions. EMBO J 3:2271–2278
Fraser SE, Green CR, Bode HR, Gilula NB (1987) Selective disruption of gap junctional communication interferes with a patterning process in Hydra. Science 237:49–55
Giloh H, Sedat JW (1982) Fluorescence microscopy: reduced photobleaching of rhodamine and fluorescein protein conjugates by n-propyl gallate. Science 217:1252–1255
Goodenough DA (1975) Methods for the isolation and structural characterization of hepatocyte gap junctions. In: Korn ED (ed) Methods in Membrane Biology. Vol. III. Plenum Publishing Corp., New York, pp 51–80
Green CR (1988) Evidence mounts for the role of gap junctions during development. Bio Essays 8:7–10
Green CR, Noirot-Timothee C, Noirot C (1983) Isolation and characterization of invertebrate smooth septate junctions. J Cell Sci 62:351–370
Henderson D, Eibl H, Weber K (1979) Structure and biochemistry of mouse hepatic gap junctions. J Mol Biol 132:193–218
Hertzberg E (1984) A detergent-independent procedure for the isolation of gap junctions from rat liver. J Biol Chem 259:9936–9943
Hertzberg EL, Gilula NB (1979) Isolation and characterization of gap junctions from rat liver. J Biol Chem 254:2138–2147
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lane NJ, Skaer HleB (1980) Intercellular junctions in insect tissues. Adv Insect Physiol 15:35–213
Lowenstein WR (1987) The cell-to-cell channel of gap junctions. Cell 48:725–726
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
MacDonald C (1985) Gap junction and cell-cell communication. Essays Biochem 21:86–118
Manjunath CK, Goings GE, Page E (1984) Detergent sensitivity and splitting of isolated liver gap junctions. J Membrane Biol 78:147–155
Manjunath CK, Nicholson BJ, Teplow D, Hood L, Page E, Revel J-P (1987) The cardiac gap junction protein (Mr 47,000) has a tissue-specific cytoplasmic domain of Mr 17,000 at its car-boxy-terminus. Biochem Biophys Res Comm 142:228–234
Nicholson B, Dermietzel R, Teplow D, Traub O, Willecke K, Revel J-P (1987) Two homologous protein components of hepatic gap junctions. Nature 329:732–734
Olmsted JB (1981) Affinity purification of antibodies from diazotized paper blots of heterogeneous protein samples. J Biol Chem 256:11955–11957
Ryerse JS (1982) Gap junctions are non-randomly distributed in Drosophila wing discs. Roux's Arch Dev Biol 191:335–339
Ryerse JS (1986) Isolation and characterization of gap junctions from Drosophila. J Cell Biol 103:74a
Spray DC (1985) Special topic: Gap junctions. Ann Rev Physiol 47:261–353
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76:4350–4354
Warner AE, Guthrie SC, Gilula NB (1984) Antibodies to gap-junctional protein selectively disrupt junctional communication in the early amphibian embryo. Nature 311:127–131
Yancey SB, Easter D, Revel J-P (1979) Cytological changes in gap junctions during liver regeneration. J Ultrastruct Res 67:229–242
Zassenhaus HP, Butow RA, Hannon YP (1982) Rapid electroelution of nucleic acids from agarose and acrylamide gels. Anal Biochem 125:125–130
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ryerse, J.S. Isolation and characterization of gap junctions from Drosophila melanogaster . Cell Tissue Res. 256, 7–16 (1989). https://doi.org/10.1007/BF00224713
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00224713