Abstract
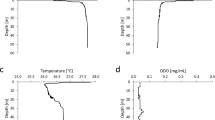
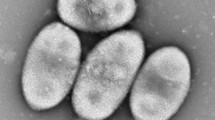
In this study, we report a novel Gram-positive bacterium, designated as strain CS13T, isolated from deep-sea sediment collected in the cold seep area of the South China Sea. Growth of strain CS13T occurred at 16–37 °C (optimum 25–28 °C), pH 7.0–9.0 (optimum, 7.0), and 0–8% (w/v) NaCl (optimum, 2–3%). Phylogenetic analysis based on 16S rRNA gene sequence indicated that strain CS13T belonged to the genus Bacillus. The closest phylogenetic neighbors of strain CS13T are Bacillus carboniphilus JCM 9731T (96.0%), Bacillus pakistanensis NCCP-168T (95.7%) and Bacillus acidicola 105-2T (95.6%). The genomic DNA G + C content of strain CS13T is 43.7 mol%. The principal respiratory quinone was menaquinone 7 (MK—7). The polar lipids of CS13T contained diphosphatidylglycerol, phosphatidylglycerol, phospholipid, and glycolipid. The major fatty acids of CS13T contained anteiso-C15:0, anteiso-C17:0, C16:0 and C18:0. Strain CS13T harboured meso-diaminopimelic acid as the diagnostic diamino acid. Phylogenetic, physiological, biochemical, and morphological analyses suggested that strain CS13T represents a novel species of genus Bacillus, and the name Bacillus fonticola sp. nov. is proposed for the type species CS13T (= CCTCC AB 2019194T = JCM 33663T).

Similar content being viewed by others
Data availability
The GenBank accession number of the 16S rRNA gene sequence of strain CS13T is MT359011. The GenBank accession number of complete genome sequence of strain CS13T is CP052135.
Abbreviations
- ANI:
-
Average nucleotide identity
- DDH:
-
DNA–DNA hybridization
- DPG:
-
Diphosphatidylglycerol
- DSMZ:
-
Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH
- FAMEs:
-
Fatty acid methyl esters
- GL:
-
Glycolipid
- MA:
-
Marine agar
- ME:
-
Minimum evolution
- MK-7:
-
Menaquinone 7
- ML:
-
Maximum likelihood
- NJ:
-
Neighbor joining
- PL:
-
Phospholipid
- PG:
-
Phosphatidylglycerol
- pNPG:
-
P-nitrophenyl-β-d-galactopyranoside
- rRNA:
-
Ribosome RNA
- tRNA:
-
Transfer RNA
References
Alber RA, Archambault J, Rosselló-Mora R, Tindall BJ, Matheny M (2005) Bacillus acidicola sp. nov., a novel mesophilic, acidophilic species isolated from acidic Sphagnum peat bogs in Wisconsin. Int J Syst Evol Microbiol 55:2125–2130. https://doi.org/10.1099/ijs.0.02337-0
Azmatunnisa M, Rahul K, Subhash Y, Sasikala C, Ramana CV (2015) Bacillus oleivorans sp. nov., a diesel oil-degrading and solvent-tolerant bacterium. Int J Syst Evol Microbiol 65:1310–1315. https://doi.org/10.1099/ijs.0.000103
Besemer J, Lomsadze A, Borodovsky M (2001) GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res 9:2607–2618. https://doi.org/10.1093/nar/29.12.2607
Cohn F (1872) Untersuchungen Über Bakterien. Beitr Biol Pflanz 1:127–224 (in German)
Cui X, Wang Y, Liu J, Chang M, Zhao Y, Zhou S, Zhuang L (2015) Bacillus dabaoshanensis sp. nov., a Cr(VI)-tolerant bacterium isolated from heavy-metal-contaminated soil. Arch Microbiol 197:513–520. https://doi.org/10.1007/s00203-015-1082-7
Fujita T, Shida O, Takagi H, Kunugita K, Pankrushina AN, Matsuhash M (1996) Description of Bacillus carboniphilus sp. nov. Int J Syst Evol Microbiol 46:116–118. https://doi.org/10.1099/00207713-46-1-116
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Hasegawa T, Takizawa M, Tanida S (1983) A rapid analysis for chemical grouping of Aerobic actinomycetes. J Gen Appl Microbiol 29:319–322. https://doi.org/10.2323/jgam.29.319
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721. https://doi.org/10.1099/ijs.0.038075-0
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lagesen K, Hallin P, Rødland EA, Staerfeldt HH, Rognes T, Ussery DW (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108. https://doi.org/10.1093/nar/gkm160
Lee I, Ouk Kim Y, Park S-C, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Liu YJ, Long LJ, Huang XF, You ZQ, Wang FZ, Li J, Kim CJ, Tian XP, Zhang S (2013) Bacillus oceani sp nov., a new slightly halophilic bacterium, isolated from a deep sea sediment environment. Anton Leeuw Int J G 104:829–836. https://doi.org/10.1007/s10482-013-9995-0
