Abstract
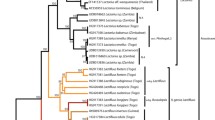
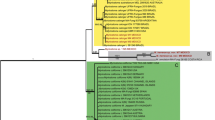
The large, white milkcaps of Lactifluus section Piperati are well known in the northern hemisphere. Historically, there was extensive debate about the number of European representatives and the diagnostic characteristics that delimit the species. Combining a morphological approach with a phylogenetic study, we aimed to resolve the problems in this section in Europe. Secondly, a molecular analysis of worldwide representatives of Lactifluus section Piperati was carried out, to verify whether there is intercontinental conspecificity. We compared nuclear ITS and LSU rDNA, nuclear protein-coding rpb2 and mitochondrial protein-coding atp6 genealogies to delimit species, using a concatenation of genes, along with Bayesian species delimitation for the European dataset. The phylogenetic analyses show the existence of two species in Europe: Lactifluus piperatus and Lactifluus glaucescens. Morphologically, the frequently used characteristics of the colouration of the latex and the macrochemical reactions of latex and context appear not to be useful as diagnostic characteristics to discriminate the species, but the microscopical characters of the pileipellis are informative. The preliminary overview of the section worldwide shows that it comprises at least 10 possible species divided over three clades, and that there is no intercontinental conspecificity.





Similar content being viewed by others
References
Barrie FR (2011) Report of the General Committee: 11. Taxon 60:1211
Basso MT (1999) Lactarius Pers. Fungi Europaei. Mykoflora, Alassio
Bataille F (1948) Les réactions macrochimiques chez les champignons suivies d’indications sur la morphologie des spores. Bull Trimest Soc Mycol Fr 63:1–172
Blum J (1966) Les Lactaires du groupe piperatus. Bull Soc Mycol Fr 82:241–247
Blum J (1976) Etudes mycologiques III. Les Lactaires. Lechevalier, Paris
Bon M (1980) Clé monographique du genre Lactarius (Pers. ex Fr.) S.F. Gray. Doc Mycol 10:1–85
Buyck B, Hofstetter V, Eberhardt U, Verbeken A, Kauff F (2008) Walking the thin line between Russula and Lactarius: the dilemma of Russula subsect. Ochricompactae. Fungal Divers 28:15–40
Buyck B, Hofstetter V, Verbeken A, Walleyn R (2010) Proposal 1919: To conserve Lactarius nom. cons. (Basidiomycota) with a conserved type. Mycotaxon 111:504–508
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Crossland C (1900) New and critical British fungi found in Western Yorkshire. Naturalist 1900:5–10
Damblon J, Darimont F, Lambinon J (1956) Contribution à l’étude de la flore mycologique de la haute et de la moyenne Belgique. Lejeunea 20:77–81
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4:699–710
Earle FS (1909) The genera of the North American gill fungi. Bull N Y Bot Gard 5:373–451
Eberhardt U (2000) Molekulare Analysen zur Verwandtschaft der agaricoiden Russulaceen im Vergleich mit Mykorrhiza- und Fruchtkörpermerkmalen. Dissertation
Fries EM (1821) Systema Mycologicum. Ex Officina Berlingiana. Lund, Sweden
Fuhrer B (2005) A field guide to Australian fungi. Bloomings Books Pty Ltd, Melbourne
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for Basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. doi:10.1111/j.1365-294X.1993.tb00005.x
Heilmann-Clausen J, Verbeken A, Vesterholt J (1998) The genus Lactarius Vol.2 – Fungi of Northern Europe. Svampetryk: Danish Mycological Society. 287 p. Svampetryk, Denmark
