Abstract
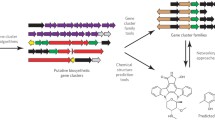
Genome mining has become an invaluable tool in natural products research to quickly identify and characterize the biosynthetic pathways that assemble secondary or specialized metabolites. Recently, evolutionary principles have been incorporated into genome mining strategies in an effort to better assess and prioritize novelty and understand their chemical diversification for engineering purposes. Here, we provide an introduction to the principles underlying evolutionary genome mining, including bioinformatic strategies and natural product biosynthetic databases. We introduce workflows for traditional genome mining, focusing on the popular pipeline antiSMASH, and methods to predict enzyme substrate specificity from genomic information. We then provide an in-depth discussion of evolutionary genome mining workflows, including EvoMining, CORASON, ARTS, and others, as adopted by our group for the discovery and prioritization of natural products biosynthetic gene clusters and their products.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bentley SD et al (2002) Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417:141–147
Chevrette MG, Currie CR (2019) Emerging evolutionary paradigms in antibiotic discovery. J Ind Microbiol Biotechnol 46:257–271
Chevrette MG et al (2020) Evolutionary dynamics of natural product biosynthesis in bacteria. Nat Prod Rep 37:566–599
Cruz-Morales P et al (2016) Phylogenomic analysis of natural products biosynthetic gene clusters allows discovery of Arseno-organic metabolites in model Streptomycetes. Genome Biol Evol 8:1906–1916
Chevrette MG et al (2019) The antimicrobial potential of Streptomyces from insect microbiomes. Nat Commun 10:516
Hurley A et al (2021) Tiny earth: a big idea for STEM education and antibiotic discovery. MBio 12:e03432-20
Montalbán-López M et al (2021) New developments in RiPP discovery, enzymology and engineering. Nat Prod Rep 38:130–239
Whitford CM, Cruz-Morales P, Keasling JD, Weber T (2021) The design-build-test-learn cycle for metabolic engineering of Streptomycetes. Essays Biochem 65(2):261–275. https://doi.org/10.1042/EBC20200132
Blin K et al (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Blin K et al (2017) antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res 45:W36–W41
Narzisi G, Mishra B (2011) Comparing De novo genome assembly: the long and short of it. PLoS One 6:e19175
Liao Y-C, Lin S-H, Lin H-H (2015) Completing bacterial genome assemblies: strategy and performance comparisons. Sci Rep 5:1–8
Davis JJ et al (2020) The PATRIC bioinformatics resource center: expanding data and analysis capabilities. Nucleic Acids Res 48:D606–D612
Aziz RK et al (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Delcher AL, Harmon D, Kasif S, White O, Salzberg SL (1999) Improved microbial gene identification with GLIMMER. Nucleic Acids Res 27:4636–4641
Hyatt D et al (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119
Devoid S et al (2013) Automated genome annotation and metabolic model reconstruction in the SEED and model SEED. Methods Mol Biol 985:17–45
Majoros WH, Pertea M, Salzberg SL (2004) TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20:2878–2879
van Santen JA, Kautsar SA, Medema MH, Linington RG (2021) Microbial natural product databases: moving forward in the multi-omics era. Nat Prod Rep 38:264–278
Sorokina M, Steinbeck C (2020) Review on natural products databases: where to find data in 2020. J Cheminform 12:20
Kautsar SA et al (2020) MIBiG 2.0: a repository for biosynthetic gene clusters of known function. Nucleic Acids Res 48:D454–D458
Blin K, Shaw S, Kautsar SA, Medema MH, Weber T (2021) The antiSMASH database version 3: increased taxonomic coverage and new query features for modular enzymes. Nucleic Acids Res 49:D639–D643
Medema MH et al (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:W339–W346
Wolf T, Shelest V, Nath N, Shelest E (2016) CASSIS and SMIPS: promoter-based prediction of secondary metabolite gene clusters in eukaryotic genomes. Bioinformatics 32:1138–1143
Kloosterman AM, Shelton KE, van Wezel GP, Medema MH, Mitchell DA (2020) RRE-Finder: a Genome-Mining Tool for Class-Independent RiPP Discovery. mSystems 5:e00267
Li W et al (2021) RefSeq: expanding the prokaryotic genome annotation pipeline reach with protein family model curation. Nucleic Acids Res 49:D1020–D1028
Kamra P, Gokhale RS, Mohanty D (2005) SEARCHGTr: a program for analysis of glycosyltransferases involved in glycosylation of secondary metabolites. Nucleic Acids Res 33:W220–W225
Caboche S, Leclère V, Pupin M, Kucherov G, Jacques P (2010) Diversity of monomers in nonribosomal peptides: towards the prediction of origin and biological activity. J Bacteriol 192:5143–5150
Stachelhaus T, Mootz HD, Marahiel MA (1999) The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem Biol 6:493–505
