Abstract
Mutations of the gene (TNNT2) encoding the thin-filament contractile protein cardiac troponin T are responsible for 15% of all cases of familial hypertrophic cardiomyopathy, the leading cause of sudden death in young athletes1,2. Mutant proteins are thought to act through a dominant-negative mode that impairs function of heart muscle3. TNNT2 mutations can also lead to dilated cardiomyopathy, a leading cause of heart failure4. Despite the importance of cardiac troponin T in human disease, its loss-of-function phenotype has not been described. We show that the zebrafish silent heart (sih) mutation affects the gene tnnt2. We characterize two mutated alleles of sih that severely reduce tnnt2 expression: one affects mRNA splicing, and the other affects gene transcription. Tnnt2, together with α-tropomyosin (Tpma) and cardiac troponins C and I (Tnni3), forms a calcium-sensitive regulatory complex within sarcomeres5. Unexpectedly, in addition to loss of Tnnt2 expression in sih mutant hearts, we observed a significant reduction in Tpma and Tnni3, and consequently, severe sarcomere defects. This interdependence of thin-filament protein expression led us to postulate that some mutations in tnnt2 may trigger misregulation of thin-filament protein expression, resulting in sarcomere loss and myocyte disarray, the life-threatening hallmarks of TNNT2 mutations in mice and humans6,7.
Similar content being viewed by others
Main
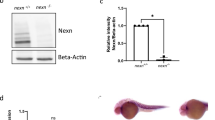
Forward genetics in zebrafish has led to the identification of several mutations affecting cardiac contractility8,9. The most severe and heart-specific of these is sih, which causes a non-contractile heart phenotype (Fig. 1). Skeletal and smooth-muscle function remain intact in sih mutant embryos, as evidenced by their ability to hatch, swim and show gut peristalsis. One γ-ray–induced allele (sihb109) and one chemically induced allele (sihtc300b) exist8. Both of these mutated alleles are fully penetrant and recessive lethal, and sihb109 and sihtc300b mutant embryos are phenotypically indistinguishable. As sih embryos age, pericardial edema develops, the endocardium peels away from the myocardium and the embryos die around seven days post fertilization. Until that time, embryos survive on diffused oxygen and are not dependent on circulating blood10.
a,b, Wildtype (a) and sih mutant (b) embryos at 32 hpf. A sihb109 mutant embryo is shown; sihtc300b mutants appear identical. Note blood cells in the heart of the wildtype embryo (a, arrow) and lack of blood cells in the heart and pericardial edema in the sihb109 mutant embryo (b, arrow). No intermediate heterozygote phenotype was observed. c,d, Ventral view; anterior to the top. Normal cardiac looping and chamber differentiation is seen at 48 hpf in wildtype (c) and sihb109 mutant (d) hearts. MF20 (TRITC) stains the entire heart tube and S46 (FITC) stains only the atrium. In double-exposure, the ventricle (v) fluoresces red and the atrium (a) fluoresces yellow. The stretched appearance of the mutant heart tube (d) is probably due to pericardial edema. Notably, even in the absence of contraction, the heart forms and loops normally.
To determine whether the sih phenotype is due to defects in cellular excitation or excitation–contraction coupling, we devised a new assay using the fluorescent calcium indicator Ca2+ green. In wildtype hearts, a wave of fluorescence representing Ca2+ influx into cardiomyocytes precedes the contractile wave and progresses from the venous to the arterial end (see Web Movie A and Web Note A online). In mutant hearts, we also observed regular waves of fluorescence, but these were not followed by contraction (Fig. 2a–d). On the basis of this assay, cellular excitation seemed to be intact in mutant cardiomyocytes, indicating that the absence of contractility results from abnormalities downstream of calcium influx.
a–d, In the sih mutant heart, Ca2+ entry into cardiomyocytes proceeded as a wave (arrowheads, a–c) starting in the atrium (a) and moving through the atrio-ventricular junction into the ventricle (v), before reaching a resting state (d). e,f, Longitudinal sections of wildtype (e) and sihb109 mutant (f) cardiomyocytes at 48 hpf as observed by electron microscopy. A-band, I-bands and Z-lines are visible in a wildtype sarcomere (e). The arrow in f points to disorganized thick filaments seen in sihb109 mutant cardiomyocytes. g,h, Cross-sections showing tightly bundled, hexagonal arrays of thick and thin filaments in a wildtype cardiomyocyte (arrow, g), and loosely scattered thick filaments, without intervening thin filaments, in a sihb109 mutant cardiomyocyte (h).
