Abstract
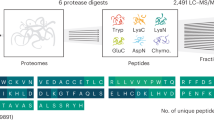
Molecular diversification of the cellular proteome through alternative splicing has emerged as an important biological principle. However, the lack of tools to specifically detect and quantify proteoforms (Smith et al., Nat Methods 10:186–187, 2013) is a major impediment to functional studies. Recently, biological mass spectrometry (MS) has undergone impressive advances (Mann, Nat Rev Mol Cell Biol 17:678, 2016), including the generation of a highly diverse set of biological applications (Aebersold and Mann, Nature 537:347–355, 2016), and has demonstrated to be an essential tool to address many biological questions (Savitski et al., Science 346:1255784, 2014; Rinner et al., Nat Methods 5:315–318, 2008). In particular, targeted LC-MS, with its high selectivity and specificity, is ideally suited for the precise and sensitive quantification of specific proteins and their proteoforms (Picotti and Aebersold, Nat Methods 9:555–566, 2012). We describe in detail the application of this workflow applied to dissect the molecular diversity of the synaptic adhesion proteins and their splicing-derived proteoforms (Schreiner et al., Elife 4:e07794, 2015).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, Mayr C, Kingsmore SF, Schroth GP, Burge CB (2008) Alternative isoform regulation in human tissue transcriptomes. Nature 456(7221):470–476. https://doi.org/10.1038/nature07509
Kim MS, Pinto SM, Getnet D, Nirujogi RS, Manda SS, Chaerkady R, Madugundu AK, Kelkar DS, Isserlin R, Jain S, Thomas JK, Muthusamy B, Leal-Rojas P, Kumar P, Sahasrabuddhe NA, Balakrishnan L, Advani J, George B, Renuse S, Selvan LD, Patil AH, Nanjappa V, Radhakrishnan A, Prasad S, Subbannayya T, Raju R, Kumar M, Sreenivasamurthy SK, Marimuthu A, Sathe GJ, Chavan S, Datta KK, Subbannayya Y, Sahu A, Yelamanchi SD, Jayaram S, Rajagopalan P, Sharma J, Murthy KR, Syed N, Goel R, Khan AA, Ahmad S, Dey G, Mudgal K, Chatterjee A, Huang TC, Zhong J, Wu X, Shaw PG, Freed D, Zahari MS, Mukherjee KK, Shankar S, Mahadevan A, Lam H, Mitchell CJ, Shankar SK, Satishchandra P, Schroeder JT, Sirdeshmukh R, Maitra A, Leach SD, Drake CG, Halushka MK, Prasad TS, Hruban RH, Kerr CL, Bader GD, Iacobuzio-Donahue CA, Gowda H, Pandey A (2014) A draft map of the human proteome. Nature 509(7502):575–581. https://doi.org/10.1038/nature13302
Smith LM, Kelleher NL, Consortium for Top Down P (2013) Proteoform: a single term describing protein complexity. Nat Methods 10(3):186–187. https://doi.org/10.1038/nmeth.2369
Baralle FE, Giudice J (2017) Alternative splicing as a regulator of development and tissue identity. Nat Rev Mol Cell Biol 18(7):437–451. https://doi.org/10.1038/nrm.2017.27
Scotti MM, Swanson MS (2016) RNA mis-splicing in disease. Nat Rev Genet 17(1):19–32. https://doi.org/10.1038/nrg.2015.3
Chen BE, Kondo M, Garnier A, Watson FL, Puettmann-Holgado R, Lamar DR, Schmucker D (2006) The molecular diversity of Dscam is functionally required for neuronal wiring specificity in Drosophila. Cell 125(3):607–620. https://doi.org/10.1016/j.cell.2006.03.034
Schreiner D, Nguyen TM, Russo G, Heber S, Patrignani A, Ahrne E, Scheiffele P (2014) Targeted combinatorial alternative splicing generates brain region-specific repertoires of neurexins. Neuron 84(2):386–398. https://doi.org/10.1016/j.neuron.2014.09.011