Minowa Y, Araki M, Kanehisa M (2007) Comprehensive analysis of distinctive polyketide and nonribosomal peptide structural motifs encoded in microbial genomes. J Mol Biol 368:1500–1517
Khayatt BI, Overmars L, Siezen RJ, Francke C (2013) Classification of the adenylation and acyl-transferase activity of NRPS and PKS systems using ensembles of substrate specific hidden Markov models. PLoS One 8:e62136
Röttig M et al (2011) NRPSpredictor2--a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res 39:W362–W367
Chevrette MG, Aicheler F, Kohlbacher O, Currie CR, Medema MH (2017) SANDPUMA: ensemble predictions of nonribosomal peptide chemistry reveal biosynthetic diversity across Actinobacteria. Bioinformatics 33:3202–3210
Helfrich EJN et al (2021) Evolution of combinatorial diversity in trans-acyltransferase polyketide synthase assembly lines across bacteria. Nat Commun 12:1422
chevrm. chevrm/transPACT: transPACT v1.0.1. (2020). https://doi.org/10.5281/zenodo.4148258
Conway KR, Boddy CN (2012) ClusterMine360: a database of microbial PKS/NRPS biosynthesis. Nucleic Acids Res 41:D402–D407
Ichikawa N et al (2013) DoBISCUIT: a database of secondary metabolite biosynthetic gene clusters. Nucleic Acids Res 41:D408–D414
Sélem-Mojica N, Aguilar C, Gutiérrez-García K, Martínez-Guerrero CE, Barona-Gómez F (2019) EvoMining reveals the origin and fate of natural product biosynthetic enzymes. Microb Genom 5:e000260
Chevrette MG et al (2019) Taxonomic and metabolic incongruence in the ancient genus. Front Microbiol 10:2170
Cruz-Morales P et al (2013) The genome sequence of Streptomyces lividans 66 reveals a novel tRNA-dependent peptide biosynthetic system within a metal-related genomic island. Genome Biol Evol 5:1165–1175
Ausland C et al (2021) dbCAN-PUL: a database of experimentally characterized CAZyme gene clusters and their substrates. Nucleic Acids Res 49:D523–D528
Alcock BP et al (2020) CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res 48:D517–D525
Palaniappan K et al (2019) IMG-ABC v.5.0: an update to the IMG/Atlas of Biosynthetic Gene Clusters Knowledgebase. Nucleic Acids Res 48:D422–D430
Bortolaia V et al (2020) ResFinder 4.0 for predictions of phenotypes from genotypes. J Antimicrob Chemother 75:3491–3500
van Santen JA et al (2019) The natural products atlas: an open access Knowledge Base for microbial natural products discovery. ACS Cent Sci 5:1824–1833
Medema MH, Takano E, Breitling R (2013) Detecting sequence homology at the gene cluster level with MultiGeneBlast. Mol Biol Evol 30:1218–1223
Navarro-Muñoz JC et al (2019) A computational framework to explore large-scale biosynthetic diversity. Nat Chem Biol 16:60–68
Kautsar SA, van der Hooft JJJ, de Ridder D, Medema MH (2021) BiG-SLiCE: A highly scalable tool maps the diversity of 1.2 million biosynthetic gene clusters. Gigascience 10:giaa154
Kautsar SA, Blin K, Shaw S, Weber T, Medema MH (2020) BiG-FAM: the biosynthetic gene cluster families database. Nucleic Acids Res 49:D490–D497
Alanjary M, Cano-Prieto C, Gross H, Medema MH (2019) Computer-aided re-engineering of nonribosomal peptide and polyketide biosynthetic assembly lines. Nat Prod Rep 36:1249–1261
Adamek M, Alanjary M, Ziemert N (2019) Applied evolution: phylogeny-based approaches in natural products research. Nat Prod Rep 36:1295–1312
Barona-Gómez F, Cruz-Morales P, Noda-García L (2012) What can genome-scale metabolic network reconstructions do for prokaryotic systematics? Antonie Van Leeuwenhoek 101:35–43
Medema MH, Fischbach MA (2015) Computational approaches to natural product discovery. Nat Chem Biol 11:639–648
Mungan MD et al (2020) ARTS 2.0: feature updates and expansion of the antibiotic resistant target seeker for comparative genome mining. Nucleic Acids Res 48:W546–W552
Alanjary M et al (2017) The antibiotic resistant target seeker (ARTS), an exploration engine for antibiotic cluster prioritization and novel drug target discovery. Nucleic Acids Res 45:W42–W48
Cimermancic P et al (2014) Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158:412–421
Choo KH, Tong JC, Zhang L (2004) Recent applications of hidden Markov models in computational biology. Genomics Proteomics Bioinformatics 2:84–96
Hannigan GD et al (2019) A deep learning genome-mining strategy for biosynthetic gene cluster prediction. Nucleic Acids Res 47:e110
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Chevrette, M.G. et al. (2022). Evolutionary Genome Mining for the Discovery and Engineering of Natural Product Biosynthesis. In: Skellam, E. (eds) Engineering Natural Product Biosynthesis. Methods in Molecular Biology, vol 2489. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2273-5_8
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2273-5_8
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2272-8
Online ISBN: 978-1-0716-2273-5
eBook Packages: Springer Protocols