Abstract
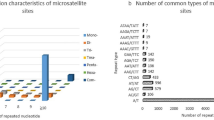
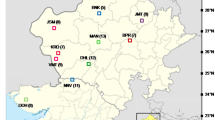
Antheraea mylitta Drury, the semi-wild silk-producing lepidopteran insect commonly known as tasar silkworm is unique to India and is distributed over a wide tropical forest range covering the states of Andhra Pradesh, Bihar, Chhattisgarh, Madhya Pradesh, Maharashtra, Orissa, and Uttaranchal. The populations found in different areas are know by their specific local names and are considered as different ecotypes, but it is difficult to separate the populations on the basis of morphological and life-cycle traits and thus molecular characterization was attempted. The present communication relates to the results obtained from the analysis of polymorphism unraveled by twelve ISSR primers for 11 populations of A. mylitta belonging to six ecotypes and 41 individuals of Railey ecotype collected from five zones of Dandakarnya forest in Madhya Pradesh. This communication, further, presents molecular evidences on genetic differences between eleven ecotype populations and highlights the genotypic diversification of a single ecotype into further separate discrete gene pools. The canonical discriminant function analysis revealed grouping of the five populations of Railey ecotype into two “clumps,” while accessions of other ecotypes stood separated from each other. The Railey populations on detailed study, further, revealed separation of two (Tokapal and Nangur) populations into discrete gene pools and the other three (Kondagaon, Darba, and Tongpal) populations, overlapped in spite of larger geographic distance between them. The analysis also identified nine markers, which can be utilized to characterize specific population and will be of help to follow the ongoing genetic changes triggered by various ecological factors and human influences on the Railey ecotype.
Access this article
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.
Similar content being viewed by others
REFERENCES
Wardle, T., in The Economic Product of India, 1881, vol. 6.
Mukherjee, S., Handbook of Sericulture, Bengal Secretariat Book Depot Calcutta, India, 1898.
Ghosh, C.C., Silk Production and Weaving in India, New Delhi: CSIR, 1949.
Jolly, M.S., Sen, S.K., and Ashan, M.M., Tasar Culture, Bombay: Ambika, 1974, pp. 1-166.
Anonymous Annual Report, Ranchi, India: Central Tasar Res. Training Inst., 1972.
Gaur, J.P., Races of Antheraea mylitta, the Tropical Tasar Silkworm, Their Distribution and Variability, Ind. J. Entomol., 1992, vol. 54. pp. 275-284.
Akai, H., Nagashima, T., and Aoyagi, S., Ultrastructure of Posterior Silk Gland Cells and Liquid Silk in Indian Tasar Silkworm, Antheraea mylitta Drury (Lepidoptera; Saturniidae), Int. J. Insect Morphol. Embryol., 1993, vol. 22, pp. 497-506.
Siddique, A.A., Chatterjee, S.N., Goel, A.K., and Sengupta, A.K., Genetic Divergence in Tasar Silkworm A. mylitta D., Seircologia, 1992, vol. 32, pp. 425-431.
Thnagavelu, K. and Sinha, A.K., Population Ecology of Antheraea mylitta Drury (Lepidoptera: Saturniidae), in Wild Silkmoth, Akai, H., Kato, Y., Kiuchi, M., and Inouch, J., Eds., Tsukuba, Japan: Int. Soc. Wild Silkmoths, Nat. Inst. Sericult. Entomol. Sci., 1992, pp. 87-92.
Alam, M.O., Suresh, M.R., and Sit, S.C., Survey, Collection and Characterization of Modal Ecorace of Indian Wild Silk Insect Antheraea mylitta D. (Lepidoptera, Saturniidae), in Wild Silkmoth, 1993, pp. 93-98.
Fang, D.Q. and Roose, M.L., Identification of Closely Related Citrus Cultivars with Inter-Simple Sequence Repeat Markers, Theor. Appl. Genet., 1997, vol. 95, pp. 408-417.
Tsumura, Y., Ohba, K., and Strauss, S.H., Diversity and Inheritance of Inter-Simple Sequence Repeat Polymorphisms in Douglas Fir (Pseudotsuga menziesii) and Sugi (Cryptomeria japonica), Theor. Appl. Genet., 1996, vol. 92, pp. 40-45.
Nagaoko, T. and Ogihara, Y., Applicability of Inter-Simple Sequence Repeat Polymorphisms in Wheat for Use as DNA Markers Comparison to RFLP and RAPD Markers, Theor. Appl. Genet., 1997, vol. 94, pp. 597-602.