The alignment of thick and thin filaments into sarcomeres creates a highly ordered ultrastructural pattern that is evident in wildtype zebrafish cardiomyocytes at 48 hours post-fertilization (hpf) by electron microscopy (Fig. 2e). By contrast, sihb109 mutant cardiomyocytes at this stage show only a few loosely organized thick filaments near the cell periphery and no thin filaments or electron dense Z-disks (Fig. 2f–h). These results suggest that sih influences the assembly and stability of thin filaments, which are required for sarcomere assembly.
Using whole-mount immunohistochemistry, we examined the expression of thick- and thin-filament contractile proteins in wildtype and sih mutant embryos from both alleles (Fig. 3). Consistent with the presence of thick filaments as seen by electron microscopy, myosin heavy chain is expressed at wildtype levels throughout the mutant heart tubes (Fig. 3b). α-cardiac actin and α-actinin, the earliest Z-disk protein, also seem to be expressed at wildtype levels (Fig. 3d and data not shown). By contrast, thin-filament proteins of the tropomyosin–troponin (Tpma–Tn) complex are altered in sih mutant hearts. Expression of Tpma seems slightly reduced (Fig. 3f), and cardiac troponin T (Tnnt2) and Tnni3 are undetectable (Fig. 3h,j). Heterozygous sih embryos show no obvious reduction in protein expression. The Tpma-Tn complex contains regularly spaced troponin subunits that are anchored to Tpma and regulate contraction in response to calcium5. The sih mutation might directly affect expression of Tpma, Tnnt2 and Tnni3, or reduce expression of one of these proteins and secondarily cause instability or degradation of the others. Some precedence for such secondary effects exists in Drosophila melanogoster indirect flight-muscle mutants, where a lack of troponin T results in secondary reduction of tropomyosin and actin11. Because the cellular and molecular phenotypes of sih mutants most closely resemble those of troponin T mutants in flies, we pursued tnnt2 as a candidate for the sih mutation, and in parallel mapped the sihgene.
All panels show results from the sihb109 allele; sihtc300b mutants show exactly the same phenotype. a–j, Lateral views, anterior to the left, dorsal to the top, heart (arrowhead). Stains: myosin heavy chain, MF20-TRITC (a,b), α-actinin-FITC (c,d), Tpma, CH1-FITC (e,f), Tnnt2, JLT-12-FITC (g,h) and Tnni3, IE7-FITC (i,j). The same wildtype embryo is shown in a and g, and the same mutant embryo is shown in b and h. Staining is also evident in skeletal muscle and serves as an internal control. Embryos were fixed at 30–32 hpf, except those shown in i and j, which were fixed at 24 hpf.
We cloned zebrafish tnnt2 by immunoscreening an adult zebrafish heart expression library (see Web Fig. A online). In wildtype embryos, tnnt2 mRNA is expressed by the 15-somite stage (16.5 hpf) in the bilateral myocardial precursors (Fig. 4a) and continues to be expressed in a heart-restricted pattern throughout development (Fig. 4c,e,g). We found that tnnt2 mRNA expression is severely reduced at all stages in sih mutant embryos from both alleles (Fig. 4b,d,f,h). tpma mRNA is expressed at wildtype levels in somites (Fig. 4i–l), but at a reduced level in the heart, at 24 hpf (arrowhead, Fig. 4j).This difference in cardiac expression becomes more exaggerated over time, whereas expression of tpma in somites remains normal (Fig. 4k,l). Considering the role of Tnnt2in contraction and its absence in mutant hearts, we hypothesized that the sih phenotype results primarily from reduction in tnnt2 expression.
a–h, All panels show the sihb109 allele, dorsal views, anterior to the top; sihtc300b mutants have the same phenotype. a,c,e,g, Wildtype tnnt2 expression. b,d,f,h, Reduced expression in sih mutant embryos. At the 15-somite stage (16.5 hpf), tnnt2 expression was not detected above background levels in the mutant lateral plate mesoderm (arrowheads, b versus a). By cone formation (21 somites), some faintly expressing cells were seen around the central lumen (arrowhead, d versus c). During heart tube elongation (24 hpf, f) and looping (30 hpf, h), an increasing number of expressing cells were detected in mutant embryos, but the level of expression in these cells remained severely reduced compared with the wild type (e,g). i–l, Progressive reduction in tpma mRNA expression in sih mutant hearts. At 24 hpf, tpma was detected at a reduced level in sih mutant hearts (arrowhead, j) compared with wildtype hearts (i). This reduction of tpma expression in mutant hearts became even more pronounced over time (30 hpf, l versus k). Note that tpma is expressed heavily in the somites at both stages. Transcript levels of cmlc2, a thick filament component, seem normal in sih mutant hearts (data not shown).