Liu GH, Narsing Rao MP, Dong ZY, Wang JP, Chen Z, Liu B, Li WJ (2019) Two novel alkaliphiles, Bacillus alkalisoli sp. nov., and Bacillus solitudinis sp. nov., isolated from saline-alkali soil. Extremophiles 23:759–764. https://doi.org/10.1007/s00792-019-01127-2
Logan NA, Lebbe L, Verhelst A, Goris J, Forsyth G, Rodríguez-Díaz M, Heyndrickx M, De Vos P (2004) Bacillus shackletonii sp. nov., from volcanic soil on Candlemas Island, South Sandwich archipelago. Int J Syst Evol Microbiol 54:373–376. https://doi.org/10.1099/ijs.0.02661-0
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 5:955–964. https://doi.org/10.1093/nar/25.5.955
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013a) Genome sequencebased species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. https://doi.org/10.1186/1471-2105-14-60
Meier-Kolthoff JP, Göker M, Sproer C, Klenk HP (2013b) When should a DDH experiment be mandatory in microbial taxonomy? Arch Microbiol 195:413–418. https://doi.org/10.1007/s00203-013-0888-4
Menes RJ, Machin EV, Iriarte A, Langleib M (2019) Bacillus natronophilus sp. nov., an alkaliphilic bacterium isolated from a soda lake. Int J Syst Evol Microbiol 70:562–568. https://doi.org/10.1099/ijsem.0.003792
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Meth 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Roohi A, Ahmed I, Paek J, Sin Y, Abbas S, Jamil M, Chang YH (2014) Bacillus pakistanensis sp. nov., a halotolerant bacterium isolated from salt mines of the Karak Area in Pakistan. Antonie Van Leeuwenhoek 105:1163–1172. https://doi.org/10.1007/s10482-014-0177-5
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC Newsl 20:1–6
Seuylemezian A, Ott L, Wolf S, Fragante J, Yip O, Pukall R, Schumann P (2020) Bacillus glennii sp. nov. and Bacillus saganii sp. nov., isolated from the vehicle assembly building at Kennedy Space Center where the Viking spacecraft were assembled. Int J Syst Evol Microbiol 70:71–76. https://doi.org/10.1099/ijsem.0.003714
Shin B, Park C, Lee BH, Lee KE, Park W (2020) Bacillus miscanthi sp. nov., a alkaliphilic bacterium from the rhizosphere of Miscanthus sacchariflorus. Int J Syst Evol Microbiol 70:1843–1849. https://doi.org/10.1099/ijsem.0.003982
Sun QL, Yu C, Luan ZD, Lian C, Hu YH, Sun L (2018) Description of Bacillus kexueae sp. nov. and Bacillus manusensis sp. nov. isolated from hydrothermal sediments. Int J Syst Evol Microbiol 68:829–834. https://doi.org/10.1099/ijsem.0.002594
Xu P, Li WJ, Tang SK, Zhang YQ, Chen GZ, Chen HH, Xu LH, Jiang CL (2005) Naxibacter alkalitolerans gen. nov., sp nov., a novel member of the family ‘Oxalobacteraceae’ isolated from China. Int J Syst Evol Microbiol 55:1149–1153. https://doi.org/10.1099/ijs.0.63407-0
Xu X, Yu L, Xu G, Wang Q, Wei S, Tang X (2020) Bacillus yapensis sp. nov., a novel piezotolerant bacterium isolated from deep-sea sediment of the Yap Trench. Pacific Ocean Antonie Van Leeuwenhoek 113:389–396. https://doi.org/10.1007/s10482-019-01348-7
Acknowledgements
The authors acknowledge the support of the Research Vessel KEXUE of the National Major Science and Technology Infrastructure from the Chinese Academy of Sciences. This work was funded by the grants of the National Key R&D Program of China (2018YFC0310801), Qingdao National Laboratory for Marine Science and Technology (QNLM2016ORP0309), The Senior User Project of RV KEXUE (KEXUE2018G20), and the Taishan Scholar Program of Shandong Province.
Author information
Authors and Affiliations
Contributions
QLS designed the work. YYS and HZZ conducted the laboratory experiments. YYS analyzed the data and drafted the manuscript. QLS supervised the study and revised the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sun, Yy., Zhou, Hz. & Sun, Ql. Bacillus fonticola sp. nov., isolated from deep sea cold seep sediment. Arch Microbiol 203, 4127–4132 (2021). https://doi.org/10.1007/s00203-021-02401-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02401-8