Raina, S.N., Rani, V., Kojima, T., et al., RAPD and ISSR Fingerprints as Useful Genetic Markers for Analysis of Genetic Diversity, Varietal Identification, and Phylogenetic Relationship in Peanut (Arachis hypogaea) Cultivars and Wild Species, Genome, 2001, vol. 44, pp. 763-772.
Reddy, D.K., Nagaraju, J., and Abraham, E.G., Genetic Characterization of the Silkworm Bombyx mori by Simple Sequence Repeat (SSR)-Anchored PCR, Heredity, 1999, vol. 83, pp. 681-687.
Ramkumar Rao, K.V.S. and Mohobia, G.P., Studies on Grainage Behavior of Nature-Grown Raily Ecorace under Natural and Captive Conditions, Ind. J. Sericult., 1992, vol. 31, pp. 165-166.
Singh, B.M.K. and Srivastava, A.K., Ecoraces of A. mylitta D. and Exploitation Strategy through Hybridization, in Current Technology Seminar on Non-Mulberry Sericulture, Ranchi: Central Tasar Res. Training Inst., 1997, base paper 6:39.
Morrison, D.F., Multivariate Statistical Methods, New York: Wiley, 1976.
Sinha, A.K. and Naqni, A.H., Need for Maintenance of Germplasm of Non-Mulberry Silkworm, in Current Technology Seminar on Non-Mulberry Sericulture, Ranchi: Central Tasar Res. Training Inst., 1997, base paper 7:16.
Kantety, R.V., Zhang, X., Bennetzen, J.L., and Zehr, B.Z., Assessment of Genetic Diversity in Dent and Popcorn (Zea mays L.) Inbred Lines Using Inter-Simple Sequence Repeat (ISSR) Amplification, Mol. Breed., 1995, vol. 1, pp. 365-373.
Provost, A. and Wilkinson, M.J., A New System of Comparing PCR Primers Applied to ISSR Fingerprinting of Potato Cultivars, Theor. Appl. Genet., 1997, vol. 98, pp. 107-112.
Chatterjee, S.N., Mohandas, T.P., and Tanushree, T., Eur. J. Entomol., 2003, vol. 1.
Akagi, H., Yokozeki, Y., Inagaki, A., et al., A Codominant DNA Marker Closely Linked to the Rice Nuclear Gene, Rf-1, Identified with Inter-SSR Fingerprinting, Genome, 1996, vol. 39, pp. 1205-1209.
Zietkiewics, E., Rafalski, A., and Labuda, D., Genome Fingerprinting by Simple Sequence Repeats (SSR)-Anchored Amplification, Genomics, 1994, vol. 20, pp. 176-183.
Davids, R., Buddhist India, London, 1903.
Breto, M.P., Ruiz, C., Pina, J.A., and Asins, M.J., Diversification of Citrus clementina Hort. Ex Tan., a Vegetatively Propagated Crop Species, Mol. Phylogenet. Evol., 2001, vol. 21, pp. 285-293.
Culley, T.M. and Wolfe, A.D., Population Genetic Structure of the Cleistogamous Plant Species Viola pubescens Aiton (Violaceae), as Indicated by Allozyme and ISSR Molecular Markers, Heridity, 2001, vol. 86, pp. 545-556.
Rao, K.V.S., Tasar Crop Planning in Bastar Plateau of Chhattisgarh, Ind. Silk, 2001, vol. 40, pp. 11-13.
Breto, M.P., Ruiz, C., Pina, J.A., and Asins, M.J., Mol. Phylogenet. Evol., 2001, vol. 21, pp. 285-293.
Charlesworth, D. and Wright, S., Breeding System and Genome Evolution, Curr. Opin. Genet. Dev., 2001, vol. 11L, pp. 685-690.
Rao, K.V.S., Ramkumar, Mahobia, G.P., et al., Studies on in Situ Conservation of Raily Ecorace of Indian Wild Tasar Insect A. mylitta D. (Lepidoptera, Saturniidae), Int. J. Wild Silkmoth Silk, 2000, vol. 5, pp. 314-317.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Chatterjee, S.N., Vijayan, K., Roy, G.C. et al. ISSR Profiling of Genetic Variability in the Ecotypes of Antheraea mylitta Drury, the Tropical Tasar Silkworm. Russian Journal of Genetics 40, 152–159 (2004). https://doi.org/10.1023/B:RUGE.0000016988.08342.c0
Issue Date:
DOI: https://doi.org/10.1023/B:RUGE.0000016988.08342.c0