Given the described cellular and molecular phenotypes of sih, we tested whether the sih gene product functions within cardiomyocytes. Using cell transplantation to generate genetic mosaics, we found that when wildtype cells contribute to mutant hearts, they contract spontaneously and express Tnnt2 (see Web Fig. B, and Web Movie B and Web Note B online). Thus, the sih gene product acts in a cell-autonomous manner in regulating cardiomyocyte contractility and Tnnt2 expression. Next, we used morpholino antisense technology to 'knock-down' Tnnt2 translation12. Injection of a tnnt2-specific morpholino oligonucleotide into wildtype eggs resulted in the absence of Tnnt2 protein expression and non-contractile embryonic heart in 98% (230/234) of injected embryos. In addition, immunohistochemistry on morpholino-injected embryos revealed a slight reduction of Tpma and absence of Tnni3 in a pattern identical to that seen in sih mutant embryos (data not shown). Thus, a lack of Tnnt2 phenocopies the sihmutation and results in misexpression of other proteins of the Tpma–Tn complex. These findings indicate that Tnnt2 is essential in sarcomere assembly.
We mapped sih and tnnt2 to the same region of linkage group 23 (Fig. 5a). We assembled a small contiguous stretch of PACs and used fine-scale recombinant mapping to locate the sih locus on PAC 65I4, which also contained tnnt2 (ref. 13). Genotyping embryos for a polymorphic marker within the 5′ upstream sequence of tnnt2 revealed no recombination in 2,095 meiotic events between this marker and the sih locus. Injection of PAC 65I4 DNA into embryos from sih heterozygote intercrosses yielded six mosaically transgenic mutant embryos, where a subset of cells could be seen beating in otherwise non-contractile hearts (see Web Movie C and Web Note C online). The rescued cells also expressed Tnnt2 (Fig. 5b,c). Together, the tight genetic linkage, morpholino phenocopy and rescue experiments provide genetic evidence that sih encodes Tnnt2.
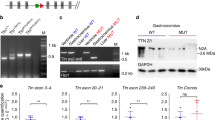
a, Linkage. Genotyping of sihb109 mutant embryos shows recombinants for RFLPs on the telomeric (tnnt2 3′ UTR; 2 in 2,095 meiotic events) and centromeric (PAC-end (65SP6); 2 in 1,872 meiotic events) sides of the sih locus. No recombinants were found for an SSCP in the 5′ upstream sequence of tnnt2. The two recombinants on the telomeric side represent intragenic recombination events. PAC 65I4 (50 kb) contains tnnt2 and spans the sih locus. b,c, PAC rescue of a sihb109 mutant embryo; panels show lateral view, anterior to the left. Rescued cardiac cells (area of detail, b) express Tnnt2 (arrow, c). d,e, Genetic defects in the two sih mutated alleles. d, An A→G change at the −2 position of the splice-acceptor sequence in intron 2 was identified in the sihtc300b allele by genomic sequencing. Consensus splice site in wildtype sequence (ag) is shown. The utilization of a downstream cryptic splice site in sihtc300b mutants (AG) results in a partial exclusion of 7 bp of exon 3 (blue bases) and a frameshift (red) that leads to a premature stop codon in exon 7 (TAA). Only the first 11 aa of Tnnt2 (exon 2) are encoded by this mutant message. e, A deletion of 13 bp from nt −258 to nt −271 of tnnt2 is found in sihb109 mutants (see Web Fig. A online for the complete upstream sequence).
