Abstract
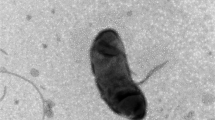
An aerobic, Gram-stain-negative, non-motile, non-spore-forming, rod-shaped and pink-colored bacterial strain, designated BRD72T, was isolated from a crater lake (Baengnokdam) at the top of Mt. Hallasan in the Republic of Korea. Cells were catalase-positive and oxidase-negative. Phylogenetic analysis based on the 16S rRNA gene sequences revealed that the isolate was a member of the genus Hymenobacter and most closely related to Hymenobacter marinus KJ035T (96.2% similarity). The isolate was found to produce carotenoid pigment, but not flexirubin-type pigment. The predominant fatty acids of strain BRD72T were summed feature 3 (C16:1 ω7c and/or C16:1 ω6c, 21.6%), iso-C15:0 (17.9%), anteiso-C15:0 (13.3%) and summed feature 4 (iso-C17:1 I and/or anteiso-C17:1 B, 11.3%). The major polar lipids were phosphatidylethanolamine, an unidentified amino lipid, and two unidentified aminophospholipids. The main respiratory quinone was menaquinone-7 (MK-7), and the main polyamine was homospermidine. The DNA G+C content was 59.8 mol%. Based on the phylogenetic, physiological, and chemotaxonomic characteristics, strain BRD72T represents a novel species, for which the name Hymenobacter baengnokdamensis sp. nov. is proposed. The type strain is BRD72T (= KCTC 72649T = JCM 33837T).

Similar content being viewed by others
References
Hirsch P, Ludwig W, Hethke C, Sittig M, Hoffmann B, Gallikowski CA (1998) Hymenobacter roseosalivarius gen. nov., sp. nov. from continental Antartica soils and sandstone: bacteria of the Cytophaga/Flavobacterium/Bacteroides line of phylogenetic descent. Syst Appl Microbiol 21:374–383
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.004332
Buczolits S, Denner EB, Vybiral D, Wieser M, Kämpfer P, Busse HJ (2002) Classification of three airborne bacteria and proposal of Hymenobacter aerophilus sp. nov. Int J Syst Evol Microbiol 52:445–456
Kim KH, Im WT, Lee ST (2008) Hymenobacter soli sp. nov., isolated from grass soil. Int J Syst Evol Microbiol 58:941–945
Hoang VA, Kim YJ, Nguyen NL, Yang DC (2013) Hymenobacter ginsengisoli sp. nov., isolated from soil of a ginseng field. Int J Syst Evol Microbiol 63:661–666
Kang H, Kim H, Joung Y, Kim KJ, Joh K (2016) Hymenobacter marinus sp. nov., isolated from coastal seawater. Int J Syst Evol Microbiol 66:2212–2217
Sheu SY, Hsieh TY, Kwon SW, Chen WM (2018) Hymenobacter rivuli sp. nov., isolated from a freshwater creek. Int J Syst Evol Microbiol 68:1220–1226
Zhang G, Niu F, Busse HJ, Ma X, Liu W, Dong M, Feng H, An L, Cheng G (2008) Hymenobacter psychrotolerans sp. nov., isolated from the Qinghai-Tibet Plateau permafrost region. Int J Syst Evol Microbiol 58:1215–1220
Klassen JL, Foght JM (2011) Characterization of Hymenobacter isolates from Victoria Upper Glacier, Antarctica reveals five new species and substantial non-vertical evolution within this genus. Extremophiles 15:45–57
Sedláček I, Pantůček R, Holochová P, Králová S, Staňková E, Vrbovská V, Šedo O, Švec P, Busse HJ (2019) Hymenobacter humicola sp. nov., isolated from soils in Antarctica. Int J Syst Evol Microbiol 69:2755–2761
Zhang L, Dai J, Tang Y, Luo X, Wang Y, An H, Fang C, Zhang C (2009) Hymenobacter deserti sp. nov., isolated from the desert of Xinjiang China. Int J Syst Evol Microbiol 59:77–82
Liang Y, Tang K, Wang Y, Yuan B, Tan F, Feng F, Liu H (2019) Hymenobacter crusticola sp. nov., isolated from biological soil crust. Int J Syst Evol Microbiol 69:547–551
Collins MD, Huston RA, Grant IR, Patterson MF (2000) Phylogenetic characterization of a novel radiation-resistant bacterium from irradiated pork: description of Hymenobacter actinosclerus sp. nov. Int J Syst Evol Microbiol 50:731–734
Dai J, Wang Y, Zhang L, Tang Y, Luo X, An H, Fang C (2009) Hymenobacter tibetensis sp. nov., a UV-resistant bacterium isolated from Qinghai-Tibet plateau. Syst Appl Microbiol 32:543–548
Klassen JL, Foght JM (2008) Differences in carotenoid composition among Hymenobacter and related strains support a tree-like model of carotenoid evolution. Appl Environ Microbiol 74:2016–2022