Mann M (2016) Origins of mass spectrometry-based proteomics. Nat Rev Mol Cell Biol 17(11):678. https://doi.org/10.1038/nrm.2016.135
Aebersold R, Mann M (2016) Mass-spectrometric exploration of proteome structure and function. Nature 537(7620):347–355. https://doi.org/10.1038/nature19949
Savitski MM, Reinhard FB, Franken H, Werner T, Savitski MF, Eberhard D, Martinez Molina D, Jafari R, Dovega RB, Klaeger S, Kuster B, Nordlund P, Bantscheff M, Drewes G (2014) Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346(6205):1255784. https://doi.org/10.1126/science.1255784
Rinner O, Seebacher J, Walzthoeni T, Mueller LN, Beck M, Schmidt A, Mueller M, Aebersold R (2008) Identification of cross-linked peptides from large sequence databases. Nat Methods 5(4):315–318. https://doi.org/10.1038/nmeth.1192
Picotti P, Aebersold R (2012) Selected reaction monitoring-based proteomics: workflows, potential, pitfalls and future directions. Nat Methods 9(6):555–566. https://doi.org/10.1038/nmeth.2015
Schreiner D, Simicevic J, Ahrne E, Schmidt A, Scheiffele P (2015) Quantitative isoform-profiling of highly diversified recognition molecules. elife 4:e07794. https://doi.org/10.7554/eLife.07794
Lange V, Picotti P, Domon B, Aebersold R (2008) Selected reaction monitoring for quantitative proteomics: a tutorial. Mol Syst Biol 4:222. https://doi.org/10.1038/msb.2008.61
Maiolica A, Junger MA, Ezkurdia I, Aebersold R (2012) Targeted proteome investigation via selected reaction monitoring mass spectrometry. J Proteome 75(12):3495–3513. https://doi.org/10.1016/j.jprot.2012.04.048
Gerber SA, Rush J, Stemman O, Kirschner MW, Gygi SP (2003) Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc Natl Acad Sci U S A 100(12):6940–6945. https://doi.org/10.1073/pnas.0832254100
Gygi SP, Rist B, Gerber SA, Turecek F, Gelb MH, Aebersold R (1999) Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat Biotechnol 17(10):994–999. https://doi.org/10.1038/13690
Ong SE, Blagoev B, Kratchmarova I, Kristensen DB, Steen H, Pandey A, Mann M (2002) Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteomics 1(5):376–386. https://doi.org/10.1074/mcp.m200025-mcp200
Picotti P, Bodenmiller B, Mueller LN, Domon B, Aebersold R (2009) Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell 138(4):795–806. https://doi.org/10.1016/j.cell.2009.05.051
Wilhelm M, Schlegl J, Hahne H, Gholami AM, Lieberenz M, Savitski MM, Ziegler E, Butzmann L, Gessulat S, Marx H, Mathieson T, Lemeer S, Schnatbaum K, Reimer U, Wenschuh H, Mollenhauer M, Slotta-Huspenina J, Boese JH, Bantscheff M, Gerstmair A, Faerber F, Kuster B (2014) Mass-spectrometry-based draft of the human proteome. Nature 509(7502):582–587. https://doi.org/10.1038/nature13319
Schaab C, Geiger T, Stoehr G, Cox J, Mann M (2012) Analysis of high accuracy, quantitative proteomics data in the MaxQB database. Mol Cell Proteomics 11(3):M111.014068. https://doi.org/10.1074/mcp.M111.014068
Perez-Riverol Y, Csordas A, Bai J, Bernal-Llinares M, Hewapathirana S, Kundu DJ, Inuganti A, Griss J, Mayer G, Eisenacher M, Perez E, Uszkoreit J, Pfeuffer J, Sachsenberg T, Yilmaz S, Tiwary S, Cox J, Audain E, Walzer M, Jarnuczak AF, Ternent T, Brazma A, Vizcaino JA (2019) The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res 47(D1):D442–D450. https://doi.org/10.1093/nar/gky1106