Genomic sequencing of the two sih mutated alleles revealed noncoding mutations in tnnt2. Thesihtc300b mutation results in an A→G change at the invariant −2 position of the splice-acceptor sequence in intron 2 and leads to pleiotropic defects in mRNA splicing. The most common transcript amplified by RT–PCR uses a cryptic splice-site that results in partial exclusion of exon 3 and a frameshift that leads to a premature stop codon in exon 7 (Fig. 5d). This mutation results in severely reduced tnnt2 expression, probably through nonsense-mediated mRNA decay14,15. The sihb109 mutation causes a deletion of 13 bp in the 5′ non-transcribed region of tnnt2 (Fig. 5e). To evaluate the effect of this deletion on tnnt2 expression, we carried out in vivo promoter analyses. An upstream gene fragment (−308) was cloned into a reporter vector encoding green fluorescent protein (GFP) and injected into wildtype embryos. On average, 17% of the injected embryos showed a mosaic pattern of intense, heart-specific GFP fluorescence (Fig. 6b,c). By contrast, no fluorescing cells were seen when a sihb109 mutant promoter construct (−308 Δ13) was injected (Fig. 6c). Moreover, injection of an 11-bp deletion construct (−308 Δ11), which corrects for phase in the DNA helix, did not rescue reporter expression (Fig. 6c). To determine whether this mutation could directly disrupt a critical cis- regulatory element, we examined the sequence of the 13-bp deletion and identified a consensus E-box transcription factor binding site (CANNTG)16. When the first two bases of the E-box were changed from CA to GT and the mutated reporter construct was injected into embryos, 33 (18%) of them expressed GFP in the heart (Fig. 6c).This number is essentially identical to the number obtained when using the wildtype promoter construct, with the notable exception that ectopic GFP expression was seen in 5 of the 33 embryos (that is, 1 or 2 fluorescing cells in the body, bloodstream, and/or head). Thus, although the identified E-box may not be necessary for tnnt2 expression, bases within this region may be relevant to heart-specific expression of tnnt2. From these data, we propose that the deletion of 13 bp causes a severe reduction in tnnt2 transcription in sihb109 mutant embryos; its effect on the binding of critical transcriptional complexes warrants future investigation.
a,b, Lateral view of head and heart (arrow, a) in a transient transgenic zebrafish embryo. Five GFP-expressing cells are seen (arrow, b). c, Data table shows the number of injected embryos evaluated for GFP expression using the wildtype (−308), sihb109 (−308 Δ13), 11-bp deletion (−308 Δ11) and E-box mutation (CA→GT) constructs. No GFP expression is observed when mutant deletion constructs are injected. GFP expression occurs at wildtype levels when the E-box mutation construct is injected. However, GFP expression was not heart-specific in 5 of 33 expressing embryos using this construct. Results are a summation of independent injection experiments.
The mutations reported here provide the first animal model of Tnnt2 deficiency. Without Tnnt2, cardiac sarcomeres fail to assemble and heart muscle is rendered nonfunctional. Reduction of Tnnt2 is accompanied by reductions in Tpma and Tnni3. A reduction in tpma mRNA in sih mutant hearts indicates the existence of a feedback mechanism to tightly coordinate the expression of these interrelated proteins.In humans, hypertrophic cardiomyopathy resulting from dominant mutations in TNNT2 can cause sudden death without evidence of clinical hypertrophy17. The precise molecular mechanisms leading to sudden death are unknown.A recent histopathological study of nine hearts from individuals with TNNT2 mutations reported lower heart weights, greater myocyte disarray and less fibrosis than hearts from individuals with hypertrophic cardiomyopathy of unknown genotype6. There may thus be a link between TNNT2 mutations, myocyte disarray and ischemia as risk factors for sudden cardiac death.
In transgenic mice, a human splice-site mutation that results in a carboxy-terminal Tnnt2 truncation has been modeled18. Only mice expressing less than 5% of total Tnnt2 as the truncated form survive beyond 24 hours; these mice have fewer and smaller cardiomyocytes and decreased heart mass. The truncated Tnnt2 molecule shows altered stability within the sarcomere and causes misregistration of Z-bands and myofibrillar disarray and degeneration7. Mechanical dysfunction due to incorporation of the abnormal protein was proposed as the mechanism leading to structural breakdown of sarcomeres and possibly increased apoptosis. Alternatively, based on the loss-of-function data presented here, we propose that if a mutation in TNNT2 results in the reduced expression of other proteins of the Tpma–Tn complex, then defective sarcomeres would be directly eliminated and lead to myocyte disarray. Identifying the multilevel controls that regulate contractile protein expression is key to understanding cardiomyocyte function and dysfunction.
