Abstract
A special methionyl-tRNA (RNAi) is universally required to initiate translation. The conservation of this reactant throughout evolution, as well as its unusual decoding properties, suggested an alternate mechanism for tRNA-mRNA interactions at initiation. We have reported that the sequence of bases neighboring the start codons of many eubacterial genes are complementary not only to the 16S rRNA 3′ end and to the anticodon of tRNAi, but, also, have the potential to base-pair the D, T or extended anticodon loops of this tRNAi. The coding properties of tRNAi and mutations that affect translation suggest that these signals may function. This hypothesis explains the observation that unusual triplets can start prokaryotic and mitochondrial genes and predicts the occurrence of other reading frames. Furthermore, it suggests a unifying model of chain initiation based on RNA-RNA contacts and displacements.
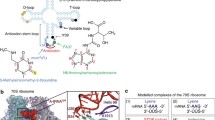
Here we examine the start domain of 290 eukaryotic genes for their ability to base-pair the tRNAi loops and the 18S rRNA. We observe that both methionine start, and methionine coding regions have the potential to pair with the 18S rRNA, but that the nucleotide distribution about start codons strongly favoured such pairings over that near internal AUGs. The 5′ extended anticodon of tRNAi is methylated, and was not represented in the mRNA with high frequency. However, the tetramer AUGg did occur with high frequency in the start domain. A modification of the tRNAi T loop also decreases its base-pairing potential. Interestingly, complementarity to the T loop did not occur with high frequency in the start sites. The early coding region, 10 to 34 nucleotides 3′ to the initiator AUG, is complementary to the tRNAi D loop in many cases, while no such affinity is found near internal AUGs.
The nucleotides around initiator AUGs were heavily biassed toward the sequence gccaccAUGgcg. No such tendency was noted around internal AUGs. Although the role of this sequence bias is unclear, the sequence gccaccAUGg has been shown by Kozak to promote initiation. Another distinguishing feature was a C-rich tract 7 to 34 nucleotides 5′ to the initiator AUGs.
Ability to pair with more than eight bases of the start consensus sequence, matching of 6 or 7 nucleotides to the D loop on the 3′ side, an C-richness on the 5′ side were used as criteria for distinguishing start AUGs. The program successfully identified over 52% of the sequences submitted to it, wrongly identified less than 4% and labelled the rest as uncertain suggesting a promising approach to reliable detection of eukaryote genetic reading frames.
Similar content being viewed by others
References
GoldL, PribnowD, SchneiderT, ShinedlingS, SingerBS & StormoG (1981) Ann. Rev. Microbiol. 35: 365–403
SteitzJA (1979) In: GodbergerRF (ed) Biological Regulation and Development, Plenum Press, New York, Vol. I, pp. 349–399
KozakM (1983) Microbiol. Rev. 47: 1–45
ShineJ & DalgarnoL (1975) Nature (London) 254: 34–38
SteitzJA & JakesK (1975) Proc. Natl. Acad. Sci. USA 72: 4734–4738
NeilsonT, KofoidEC & GanozaMC (1980) Nucl. Acids Res. Symp. Ser. 7: 321–323
DunnJJ, Buzash-PollertE & StudierFW (1978) Proc. Natl. Acad. Sci. USA 75: 2741–2745
PtashneM, BackmanK, HumayunMZ, JeffreyA, MaurerR, MeyerB & SauerRT (1976) Science 194: 156–161
BeckE, SommerR, AuerswaldEA, KurzC, ZinkB, OsterburgG, ShalderH, SugimotoK, SugisakiH, OkamotoT & TakanamiM (1978) Nucl. Acids Res. 5: 4495–4503
FarabaughPJ (1978) Nature (London) 274: 765–769
GodsonGN, BarrellBG, StadenR & FiddesJC (1978) Nature (London) 276: 236–247
PirrotaB (1979) Nucl. Acids Res. 6: 1495–1508
SingletonCK, RoederWD, BogosianG, SomervilleRL & WeithHL (1980) Nucl. Acids Res. 8: 1551–1560
RobertsTM, KacichR & PtashneM (1979) Proc. Natl. Acad. Sci. USA. 76: 760–764
HagenbuchleO, SanterM & SteitzJA (1978) Cell 13: 551–563
SarganDR, GregorySP & ButterworthPHW (1982) FEBS Lett. 47: 133–136
SalserW (1978) Cold Spring Harbor Symp. Quant. Biol. 42: 985–1002
BothGW (1979) FEBS Lett. 101: 220–224
MarounLE, DegnerM, PrecupJW & FranciskovichPP (1987) J. Theor. Biol. 91: 85–98
AzadAA & DeaconNJ (1979) Biochem. Biophys. Res. Commun. 86: 568–576
NakashimaK, DarzynkiewiczE & ShatkinA (1980) Nature (London) 286: 226–231
SchroederHW, LiarakosCD, GuptaRC, RanderathK & O'MalleyBW (1979) J. Biochem. 18: 5798–5808
YamaguchiK, HidakaS & MiuraKL (1982) Proc. Natl. Acad. Sci. USA. 79: 1012–1016
DeWachterR (1979) Nucl. Acids Res. 7: 2045–2054
KozakM (1987) Nucl. Acids Res. 15: 8125–8148
ShermanF & StewartJW (1983) In: StruthersJN, JonesEW & BirachJR, (eds) The Molecular Biology of Saccharomyces cerevisiae: Metabolism and Gene Expression, Cold Spring Harbor Laboratory, New York, pp. 301–334
BaimSB, GoodhueCT, PietrasDF, EusticeDC, LabhardM, FriedmanLR, HampseyMD, StilesIT & ShermanF (1985) In: CalenderR & GoldL (eds) Sequence Specificity in Transcription and Translation, Alan R. Liss Inc., New York, pp. 351–362
BaralleFE & BrownleeGG (1978) Nature (London) 274: 84–87
GenBank ref. 44.0 (August 1986), IBM PC-format floppydisk version. Bolt, Beranek and Newman, Inc., distr. by IRL Press, McLean, VA, U.S.A.