Liu L, Zhou EM, Jiao JY, Manikprabhu D, Ming H, Huang MJ, Yin YR, Li WJ (2015) Hymenobacter mucosus sp. nov., isolated from a karst cave soil sample. Int J Syst Evol Microbiol 65:4121–4127
Marizcurrena JJ, Herrera LM, Costábile A, Morales D, Villadóniga C, Eizmendi A, Davyt D, Castro-Sowinski S (2019) Validating biochemical features at the genome level in the Antarctic bacterium Hymenobacter sp. strain UV11. FEMS Microbiol Lett 366:fnz177
Chun J, Goodfellow M (1995) A phylogenetic analysis of the genus Nocardia with 16S rRNA gene sequences. Int J Syst Bacteriol 45:240–245
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Felsenstein J (1993) PHYLIP (phylogeny inference package), version 3.5c., Distributed by the author. Department of Genome Sciences, University of Washington, Seattle.
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Chin CS, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE et al (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10:563–569
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069
Huerta-Cepas J, Szklarczyk D, Forslund K, Cook H, Heller D, Walter MC, Rattei T, Mende DR, Sunagawa S, Kuhn M et al (2016) eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucl Acids Res 44:286–293
Dawson RMC, Elliot DC, Elliot WH, Jones KM (1986) Data for biochemical research, 3rd edn. Oxford Science Publ, Oxford
Miller PH, Wiggs LS, Miller JM (1995) Evaluation of AnaeroGen system for growth of anaerobic bacteria. J Clin Microbiol 33:2388–2391
Bernardet JF, Nakagawa Y, Holmes B (2002) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52:1049–1070
Powers EM (1995) Efficacy of the Ryu nonstaining KOH technique for rapidly determining gram reactions of food-borne and waterborne bacteria and yeasts. Appl Environ Microbiol 61:3756–3758
Kovacs N (1956) Identification of Pseudomonas pyocyanea by the oxidase reaction. Nature 178:703
Smibert RM, Krieg NR (1994) General characterization. In: Gebhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. DE: MIDI Inc, Newark
Minnikin DE, Patel PV, Alshamaony L, Goodfellow M (1977) Polar lipid composition in the classification of Nocardia and related bacteria. Int J Syst Bacteriol 27:104–117
Schenkel E, Berlaimont V, Dubois J, Helson-Cambier M, Hanocq M (1995) Improved high-performance liquid chromatographic method for the determination of polyamines as their benzoylated derivatives: application to P388 cancer cells. J Chromatogr B 668:189–197
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Ahn DH, Han SR, Oh TJ, Park H (2016) Complete genome sequence of ionizing radiation-resistant Hymenobacter sp. strain PAMC26628 isolated from an Arctic lichen. J Biotechnol 223:50–51
Kim MK, Joo ES, Lee SY, Lee DS, Srinivasan S, Jung HY (2016) Complete genome sequence of Hymenobacter sp. DG25B, a novel bacterium with gamma-radiation resistance isolated from soil in South Korea. J Biotechnol 217:98–99
Acknowledgements
This research was supported by the Nuclear R&D program of the Ministry of Science and ICT (MSIT), Republic of Korea.
Author information
Authors and Affiliations
Contributions
JHL performed the experiments and wrote the manuscript. JHJ analyzed the genome sequence. HSS and JZ isolated and identified the strain. HNC and SC tested the chemotaxonomic characterizations. MKK and CNS guided the experiments and data analysis. SL conceived the study and was in charge of overall direction and planning. All authors approved the manuscript and its submission.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Accession number
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain BRD72T is MK752551. The GenBank/EMBL/DDBJ accession number for the whole genome sequence of strain BRD72T is CP044285.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J.H., Kim, MK., Jung, JH. et al. Hymenobacter baengnokdamensis sp. nov., Isolated from the Soil of a Crater Lake in Korea. Curr Microbiol 77, 4167–4173 (2020). https://doi.org/10.1007/s00284-020-02225-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02225-7