Methods
Zebrafish.
We maintained and staged zebrafish as described19. The sihb109 mutation was identified in a γ-ray mutagenesis screen in the laboratory of Charles Kimmel (Eugene, Oregon).
Calcium activation.
We dissected hearts from wildtype and sihb109 embryos at 32 hpf and loaded them with Ca2+ green (Molecular Probes) according to the manufacturers instructions. We then viewed and recorded the hearts using a Zeiss Axiophot microscope.
Electron microscopy.
We fixed wildtype and homozygous mutant sihb109 embryos at 48 hpf in 2% glutaraldehyde in 0.1M sodium cacodylate buffer (pH 7.2). Post-fixation was carried out in 0.5% OsO4 plus 0.8% K3Fe(CN)6. We then placed embryos in 2% uranyl acetate for 1 h in the dark and dehydrated them in a graded acetone solution. We used Epon-Araldite resin for embedding and cured the resin for 48 h at 60 °C. We cut thin sections (70 nm), post-stained them with uranyl acetate and lead citrate and observed them under a JEOL 100CX electron microscope.
Immunohistochemistry.
We carried out whole-mount immunohistochemistry as described20. We used monoclonal antibodies against myosin heavy chain, MF20 (ref. 21), atrial-specific myosin heavy chain, S46 (gift from F. Stockdale, Stanford Univ.); α-actinin (Sigma); tropomyosin, CH1 (ref. 22), cardiac troponin T, JLT12 (Sigma); cardiac troponin I, IE7 (gift from J. Potter, Univ. of Miami); and α-sarcomeric actin (Sigma). Anti-IgG2b TRITC recognized MF20. FITC conjugated anti-mouse antibodies recognized S46, α-actinin, CH1 and JLT12 (IgG1-FITC), IE7 (IgG-FITC) and α-sarcomeric actin (IgM-FITC).
Isolation of tnnt2 cDNA.
We used a heart-specific monoclonal antibody against Tnnt2, mAb13-11 (ref. 23, gift from P. Anderson, Duke Univ.)to screen 5×105 recombinants from a zebrafish heart cDNA library in Stratagene Uni-ZAP XR (gift from R. Breitbart). We applied nitrocellulose filters impregnated with isopropyl-b-D-thiogalactopyranoside to plaques for 4 h at 37 °C. We blocked filters (3% nonfat dry milk in Tris buffered saline with 0.1% tween) and then incubated them in mAb13-11 (1:2,000). After washes we incubated filters with a goat anti-mouse HRP conjugated secondary antibody (1:5,000). We carried out electro-chemi-luminescence detection with ECL reagents (Amersham). Positive pBluescript phagemids were in vivo excised and sequenced. Sequencing of three clones revealed identical, full-length cDNA sequences that have significant homology to other vertebrate tnnt2 genes (see Web Fig. A online).
Morpholino antisense 'knock-down'.
We designed a morpholino oligonucleotide (Gene Tools) to bind the Tnnt2 translation start codon and flanking 5′ sequence (5′-CATGTTTGCTCTGATCTGACACGCA-3′). We injected 4 ng of the morpholino oligo into embryos at the 1–4 cell stage and examined the cardiac phenotype of the injected embryos at 30 hpf.
Genetic mapping.
We genotyped a combination of diploid mutant embryos and haploid mutant and wildtype embryos from sihb109 AB/WIK hybrid strains on the telomeric side of the sih locus using a restriction fragment length polymorphism (RFLP) in the 3′ UTR of tnnt2. We isolated PAC 65I4 by PCR from PAC library pools using tnnt2-specific primers and sized it on a pulse-field gel13. We directly sequenced the PAC ends. We used an RFLP in one end of the PAC (65SP6) to genotype on the centromeric side of the sih locus. We then obtained 5′ upstream sequence of tnnt2 by direct PAC sequencing and used a single-strand conformation polymorphism (SSCP) within this sequence for final genotyping of all recombinant embryos.
Mutation detection.