SchneiderC, OwenMJ, BanvilleD & WilliamsJG (1984) Nature 311: 675–678
SorgeJ, WestC, WestwoodB & BeutlerE (1985) Proc. Natl. Acad. Sci. USA. 82: 7289–7293
OuJ-H, MasiarzF, KanYM, GoldfineID, RothRA & RutterWJ (1985) Cell 40: 747–758
HurleyJB, FongHKW, TeplowDB, DreyerWJ, SimonMI (1984) Proc. Natl. Acad. Sci. USA 81: 6948–6952
KimuraS, GonzalesFJ & NebertDW (1984) J. Biol. Chem. 259: 10705–10713
LinzerDIH & TalamantesF (1985) J. Biol. Chem. 260: 9574–9579
KozacM (1978) Cell 15: 1109–1123
FiersW, ContrerasR, HaegemanG, RogiersR, Van deVoordeA, VanHeuverswynH, VanHerrewegheJ, VolckaertG & YsebaertM (1978) Nature (London) 273: 113–120
ContrerasR, RogiersR, Van deVoordeA & FiersW (1977) Cell 12: 529–538
GanozaMC, KofoidEC, MarlièreP & LouisBG (1987) Nucl. Acids Res. 15: 345–360
Louis BG & Ganoza MC (1987) Cold Spring Harbour Abstracts: Translational Control, Cold Spring Harbour, NY, USA, p. 110
GanozaMC, MarlièreP, KofoidEC & LouisBG (1985) Proc. Natl. Acad. Sci. USA. 82: 4587–4595
GanozaMC, FraserA & NeilsonT (1978) Biochemistry 17: 2769–2775
GanozaMC, SullivanP, CunninghamC, KofoidEC, HaderP & NeilsonT (1982) J. Biol. Chem. 257: 8228–8232
EckhardtH & LuhrmannR (1981) Biochemistry 20: 2075–2080
TaniguchiT & WeissmannC (1978) J. Mol. Biol. 118: 533–565
SchmittM, ManderscheidU, KyriatsoulisA, BrinkmannU & GassenHG (1980) Eur. J. Biochem. 109: 291–299
GanozaMC (1977) Can. J. Biochem. 55: 257–281
EMBL Gene Sequence Library, Rel. 3.0 (1983). Eur. Mol. Biol. Organization, Heidelberg
StormoG, SchneiderTD & GoldL (1982) Nucl. Acids Res. 10: 2791–2996
GanozaMC (1988) In: Kleinkauf, vonDohren & Jalnicke, (eds) The Roots of Modern Biochemistry, Walter de Gruyter and Co., Berlin, New York, pp. 541–549
HamiltonR, WatanabeCK & deBoerH (1987) Nucl. Acids Res. 15: 3581–3593
RichA & Raj BhandaryUL (1976) Annu. Rev. Biochem. 45: 805–860
JayG & KaempferR (1975) J. Biol. Chem. 250: 5742–5748
CunninghamC & GanozaMC (1984) Mol. Biol. Rep. 10: 115–121
UhlenbeckOC (1972) J. Mol. Biol. 65: 25–41
WooNH, RoeBA & RichA (1980) Nature (London) 286: 346–351
SchevitzRW, PodjarnyAD, KrisnamachariN, HughesJJ & SiglerPB (1979) Nature (London) 278: 188–190
GaussDH & SprizlM (1981) Nucl. Acids Res. 9: 1–23
FickettJW (1982) Nucl. Acids Res. 10: 5303–5318
MichelCJ (1986) J. Theor. Biol. 120: 223–236
BibbMJ, FindlayPR & JohnsonMW (1984) Gene 30: 157–166
StadenR & McLachlanAD (1982) Nucl. Acids Res. 10: 141–156
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Louis, B.G., Ganoza, M.C. Signals determining translational start-site recognition in eukaryotes and their role in prediction of genetic reading frames. Mol Biol Rep 13, 103–115 (1988). https://doi.org/10.1007/BF00539058
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00539058