Abstract
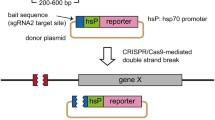
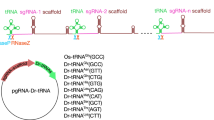
The controlled expression of Cas9 and/or sgRNA in transgenic zebrafish made it possible to knock out a gene in a spatially and/or temporally controlled manner. This transgenic approach can be more useful if multiple sgRNAs are efficiently expressed since we can improve the biallelic frame-shift mutation rate and circumvent the functional redundancy of genes and genetic compensation. We developed the tRNA-based system to express multiple functional sgRNAs from a single transcript in zebrafish and found that it is applicable to the transgenic expression of multiple sgRNAs. In this chapter, we describe a procedure for the generation of plasmids containing multiple sgRNAs flanked by tRNAs and a method to induce multiple CRISPR/Cas9-mediated genome modifications in transgenic zebrafish.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Cong L, Ran F, Cox D, Lin S, Barretto R, Habib N, Hsu P, Wu X, Jiang W, Marraffini L et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–822
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157:1262–1278
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X, Xiong JW, Xi JJ (2013) Genome editing with RNA-guided Cas9 nuclease in Zebrafish embryos. Cell Res 23:465–472
Hwang WY, Fu Y, Reyon D, Maeder ML, Tsai SQ, Sander JD, Peterson RT, Yeh JRJ, Joung JK (2013) Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 31:227–229
Ablain J, Durand EM, Yang S, Zhou Y, Zon LI (2015) A CRISPR/Cas9 vector system for tissue-specific gene disruption in zebrafish. Dev Cell 32:756–764
Yin L, Maddison LA, Li M, Kara N, Lafave MC, Varshney GK, Burgess SM, Patton JG, Chen W (2015) Multiplex conditional mutagenesis using transgenic expression of Cas9 and sgRNAs. Genetics 200:431–441
Di Donato V, De Santis F, Auer TO, Testa N, Sánchez-Iranzo H, Mercader N, Concordet JP, Del BF (2016) 2C-Cas9: a versatile tool for clonal analysis of gene function. Genome Res 26:681–692
Howe K, Clark MD, Torroja CF, Torrance J, Berthelot C, Muffato M, Collins JJEJE, Humphray S, McLaren K, Matthews L et al (2013) The zebrafish reference genome sequence and its relationship to the human genome. Nature 496:498–503
Rossi A, Kontarakis Z, Gerri C, Nolte H, Hölper S, Krüger M, Stainier DYR (2015) Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524:230–233
Xie K, Minkenberg B, Yang Y (2015) Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc Natl Acad Sci U S A 112:3570–3575
Shiraki T, Kawakami K (2018) A tRNA-based multiplex sgRNA expression system in zebrafish and its application to generation of transgenic albino fish. Sci Rep 8:1–14
Ansai S, Kinoshita M (2014) Targeted mutagenesis using CRISPR/Cas system in medaka. Biol Open 3:362–371
Moreno-Mateos MA, Vejnar CE, Beaudoin JD, Fernandez JP, Mis EK, Khokha MK, Giraldez AJ (2015) CRISPRscan: designing highly efficient sgRNAs for CRISPR-Cas9 targeting in vivo. Nat Methods 12(10):982–988
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, Park J, Blackburn EH, Weissman JS, Qi LS et al (2013) Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155:1479–1491
Hwang WY, Fu Y, Reyon D, Maeder ML, Kaini P, Sander JD, Joung JK, Peterson RT, Yeh JRJ (2013) Heritable and precise zebrafish genome editing using a CRISPR-Cas system. PLoS One 8:1–9
Gagnon JA, Valen E, Thyme SB, Huang P, Ahkmetova L, Pauli A, Montague TG, Zimmerman S, Richter C, Schier AF (2014) Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS One 9:5–12
Knapp DJHF, Michaels YS, Jamilly M, Ferry QRV, Barbosa H, Milne TA, Fulga TA (2019) Decoupling tRNA promoter and processing activities enables specific Pol-II Cas9 guide RNA expression. Nat Commun 10:1490
Lee RTH, Ng ASM, Ingham PW (2016) Ribozyme mediated gRNA Generation for in vitro and in vivo CRISPR/Cas9 mutagenesis. PLoS One 11:1–12
Acknowledgments
This work was supported by grants from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) in Japan: KAKENHI Grant Number 19K16196 and 21H02463.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Shiraki, T., Kawakami, K. (2024). Generation of Transgenic Fish Harboring CRISPR/Cas9-Mediated Somatic Mutations Via a tRNA-Based Multiplex sgRNA Expression. In: Amatruda, J.F., Houart, C., Kawakami, K., Poss, K.D. (eds) Zebrafish. Methods in Molecular Biology, vol 2707. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3401-1_20
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3401-1_20
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3400-4
Online ISBN: 978-1-0716-3401-1
eBook Packages: Springer Protocols