We used pools of 100 homozygous mutant embryos from both alleles (sihb109 and sihtc300b) to extract mRNA (Trizol, Gibco BRL) and synthesize cDNA (SuperScript II Reverse Transcription, Gibco BRL). We amplified and cloned two overlapping fragments of tnnt2 coding region from the cDNA pools. Sequencing of clones from the sihtc300b allele revealed one abnormally spliced transcript, which contained sequence for intron 2, and three independent clones of a second splice form, which partially excluded exon 3. The splice form that contained intron 2 had a G at the −2 position of the splice-acceptor sequence (consensus=A). We confirmed an A→G change at this position by sequencing two independent clones amplified from mutant genomic DNA (DNA). Coding sequence analysis of the sihb109 allele revealed no mutations. We therefore amplified and cloned a 327-bp fragment of tnnt2 5′ upstream sequence (containing the SSCP described above) from genomic DNA prepared from individual wildtype and sihb109 haploid embryos. We identified a deletion of 13 bp from nt −258 to nt −271 upstream of the transcription start site, which we confirmed in three independent clones from sihb109 embryos.
Phenotypic rescue.
We isolated PAC DNA using a Maxi-Kit (Qiagen) and injected it at concentrations ranging from 50 μg ml−1 to 190 μg ml−1 into embryos from sihb109 and sihtc300b heterozygote intercrosses. At 30 hpf, we sorted mutant embryos and observed them for evidence of beating cells in the heart. We then prepared injected embryos for whole-mount immunohistochemistry using antibody mAB13-11 (ref. 23). We genotyped rescued embryos from the sihb109 allele using an allele-specific RFLP that lies outside of the PAC.
Promoter analysis.
We amplified a 327-bp fragment from nt −308 to nt +19 of tnnt2 (see Web Fig. A online) from homozygous wildtype WIK and sihb109 genomic DNA and cloned it into the pEGFP-1 promoterless reporter vector (Clontech). We used the QuikChange Site-Directed Mutagenesis Kit (Stratagene) to generate two additional constructs using the mutant (−308 Δ13) and wildtype (−308) constructs as templates, respectively. To create an 11-bp deletion (–308 Δ11), we re-inserted two bases (GG) into the (−308 Δ13) construct. To alter the E-box consensus site (CANNTG) contained within the 13-bp sequence, we changed two bases within the wildtype promoter (CA to GT).We confirmed all constructs by sequencing. We diluted plasmid DNA to a working concentration of 100 μg ml−1 for microinjection. We scored injected embryos for GFP expression at 30–48 hpf.
GenBank accession numbers:
Zebrafish tnnt2, AF282384; PAC 65I4, AL662878.
Note: Supplementary information is available on the Nature Genetics website.
References
Thierfelder, L. et al. α-tropomyosin and cardiac troponin T mutations cause familial hypertrophic cardiomyopathy: a disease of the sarcomere. Cell 77, 701–712 (1994).
Maron, B.J. et al. Sudden death in young competitive athletes. Clinical, demographic, and pathological profiles. JAMA 276, 199–204 (1996).
Seidman, J.G. & Seidman, C. The genetic basis for cardiomyopathy: from mutation identification to mechanistic paradigms. Cell 104, 557–567 (2001).
Kamisago, M. et al. Mutations in sarcomere protein genes as a cause of dilated cardiomyopathy. N. Engl. J. Med. 343, 1688–1696 (2000).
Tobacman, L.S. Thin filament-mediated regulation of cardiac contraction. Annu. Rev. Physiol. 58, 447–481 (1996).
Varnava, A.M. et al. Hypertrophic cardiomyopathy: histopathological features of sudden death in cardiac troponin T disease. Circulation 104, 1380–1384 (2001).
Tardiff, J.C. et al. Cardiac troponin T mutations result in allele-specific phenotypes in a mouse model for hypertrophic cardiomyopathy. J. Clin. Invest. 104, 469–481 (1999).
Chen, J.N. et al. Mutations affecting the cardiovascular system and other internal organs in zebrafish. Development 123, 293–302 (1996).
Stainier, D.Y. et al. Mutations affecting the formation and function of the cardiovascular system in the zebrafish embryo. Development 123, 285–292 (1996).
Burggren, W.W. & Pinder, A.W. Ontogeny of cardiovascular and respiratory physiology in lower vertebrates. Annu. Rev. Physiol. 53, 107–135 (1991).
Fyrberg, E., Fyrberg, C.C., Beall, C. & Saville, D.L. Drosophila melanogaster troponin-T mutations engender three distinct syndromes of myofibrillar abnormalities. J. Mol. Biol. 216, 657–675 (1990).
