Abstract
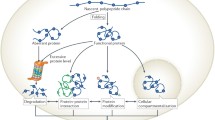
Biological systems often respond to environmental changes by rapidly altering the activity of specific enzymes: for example through desequesterization of enzyme activities by dissociation from inhibitors, activation/deactivation through posttranslational modification, or relocation of the enzyme to different organelles. This means that expression levels of enzymes do not necessarily correlate with the activities observed for these enzymes. In this chapter we review some of the approaches used to selectively image only the active sub-populations of given enzymes, the so-called activity-based protein profiling. A focus lies on recent developments that are taking this approach from chemical novelty to biochemical stalwart.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Willems LI, Overkleeft HS, van Kasteren SI (2014) Current developments in activity-based protein profiling. Bioconjug Chem 25(7):1181–1191
Cravatt BF, Wright AT, Kozarich JW (2008) Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu Rev Biochem 77(1):383–414, doi:10.1146/annurev.biochem.75.101304.124125
Ostrowski K, Barnard EA (1961) Application of isotopically labelled specific inhibitors as a method in enzyme cytochemistry. Exp Cell Res 25:465–468
Fenteany G, Standaert RF, Lane WS, Choi S, Corey EJ, Schreiber SL (1995) Inhibition of proteasome activities and subunit-specific amino-terminal threonine modification by lactacystin. Science 268(5211):726–731
Shi GP, Munger JS, Meara JP, Rich DH, Chapman HA (1992) Molecular cloning and expression of human alveolar macrophage cathepsin S, an elastinolytic cysteine protease. J Biol Chem 267(11):7258–7262
Bernstein KE, Welsh SL, Inman JK (1990) A deeply recessed active site in angiotensin-converting enzyme is indicated from the binding characteristics of biotin-spacer-inhibitor reagents. Biochem Biophys Res Commun 167(1):310–316, doi:http://dx.doi.org/10.1016/0006-291X(90)91766-L
Steven FS, Griffin MM, Williams LA, Clarke NW, Maier H (1991) Labelling of tumour cells with a biotinylated inhibitor of a cell surface protease. J Enzyme Inhib 4(4):337–346
Sin N, Meng L, Wang MQ, Wen JJ, Bornmann WG, Crews CM (1997) The anti-angiogenic agent fumagillin covalently binds and inhibits the methionine aminopeptidase, MetAP-2. Proc Natl Acad Sci U S A 94(12):6099–6103
Bogyo M, McMaster JS, Gaczynska M, Tortorella D, Goldberg AL, Ploegh H (1997) Covalent modification of the active site threonine of proteasomal beta subunits and the Escherichia coli homolog HslV by a new class of inhibitors. Proc Natl Acad Sci U S A 94(13):6629–6634
Borodovsky A, Kessler BM, Casagrande R, Overkleeft HS, Wilkinson KD, Ploegh HL (2001) A novel active site-directed probe specific for deubiquitylating enzymes reveals proteasome association of USP14. EMBO J 20(18):5187–5196
Ekkebus R, van Kasteren SI, Kulathu Y, Scholten A, Berlin I, Geurink PP, de Jong A, Goerdayal S, Neefjes J, Heck AJR, Komander D, Ovaa H (2013) On terminal alkynes that can react with active-site cysteine nucleophiles in proteases. J Am Chem Soc 135(8):2867–2870. doi:10.1021/ja309802n
Sommer S, Weikart ND, Linne U, Mootz HD (2013) Covalent inhibition of SUMO and ubiquitin-specific cysteine proteases by an in situ thiol-alkyne addition. Bioorg Med Chem 21(9):2511–2517. doi:10.1016/j.bmc.2013.02.039
McGouran JF, Gaertner SR, Altun M, Kramer HB, Kessler BM (2013) Deubiquitinating enzyme specificity for ubiquitin chain topology profiled by di-ubiquitin activity probes. Chem Biol 20(12):1447–1455. doi:10.1016/j.chembiol.2013.10.012
