Abstract
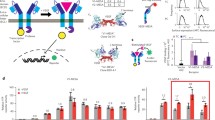
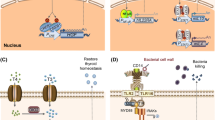
Synthetic receptors control cell behavior in response to environmental stimuli for applications in basic research and cell therapy. However, the integration of synthetic receptors in unexplored contexts is cumbersome, especially for nonspecialist laboratories. Here, I provide a detailed protocol on how to use receptors of the generalized extracellular molecule sensor (GEMS) platform. GEMS is a modular receptor system that can be adapted to sense molecules of choice by using affinity domains that dimerize in response to the target. GEMS consist of an erythropoietin receptor scaffold that has been mutated to no longer bind to erythropoietin. N-terminal fusions with affinity domains, such as single chain variable fragments (scFvs), that bind to two epitopes on the same target activate the receptor. The intracellular receptor domain can be chosen from several signal transduction domains of single-pass transmembrane receptors to activate endogenous signaling pathways. As of now, GEMS have been used for sensing prostate specific antigen (PSA), the synthetic azo dye RR120, caffeine, nicotine, rapamycin, the SunTag peptide, and a de novo designed protein displaying two viral epitopes. The tested intracellular domains were derived from FGFR1, IL-6RB, and VEGFR2, and were used to drive transgene expression from reporter plasmids responsive to the endogenous transcription factors STAT3, NFAT, NF-κB, and a synthetic transcription factor activated by the MAPK pathway. In this protocol, I focus on transient transfections of HEK293T cells and include several general notes about cell handling. While the described methods are optimized for experiments with GEMS, most of the described techniques are general procedures to set up synthetic biology experiments in mammalian cell culture. I outline how to generate stable cell lines and share tips on how to adapt GEMS for new ligands. The main objective of this protocol is to make the GEMS technology accessible also to nonspecialist laboratories to facilitate the use of synthetic receptors in new research contexts.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Scheller L, Fussenegger M (2019) From synthetic biology to human therapy: engineered mammalian cells. Curr Opin Biotechnol 58:108–116. https://doi.org/10.1016/j.copbio.2019.02.023
Lim WA, June CH (2017) The principles of engineering immune cells to treat cancer. Cell 168:724–740. https://doi.org/10.1016/j.cell.2017.01.016
Morsut L, Roybal KT, Xiong X et al (2016) Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell 164:780–791. https://doi.org/10.1016/j.cell.2016.01.012
Schwarz KA, Daringer NM, Dolberg TB, Leonard JN (2017) Rewiring human cellular input–output using modular extracellular sensors. Nat Chem Biol 13:202–209. https://doi.org/10.1038/nchembio.2253
Scheller L, Strittmatter T, Fuchs D et al (2018) Generalized extracellular molecule sensor platform for programming cellular behavior. Nat Chem Biol 14:723–729. https://doi.org/10.1038/s41589-018-0046-z
Bojar D, Scheller L, GC-E H et al (2018) Caffeine-inducible gene switches controlling experimental diabetes. Nat Commun 9:2318. https://doi.org/10.1038/s41467-018-04744-1
Yang C, Sesterhenn F, Bonet J et al (2020) Bottom-up de novo design of functional proteins with complex structural features. Nat Chem Biol https://doi.org/10.1038/s41589-020-00699-x
Querques I, Mades A, Zuliani C et al (2019) A highly soluble sleeping beauty transposase improves control of gene insertion. Nat Biotechnol 37:1502–1512. https://doi.org/10.1038/s41587-019-0291-z
Dunbar J, Deane CM (2015) ANARCI: antigen receptor numbering and receptor classification. Bioinformatics 32(2):298–300. https://doi.org/10.1093/bioinformatics/btv552
Berman HM (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
The UniProt Consortium (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515. https://doi.org/10.1093/nar/gky1049
Krawczyk K, Scheller L, Kim H, Fussenegger M (2020) Rewiring of endogenous signaling pathways to genomic targets for therapeutic cell reprogramming. Nat Commun 11:608. https://doi.org/10.1038/s41467-020-14397-8
Blainey P, Krzywinski M, Altman N (2014) Points of significance: replication. Nat Methods 11:879–880. https://doi.org/10.1038/nmeth.3091
Beal J, Haddock-Angelli T, Baldwin G et al (2018) Quantification of bacterial fluorescence using independent calibrants. PLoS One 13:e0199432. https://doi.org/10.1371/journal.pone.0199432
Acknowledgments
I thank Tobias Strittmatter and Sailan Shui for helpful discussions and help with the illustrations.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Scheller, L. (2021). Synthetic Receptors for Sensing Soluble Molecules with Mammalian Cells. In: Kojima, R. (eds) Mammalian Cell Engineering. Methods in Molecular Biology, vol 2312. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1441-9_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1441-9_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1440-2
Online ISBN: 978-1-0716-1441-9
eBook Packages: Springer Protocols