Nasevicius, A. & Ekker, S.C. Effective targeted gene 'knockdown' in zebrafish. Nature Genet. 26, 216–220 (2000).
Amemiya, C.T., Zhong, T.P., Silverman, G.A., Fishman, M.C. & Zon, L.I. Zebrafish YAC, BAC, and PAC genomic libraries. Methods Cell Biol. 60, 235–258 (1999).
O'Neill, J.P., Rogan, P.K., Cariello, N. & Nicklas, J.A. Mutations that alter RNA splicing of the human HPRT gene: a review of the spectrum. Mutat. Res. 411, 179–214 (1998).
Maquat, L.E. & Carmichael, G.G. Quality control of mRNA function. Cell 104, 173–176 (2001).
Edmondson, D.G. & Olson, E.N. Helix-loop-helix proteins as regulators of muscle-specific transcription. J. Biol. Chem. 268, 755–758 (1993).
Moolman, J.C. et al. Sudden death due to troponin T mutations. J. Am. Coll. Cardiol. 29, 549–555 (1997).
Tardiff, J.C. et al. A truncated cardiac troponin T molecule in transgenic mice suggests multiple cellular mechanisms for familial hypertrophic cardiomyopathy. J. Clin. Invest. 101, 2800–2811 (1998).
Westerfield, M. The Zebrafish Book (Univ. of Oregon Press, Eugene, 1995).
Alexander, J., Stainier, D.Y. & Yelon, D. Screening mosaic F1 females for mutations affecting zebrafish heart induction and patterning. Dev. Genet. 22, 288–299 (1998).
Bader, D., Masaki, T. & Fischman, D.A. Immunochemical analysis of myosin heavy chain during avian myogenesis in vivo and in vitro. J. Cell. Biol. 95, 763–770 (1982).
Lin, J.J., Chou, C.S. & Lin, J.L. Monoclonal antibodies against chicken tropomyosin isoforms: production, characterization, and application. Hybridoma 4, 223–242 (1985).
Malouf, N.N., McMahon, D., Oakeley, A.E. & Anderson, P.A. A cardiac troponin T epitope conserved across phyla. J. Biol. Chem. 267, 9269–9274 (1992).
Yelon, D., Horne, S.A. & Stainier, D.Y. Restricted expression of cardiac myosin genes reveals regulated aspects of heart tube assembly in zebrafish. Dev. Biol. 214, 23–37 (1999).
Ohara, O., Dorit, R.L. & Gilbert, W. One-sided polymerase chain reaction: the amplification of cDNA. Proc. Natl Acad. Sci. USA 86, 5673–5677 (1989).
Acknowledgements
We thank A. Navarro and S. Waldron for their dedicated care of our fish, K. MacDonald for his expertise in generating electron micrographs, and M. Brook and F. Aburto for their assistance with the Quicktime movies. We thank C. Kimmel, F. Stockdale, J. Potter, P. Anderson and R. Breitbart for sharing valuable reagents; W. Tidyman, C. Ordahl and B. Black for helpful discussions about cardiac gene regulation and P. Wolters, M. Zeiger, R. Reijo-Pera and members of the Stainier lab for their comments on this manuscript. A.S. was supported as a March of Dimes fellow of the Pediatric Scientist Development Program and by the American Heart Association Western States Affiliate. This work was supported in part by grants from the National Institutes of Health (to A.S., M.C.F. and D.Y.R.S., and to Charles Kimmel supporting the mutagenesis screen), as well as grants from the Packard Foundation and the American Heart Association (to D.Y.R.S.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Sehnert, A., Huq, A., Weinstein, B. et al. Cardiac troponin T is essential in sarcomere assembly and cardiac contractility. Nat Genet 31, 106–110 (2002). https://doi.org/10.1038/ng875
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng875
This article is cited by
-
Glia maturation factor-γ is required for initiation and maintenance of hematopoietic stem and progenitor cells
Stem Cell Research & Therapy (2023)
-
A bioelectrical phase transition patterns the first vertebrate heartbeats
Nature (2023)
-
Multiple pkd and piezo gene family members are required for atrioventricular valve formation
Nature Communications (2023)
-
Role of Biomarkers in the Management of Immune-Checkpoint Inhibitor-Related Myocarditis
Current Cardiology Reports (2023)
-
Transcriptome studies of inherited dilated cardiomyopathies
Mammalian Genome (2023)