Kusebauch U, Campbell DS, Deutsch EW, Chu CS, Spicer DA, Brusniak MY, Slagel J, Sun Z, Stevens J, Grimes B, Shteynberg D, Hoopmann MR, Blattmann P, Ratushny AV, Rinner O, Picotti P, Carapito C, Huang CY, Kapousouz M, Lam H, Tran T, Demir E, Aitchison JD, Sander C, Hood L, Aebersold R, Moritz RL (2016) Human SRMAtlas: a resource of targeted assays to quantify the complete human proteome. Cell 166(3):766–778. https://doi.org/10.1016/j.cell.2016.06.041
Zauber H, Kirchner M, Selbach M (2018) Picky: a simple online PRM and SRM method designer for targeted proteomics. Nat Methods 15(3):156–157. https://doi.org/10.1038/nmeth.4607
Brusniak MY, Kwok ST, Christiansen M, Campbell D, Reiter L, Picotti P, Kusebauch U, Ramos H, Deutsch EW, Chen J, Moritz RL, Aebersold R (2011) ATAQS: a computational software tool for high throughput transition optimization and validation for selected reaction monitoring mass spectrometry. BMC Bioinformatics 12:78. https://doi.org/10.1186/1471-2105-12-78
Nahnsen S, Kohlbacher O (2012) In silico design of targeted SRM-based experiments. BMC Bioinformatics 13(Suppl 16):S8. https://doi.org/10.1186/1471-2105-13-S16-S8
Reiter L, Rinner O, Picotti P, Huttenhain R, Beck M, Brusniak MY, Hengartner MO, Aebersold R (2011) mProphet: automated data processing and statistical validation for large-scale SRM experiments. Nat Methods 8(5):430–435. https://doi.org/10.1038/nmeth.1584
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B, Kern R, Tabb DL, Liebler DC, MacCoss MJ (2010) Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26(7):966–968. https://doi.org/10.1093/bioinformatics/btq054
Gomez AM, Traunmuller L, Scheiffele P (2021) Neurexins: molecular codes for shaping neuronal synapses. Nat Rev Neurosci 22(3):137–151. https://doi.org/10.1038/s41583-020-00415-7
Shi T, Song E, Nie S, Rodland KD, Liu T, Qian WJ, Smith RD (2016) Advances in targeted proteomics and applications to biomedical research. Proteomics 16(15–16):2160–2182. https://doi.org/10.1002/pmic.201500449
Kusebauch U, Deutsch EW, Campbell DS, Sun Z, Farrah T, Moritz RL (2014) Using PeptideAtlas, SRMAtlas, and PASSEL: comprehensive resources for discovery and targeted proteomics. Curr Protoc Bioinformatics 46:13.25.11–13.25.28. https://doi.org/10.1002/0471250953.bi1325s46
Whiteaker JR, Halusa GN, Hoofnagle AN, Sharma V, MacLean B, Yan P, Wrobel JA, Kennedy J, Mani DR, Zimmerman LJ, Meyer MR, Mesri M, Rodriguez H, Clinical Proteomic Tumor Analysis C, Paulovich AG (2014) CPTAC assay portal: a repository of targeted proteomic assays. Nat Methods 11 (7):703–704. https://doi.org/10.1038/nmeth.3002
Sharma V, Eckels J, Schilling B, Ludwig C, Jaffe JD, MacCoss MJ, MacLean B (2018) Panorama public: a public repository for quantitative data sets processed in skyline. Mol Cell Proteomics 17(6):1239–1244. https://doi.org/10.1074/mcp.RA117.000543
Bhowmick P, Mohammed Y, Borchers CH (2018) MRMAssayDB: an integrated resource for validated targeted proteomics assays. Bioinformatics 34(20):3566–3571. https://doi.org/10.1093/bioinformatics/bty385
Bhowmick P, Roome S, Borchers CH, Goodlett DR, Mohammed Y (2021) An update on MRMAssayDB: a comprehensive resource for targeted proteomics assays in the community. J Proteome Res 20(4):2105–2115. https://doi.org/10.1021/acs.jproteome.0c00961
