Abstract
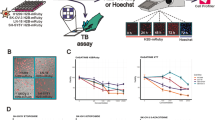
In recent years, the assembly and annotation of chemogenomic libraries have gained interest by the phenotypic screening community. Apart from basic annotations of the compound potency and selectivity, these compound libraries benefit in particular from annotation regarding the effect of the inhibitors on cellular viability to distinguish between on-target effects of a compound and unspecific cytotoxicity. Here, we provide a protocol to determine viability as a first determinant in compound quality control, using the Incucyte live-cell imaging system. The compounds are classified according to their calculated growth rate to determine a cytotoxic, cytostatic, or healthy outcome. All compounds affecting the growth rate can be further evaluated regarding their specific effects on cell health in a high-content live-cell multiplex assay, described in Chapter 5.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Tjaden A, Chaikuad A, Kowarz E, Marschalek R, Knapp S, Schröder M, Müller S (2022) Image based annotation of chemogenomic libraries for phenotypic screening. Molecules 27:1439
Müller S et al (2022) Target 2035 – update on the quest for a probe for every protein. RSC Med Chem 13:13–21
Moffat JG, Vincent F, Lee JA, Eder J, Prunotto M (2017) Opportunities and challenges in phenotypic drug discovery: an industry perspective. Nat Rev Drug Discov 16:531–543
Wagner BK, Schreiber SL (2016) The power of sophisticated phenotypic screening and modern mechanism-of-action methods. Cell Chem Biol 23:3–9
Hafner J, Niepel M, Chung M, Sorger PK (2016) Growth rate inhibition metrics correct for confounders in measuring sensitivity to cancer drugs. Nat Methods 13:521–527
Hughes P, Marshall D, Reid Y, Parkes H, Gelber C (2018) The costs of using unauthenticated, over-passaged cell lines: how much more data do we need? BioTechniques 43. https://www.future-science.com/doi/10.2144/000112598
Kozikowski BA et al (2003) The effect of room-temperature storage on the stability of compounds in DMSO. J Biomol Screen 8:205–209
Sartorius (2022) Incucyte live-cell analysis systems user manual. https://www.sartorius.com/download/1087348/incucyte-live-cell-analysis-systems-user-manual-en-l-8000-04-1%2D%2Ddata.pdf. Accessed 17 June 2022
EMBL-EBI (2022) BioImage Archive https://www.ebi.ac.uk/bioimage-archive/. Accessed 4 July 2022
Al-Aamri H et al (2019) Time dependent response of daunorubicin on cytotoxicity, cell cycle and DNA repair in acute lymphoblastic leukaemia. BMC Cancer 19. https://doi.org/10.1186/s12885-019-5377-y
Bruno S et al (1992) Different effects of staurosporine, an inhibitor of protein kinases, on the cell cycle and chromatin structure of normal and leukemic lymphocytes. Cancer Res 51:470–473
Seina J et al (2001) Cytotoxicity of digitoxin and related cardiac glycosides in human tumor cells. Anti-Cancer Drugs 12:475–483
Sanchez-Martinez C, Gelbert LM, Lallena MJ, Dios A (2015) Cyclin dependent kinase (CDK) inhibitors as anticancer drugs. Bioorg Med Chem Lett 25. https://doi.org/10.1016/j.bmcl.2015.05.100
Picaud S et al (2016) Promiscuous targeting of bromodomains by bromosporine identifies BET proteins as master regulators of primary transcription response in leukemia. Sci Adv 2. https://doi.org/10.1126/sciadv.1600760
Wen N et al (2019) Bromodomain inhibitor JQ1 induces cell cycle arrest and apoptosis of glioma stem cells through VEGF/PI3K/AKT signalling pathway. Int J Oncol 55. https://doi.org/10.3892/ijo.2019.4863
Wang TH et al (2000) Paclitaxel-induced cell death. Am Cancer Soc J 88. https://doi.org/10.1002/1097-0142(20000601)88:11<2619::AID-CNCR26>3.0.CO;2-J
Francipane MG, Lagasse E (2013) Selective targeting of human colon cancer stem-like cells by the mTOR inhibitor Torin-1. Oncotarget 4. https://doi.org/10.18632/oncotarget.1310
Violante GA et al (2002) Evaluation of the cytotoxicity effect of dimethyl sulfoxide (DMSO) on CaCo2/TC/colon tumor cell cultures. Biol Pharmaceut Bull 25:1600–1603
Acknowledgement
The authors are grateful for support by the Structural Genomics Consortium (SGC), a registered charity (No: 1097737) that receives funds from Bayer AG, Boehringer Ingelheim, Bristol Myers Squibb, Genentech, Genome Canada through Ontario Genomics Institute, Janssen, Merck KGaA, Pfizer and Takeda and by the German Cancer Research Center DKTK and the Frankfurt Cancer Institute (FCI). This project has received funding from the Innovative Medicines Initiative 2 Joint Undertaking (JU) under grant agreement No 875510. The JU receives support from the European Union’s Horizon 2020 research and innovation program and EFPIA and Ontario Institute for Cancer Research, Royal Institution for the Advancement of Learning McGill University, Kungliga Tekniska Hoegskolan, Diamond Light Source Limited. Disclaimer: This communication reflects the views of the authors, and the JU is not liable for any use that may be made of the information contained herein. A.T. is supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – grant number 259130777 (SFB1177). We would also like to thank Dr. Alexandra Stolz and her team for assistance with experimental setup and image optimization.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Elson, L., Tjaden, A., Knapp, S., Müller, S. (2023). Characterization of Cellular Viability Using Label-Free Brightfield Live-Cell Imaging. In: Merk, D., Chaikuad, A. (eds) Chemogenomics. Methods in Molecular Biology, vol 2706. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3397-7_6
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3397-7_6
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3396-0
Online ISBN: 978-1-0716-3397-7
eBook Packages: Springer Protocols