Mulder MP, El Oualid F, Ter Beek J, Ovaa H (2014) A native chemical ligation handle that enables the synthesis of advanced activity-based probes: diubiquitin as a case study. ChemBioChem. doi:10.1002/cbic.201402012
Li G, Liang Q, Gong P, Tencer AH, Zhuang Z (2014) Activity-based diubiquitin probes for elucidating the linkage specificity of deubiquitinating enzymes. Chem Commun 50(2):216–218. doi:10.1039/c3cc47382a
Liu Y, Patricelli MP, Cravatt BF (1999) Activity-based protein profiling: the serine hydrolases. Proc Natl Acad Sci U S A 96(26):14694–14699
Greenbaum D, Medzihradszky KF, Burlingame A, Bogyo M (2000) Epoxide electrophiles as activity-dependent cysteine protease profiling and discovery tools. Chem Biol 7(8):569–581
Blum G, Mullins SR, Keren K, Fonovic M, Jedeszko C, Rice MJ, Sloane BF, Bogyo M (2005) Dynamic imaging of protease activity with fluorescently quenched activity-based probes. Nat Chem Biol 1(4):203–209. doi:10.1038/nchembio728
Blum G, von Degenfeld G, Merchant MJ, Blau HM, Bogyo M (2007) Noninvasive optical imaging of cysteine protease activity using fluorescently quenched activity-based probes. Nat Chem Biol 3(10):668–677. doi:10.1038/nchembio.2007.26
Edgington LE, Verdoes M, Ortega A, Withana NP, Lee J, Syed S, Bachmann MH, Blum G, Bogyo M (2013) Functional imaging of legumain in cancer using a New quenched activity-based probe. J Am Chem Soc 135(1):174–182. doi:10.1021/ja307083b
Weerapana E, Wang C, Simon GM, Richter F, Khare S, Dillon MBD, Bachovchin DA, Mowen K, Baker D, Cravatt BF (2010) Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468(7325):790–U779. doi:10.1038/nature09472
Borodovsky A, Ovaa H, Kolli N, Gan-Erdene T, Wilkinson KD, Ploegh HL, Kessler BM (2002) Chemistry-based functional proteomics reveals novel members of the deubiquitinating enzyme. Chem Biol 9(10):1149–1159. doi:10.1016/s1074-5521(02)00248-x
Niphakis MJ, Cravatt BF (2014) Enzyme inhibitor discovery by activity-based protein profiling. Annu Rev Biochem 83(1):341–377, doi:10.1146/annurev-biochem-060713-035708
Willems LI, Jiang J, Li KY, Witte MD, Kallemeijn WW, Beenakker TJ, Schroder SP, Aerts JM, van der Marel GA, Codee JD, Overkleeft HS (2014) From covalent glycosidase inhibitors to activity-based glycosidase probes. Chemistry 20(35):10864–10872. doi:10.1002/chem.201404014
Speers AE, Adam GC, Cravatt BF (2003) Activity-based protein profiling in vivo using a copper(I)-catalyzed azide-alkyne [3 + 2] cycloaddition. J Am Chem Soc 125(16):4686–4687
Ovaa H, van Swieten PF, Kessler BM, Leeuwenburgh MA, Fiebiger E, van den Nieuwendijk AM, Galardy PJ, van der Marel GA, Ploegh HL, Overkleeft HS (2003) Chemistry in living cells: detection of active proteasomes by a two-step labeling strategy. Angew Chem Int Ed Engl 42(31):3626–3629. doi:10.1002/anie.200351314
Arastu-Kapur S, Ponder EL, Fonovic UP, Yeoh S, Yuan F, Fonovic M, Grainger M, Phillips CI, Powers JC, Bogyo M (2008) Identification of proteases that regulate erythrocyte rupture by the malaria parasite Plasmodium falciparum. Nat Chem Biol 4(3):203–213. doi:10.1038/nchembio.70
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
van Kasteren, S.I., Florea, B.I., Overkleeft, H.S. (2017). Activity-Based Protein Profiling: From Chemical Novelty to Biomedical Stalwart. In: Overkleeft, H., Florea, B. (eds) Activity-Based Proteomics. Methods in Molecular Biology, vol 1491. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6439-0_1
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6439-0_1
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6437-6
Online ISBN: 978-1-4939-6439-0
eBook Packages: Springer Protocols