Heim R (1966) Breves diagnoses latinae novitatum genericarum specificarumque nuper descriptarum. Rev Mycol 30:231–241
Heinemann P (1960) Les Lactaires (2° édition). Naturalistes-Belges 41:133–156
Heled J, Drummond AJ (2010) Bayesian inference of species trees from multilocus data. Mol Biol Evol 27:570–580. doi:10.1093/molbev/msp274
Hesler LR, Smith AH (1979) North American species of Lactarius. University of Michigan Press, Ann Arbor
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:286–298. doi:10.1093/bib/bbn013
Knowles LL, Carstens BC (2007) Delimiting species without monophyletic gene trees. Syst Biol 56:887–895. doi:10.1080/10635150701701091
Kretzer AM, Bruns TD (1999) Use of atp6 in fungal phylogenetics: an example from the Boletales. Mol Phylogenet Evol 13:483–492
Kubatko LS, Degnan JH (2007) Inconsistency of phylogenetic estimates from concatenated data under coalescence. Syst Biol 56:17–24. doi:10.1080/10635150601146041
Le HT (2007) Biodiversity of the genus Lactarius (Basidiomycota) in northern Thailand. PhD dissertation, Chiang Mai University
Le HT, Nuytinck J, Verbeken A, Lumyong S, Desjardin DE (2007) Lactarius in Northern Thailand: 1. Lactarius subgenus Piperites. Fungal Divers 24:173–224
Leache AD, Fujita MK (2010) Bayesian species delimitation in West African forest geckos (Hemidactylus fasciatus). Proc R Soc Lond Ser B-Biol Sci 277:3071–3077. doi:10.1098/rspb.2010.0662
Lecomte M (2010) Lactarius piperatus et L. glaucescens, peut-être pas si simple que cela! Bull Assoc Mycol Francoph Belg 2010:37–46
Liu YJJ, Whelen S, Benjamin DH (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol 16:1799–1808
Maddison WP (1997) Gene trees in species trees. Syst Biol 46:523–536. doi:10.2307/2413694
Marchand A (ed) (1980) Champignons du nord et du midi 6. Lactaires et Pholoiotes. Société Mycologique des Pyrénées Méditerranéennes, Perpignan (66 000), Perpignan
Matheny PB (2005) Improving phylogenetic inference of mushrooms with RPB1 and RPB2 nucleotide sequences (Inocybe; Agaricales). Mol Phylogenet Evol 35:1–20. doi:10.1016/j.ympev.2004.11.014
McNeill J, Turland NJ, Monro AM, Lepschi BJ (2011) XVIII International Botanical Congress: preliminary mail vote and report of Congress action on nomenclature proposals. Taxon 60:1507–1520
Neuhoff W (1956) Die Milchlinge (Lactarii). In Die Pilze Mitteleuropas Bd. IIb. Julius Klinckhardt, Bad Heilbrunn
Norvell LL (2011) Report of the Nomenclature Committee for Fungi: 16. Taxon 60:223–226
Nuytinck J, Verbeken A (2003) Lactarius sanguifluus versus Lactarius vinosus – molecular and morphological analyses. Mycol Prog 2:227–234
Nylander JAA (2004) Mr.Modeltest v2. Program distributed by the author. Evolutionary Biology Centre:Uppsala University
Rambaut A, Drummond AJ (2007) Tracer v1.5. Available from http://beast.bio.ed.ac.uk/Tracer
Rannala B, Yang ZH (2003) Bayes estimation of species divergence times and ancestral population sizes using DNA sequences from multiple loci. Genetics 164:1645–1656
Romagnesi H (1956) Nouvel atlas des champignons. Tome I. Bordas, Paris
Romagnesi H (1961) Nouvel atlas des champignons. Tome III. Bordas, Paris
Romagnesi H (1980) Nouvelles observations sur les Lactaires blancs (Albati Bataille). Bull Soc Mycol Fr 96:73–95
Ronquist F, Teslenko M, van der Mark P, Ayres D, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. doi:10.1093/sysbio/sys029
Schaefer Z (1979) Beitrag zum Studium der Sektion Albates der Lactarien. Ceska Mykol 33:1–12
Singer R (1962) The Agaricales in modern taxonomy, 2nd edn. J. Cramer, Weinheim