Schmidt T, Samaras P, Frejno M, Gessulat S, Barnert M, Kienegger H, Krcmar H, Schlegl J, Ehrlich HC, Aiche S, Kuster B, Wilhelm M (2018) ProteomicsDB. Nucleic Acids Res 46(D1):D1271–D1281. https://doi.org/10.1093/nar/gkx1029
Gessulat S, Schmidt T, Zolg DP, Samaras P, Schnatbaum K, Zerweck J, Knaute T, Rechenberger J, Delanghe B, Huhmer A, Reimer U, Ehrlich HC, Aiche S, Kuster B, Wilhelm M (2019) Prosit: proteome-wide prediction of peptide tandem mass spectra by deep learning. Nat Methods 16(6):509–518. https://doi.org/10.1038/s41592-019-0426-7
Desiere F, Deutsch EW, King NL, Nesvizhskii AI, Mallick P, Eng J, Chen S, Eddes J, Loevenich SN, Aebersold R (2006) The PeptideAtlas project. Nucleic Acids Res 34(Database issue):D655–D658. https://doi.org/10.1093/nar/gkj040
Cox J, Mann M (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat Biotechnol 26(12):1367–1372. https://doi.org/10.1038/nbt.1511
Addona TA, Abbatiello SE, Schilling B, Skates SJ, Mani DR, Bunk DM, Spiegelman CH, Zimmerman LJ, Ham AJ, Keshishian H, Hall SC, Allen S, Blackman RK, Borchers CH, Buck C, Cardasis HL, Cusack MP, Dodder NG, Gibson BW, Held JM, Hiltke T, Jackson A, Johansen EB, Kinsinger CR, Li J, Mesri M, Neubert TA, Niles RK, Pulsipher TC, Ransohoff D, Rodriguez H, Rudnick PA, Smith D, Tabb DL, Tegeler TJ, Variyath AM, Vega-Montoto LJ, Wahlander A, Waldemarson S, Wang M, Whiteaker JR, Zhao L, Anderson NL, Fisher SJ, Liebler DC, Paulovich AG, Regnier FE, Tempst P, Carr SA (2009) Multi-site assessment of the precision and reproducibility of multiple reaction monitoring-based measurements of proteins in plasma. Nat Biotechnol 27(7):633–641. https://doi.org/10.1038/nbt.1546
Peterson AC, Russell JD, Bailey DJ, Westphall MS, Coon JJ (2012) Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol Cell Proteomics 11(11):1475–1488. https://doi.org/10.1074/mcp.O112.020131
Gallien S, Duriez E, Crone C, Kellmann M, Moehring T, Domon B (2012) Targeted proteomic quantification on quadrupole-orbitrap mass spectrometer. Mol Cell Proteomics 11(12):1709–1723. https://doi.org/10.1074/mcp.O112.019802
Schnatbaum K, Zerweck J, Nehmer J, Wenschuh H, Schutkowski M, Reimer U (2011) SpikeTides™—proteotypic peptides for large-scale MS-based proteomics. Nat Methods 8(3):i–ii. https://doi.org/10.1038/nmeth.f.337
Glatter T, Ludwig C, Ahrne E, Aebersold R, Heck AJ, Schmidt A (2012) Large-scale quantitative assessment of different in-solution protein digestion protocols reveals superior cleavage efficiency of tandem Lys-C/trypsin proteolysis over trypsin digestion. J Proteome Res 11(11):5145–5156. https://doi.org/10.1021/pr300273g
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic Supplementary Material
Supplementary Table 1
Suppl_Table_1 (XLSX 22 kb)
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Schmidt, A., Schreiner, D. (2022). Quantitative Detection of Protein Splice Variants by Selected Reaction Monitoring (SRM) Mass Spectrometry. In: Scheiffele, P., Mauger, O. (eds) Alternative Splicing. Methods in Molecular Biology, vol 2537. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2521-7_14
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2521-7_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2520-0
Online ISBN: 978-1-0716-2521-7
eBook Packages: Springer Protocols