Abstract
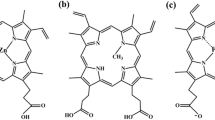
Mesoporphyrin IX (MPIX) contains a planar macrocycle center that can interact with various divalent metal ions through the exposed binding sites, leading to the metalation of MPIX. The DNA aptamers for porphyrin molecules usually display different catalytic functions (termed deoxyribozymes or DNAzymes), which can accelerate such chemical reactions. Inspired by this, an affinity chromatography selection approach was designed for identifying a porphyrin metalation DNAzyme. In our experiment, N-methyl mesoporphyrin IX (NMM), an analog of MPIX, is used as the target molecule, owing to its stable and high fluorescence enhancement after combining with specific oligonucleotides. Our results showed that the selected aptamer Nm1 is capable of binding to NMM with a low micromolar dissociation constant (0.75 ± 0.08 μM) and displays a catalytic activity for MPIX metalation with 3.3-fold rate enhancement. The protocol for isolation of such a porphyrin metalation DNAzyme is described in detail here.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346(6287):818–822
Tuerk C, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249(4968):505–510
Hermann T, Patel DJ (2000) Adaptive recognition by nucleic acid aptamers. Science 287(5454):820–825
Tan W, Wang H, Chen Y, Zhang X, Zhu H, Yang C, Yang R, Liu C (2011) Molecular aptamers for drug delivery. Trends Biotechnol 29(12):634–640
Yan S-R, Foroughi MM, Safaei M, Jahani S, Ebrahimpoor N, Borhani F, Baravati NRZ, Aramesh-Boroujeni Z, Foong LK (2020) A review: recent advances in ultrasensitive and highly specific recognition aptasensors with various detection strategies. Int J Biol Macromol 155:184–207
Willner I, Shlyahovsky B, Zayats M, Willner B (2008) DNAzymes for sensing, nanobiotechnology and logic gate applications. Chem Soc Rev 37(6):1153–1165
Teller C, Shimron S, Willner I (2009) Aptamer−DNAzyme hairpins for amplified biosensing. Anal Chem 81(21):9114–9119
Li W, Li Y, Liu Z, Lin B, Yi H, Xu F, Nie Z, Yao S (2016) Insight into G-quadruplex-hemin DNAzyme/RNAzyme: adjacent adenine as the intramolecular species for remarkable enhancement of enzymatic activity. Nucleic Acids Res 44(15):7373–7384
Norouzi A, Ravan H, Mohammadi A, Hosseinzadeh E, Norouzi M, Fozooni T (2018) Aptamer–integrated DNA nanoassembly: a simple and sensitive DNA framework to detect cancer cells. Anal Chim Acta 1017:26–33
Conn MM, Prudent JR, Schultz PG (1996) Porphyrin metalation catalyzed by a small RNA molecule. J Am Chem Soc 118(29):7012–7013
Ren R, Wang L-L, Ding T-R, Li X-M (2014) Enzyme-free amplified detection of nucleic acids based on self-sustained replication of RNAzyme and its application in tumor cell detection. Biosens Bioelectron 54:122–127
Li Y, Sen D (1996) A catalytic DNA for porphyrin metallation. Nat Struct Biol 3(9):743–747
Paniel N, Istamboulié G, Triki A, Lozano C, Barthelmebs L, Noguer T (2017) Selection of DNA aptamers against penicillin G using capture-SELEX for the development of an impedimetric sensor. Talanta 162:232–240
Duan N, Gong W, Wu S, Wang Z (2017) An ssDNA library immobilized SELEX technique for selection of an aptamer against ractopamine. Anal Chim Acta 961:100–105
Komarova N, Andrianova M, Glukhov S, Kuznetsov A (2018) Selection, characterization, and application of ssDNA aptamer against furaneol. Molecules 23(12):3159
Guo L, Nie D, Qiu C, Zheng Q, Wu H, Ye P, Hao Y, Fu F, Chen G (2012) A G-quadruplex based label-free fluorescent biosensor for lead ion. Biosens Bioelectron 35(1):123–127
Yett A, Lin LY, Beseiso D, Miao J, Yatsunyk LA (2019) N-methyl mesoporphyrin IX as a highly selective light-up probe for G-quadruplex DNA. J Porphyr Phthalocyanines 23:1195–1215
Zadeh JN, Steenberg CD, Bois JS, Wolfe BR, Pierce MB, Khan AR, Dirks RM, Pierce NA (2011) NUPACK: analysis and design of nucleic acid systems. J Comput Chem 32(1):170–173
Huang CY (1982) Determination of binding stoichiometry by the continuous variation method: the job plot. In: Purich DL (ed) Methods enzymol, vol 87. Academic Press, Cambridge, pp 509–525
Yang L, Ding P, Luo Y, Wang J, Lv H, Li W, Cao Y, Pei R (2019) Exploration of catalytic nucleic acids on porphyrin metalation and peroxidase activity by in vitro selection of aptamers for N-methyl mesoporphyrin IX. ACS Comb Sci 21(2):83–89
Yang J, Bowser MT (2013) Capillary electrophoresis–SELEX selection of catalytic DNA aptamers for a small-molecule porphyrin target. Anal Chem 85(3):1525–1530
Owczarzy R, Tataurov AV, Wu Y, Manthey JA, McQuisten KA, Almabrazi HG, Pedersen KF, Lin Y, Garretson J, McEntaggart NO (2008) IDT SciTools: a suite for analysis and design of nucleic acid oligomers. Nucleic Acids Res 36(2):163–169
Acknowledgments
This work was financially supported by the Natural Science Foundation of China (21575154, 21775160, 81801837, 31800685), the CAS/SAFEA International Innovation Teams program, and the Science Foundation of Jiangsu Province (BE2016680, BE2018665, BK20180250, BK20180258, BK20180261).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Yang, L., Cao, Y., Pei, R. (2022). Selection of Aptamer for N-Methyl Mesoporphyrin IX to Develop Porphyrin Metalation DNAzyme. In: Steger, G., Rosenbach, H., Span, I. (eds) DNAzymes. Methods in Molecular Biology, vol 2439. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2047-2_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2047-2_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2046-5
Online ISBN: 978-1-0716-2047-2
eBook Packages: Springer Protocols