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690. doi:10.1093/bioinformatics/btl446
Stamatakis A, Hoover P, Rougemont J (2008) A Rapid Bootstrap Algorithm for the RAxML Web Servers. Syst Biol 57:758–771. doi:10.1080/10635150802429642
Stubbe D, Nuytinck J, Verbeken A (2010) Critical assessment of the Lactarius gerardii species complex (Russulales). Fungal Biol 114:271–283. doi:10.1016/j.funbio.2010.01.008
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Thiers HD (1957) The Agaric flora of Texas. I. New species of Agarics and Boletes. Mycologia 49:707–722. doi:10.2307/3755988
Triantafyllou M, Polemis E, Dimou DM, Gonou-Zagou Z, Delivorias P, Zervakis GI (2011) A reappraisal of existing knowledge on the diversity of the genus Lactarius Pers. in Greece. Book of Abstracts, XVI congress of European Mycologists
Van de Putte K, Nuytinck J, Das K, Verbeken A (2012) Exposing hidden diversity by concordant genealogies and morphology-a study of the Lactifluus volemus (Russulales) species complex in Sikkim Himalaya (India). Fungal Divers 55:171–194. doi:10.1007/s13225-012-0162-0
Van de Putte K, Nuytinck J, Stubbe D, Huyen TL, Verbeken A (2010) Lactarius volemus sensu lato (Russulales) from northern Thailand: morphological and phylogenetic species concepts explored. Fungal Divers 45:99–130. doi:10.1007/s13225-010-0070-0
Vellinga EC (1988) Glossary. In: Bas CK, Kype TW, Noordeloos ME, Velliga EC (eds) Flora Agaricina Neerlandica, vol 1. AA Balkema, Rotterdam, pp 54–64
Verbeken A (1998a) Studies in tropical African Lactarius species. 5. A synopsis of the subgenus Lactifluus (Burl.) Hesler & A.H. Sm. emend. Mycotaxon 66:363–386
Verbeken A (1998b) Studies in tropical African Lactarius species. 6. A synopsis of the subgenus Lactariopsis (Henn.) R. Heim emend. Mycotaxon 66:387–418
Verbeken A, Fraiture A, Walleyn R (1997) Pepermelkzwammen en schaapjes in België (Bijdragen tot de kennis van het genus Lactarius in België. 4. De sectie Albati ss. auct. pl. Mededelingen Antwerpse Mycologische Kring 1997:48–64
Verbeken A, Horak E (1999) Lactarius (Basidiomycota) in Papua New Guinea. 1. Species of tropical lowland habitats. Aust Syst Bot 12:767–779
Verbeken A, Van de Putte K, De Crop E (2012) New combinations in Lactifluus. 3. L. subgenera Lactifluus and Piperati. Mycotaxon 120:443–450. doi:10.5248/120.443
Verbeken A, Walleyn R (2010) Monograph of Lactarius in tropical Africa. Fungus Flora of Tropical Africa, vol 2. National Botanic Garden, Belgium
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: A guide to methods and applications. Academic, New York, pp 315–322
Yang ZH, Rannala B (2010) Bayesian species delimitation using multilocus sequence data. Proc Natl Acad Sci U S A 107:9264–9269. doi:10.1073/pnas.0913022107
Acknowledgments
The first author is funded by the “Bijzonder Onderzoeksfonds Ghent University” (BOF). The Bayesian analyses were carried out using the Stevin Supercomputer Infrastructure at Ghent University, funded by Ghent University, the Hercules Foundation and the Flemish Government – department EWI. We thank Omer Van de Kerckhove and Marina Triantafyllou for the helpful discussions concerning the European Piperati.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
De Crop, E., Nuytinck, J., Van de Putte, K. et al. Lactifluus piperatus (Russulales, Basidiomycota) and allied species in Western Europe and a preliminary overview of the group worldwide. Mycol Progress 13, 493–511 (2014). https://doi.org/10.1007/s11557-013-0931-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-013-0931-5