-
PDF
- Split View
-
Views
-
Cite
Cite
Sabyasachi Sanyal, Jason Matthews, Didier Bouton, Han-Jong Kim, Hueng-Sik Choi, Eckardt Treuter, Jan-Åke Gustafsson, Deoxyribonucleic Acid Response Element-Dependent Regulation of Transcription by Orphan Nuclear Receptor Estrogen Receptor-Related Receptor γ, Molecular Endocrinology, Volume 18, Issue 2, 1 February 2004, Pages 312–325, https://doi.org/10.1210/me.2003-0165
Close - Share Icon Share
Abstract
The estrogen receptor-related receptor γ (ERRγ/ERR3/NR3B3) is the newest member of the ERR subfamily that also includes ERRα and ERRβ. All three isoforms share a high degree of amino acid identity especially in the DNA binding domain. ERRγ is a constitutively active transcriptional activator that regulates reporter elements driven by steroidogenic factor 1 response element (SF-1RE) and estrogen response element. However, it has the highest potency on a derivative of SF-1RE present in the small heterodimer partner gene promoter called sft4 and unlike ERRα and -β, it fails to activate a palindromic thyroid hormone response element. To investigate the mechanism behind this response element-specific differential transcriptional activity of ERRγ, the interactions of ERRγ and the aforementioned response elements was monitored. EMSA and chromatin immunoprecipitation assays demonstrated that ERRγ binds to sft4, SF-1RE, and palindromic thyroid hormone response element albeit with different degrees of affinity, but causes hyperacetylation of sft4 and SF-1RE templates only. Limited proteolysis assays showed that ERRγ, bound to different elements, shows differential trypsin sensitivity. A search for novel coregulators of ERRγ led to the identification of receptor interacting protein 140 as a potent corepressor and peroxisome proliferator-activated receptor γ coactivator 1 as a potent coactivator of ERRγ. DNA-dependent pull-down and transient transfection assays demonstrated that, on different DNA elements, ERRγ exhibits differential cofactor interactions, which in turn dictate its transcriptional activity. Because ERRγ shows a similar tissue distribution as peroxisome proliferator-activated receptor γ coactivator 1 and receptor interacting protein 140, these two coregulators may act as key components of ERRγ-mediated transcription.
THE ESTROGEN RECEPTOR (ER)-RELATED RECEPTOR (ERR) subfamily includes the first orphan nuclear receptors described. These proteins form a distinct subgroup in the nuclear receptor superfamily with striking sequence similarity in the DNA binding domain with ERs. The sequence similarity in the ligand binding domain (LBD), however, is more limited and the ERRs do not bind 17β-estradiol (1). Several recent reports have shown that synthetic estrogenic compounds such as diethylstilbestrol and 4-hydroxytamoxifen (OHT) act as inverse agonists for the ERR family members by disrupting ERR-coactivator interactions (2–4). All three ERR isoforms, designated as ERRα, -β, and -γ, share remarkably high degree of amino acid identity, which is more evident between ERRβ and -γ. ERRα and -γ share a similar tissue distribution with high expression in heart, brain, kidney, and skeletal muscle (5–8). In addition, ERRγ is highly expressed in pancreas and stomach (8). Expression of ERRβ is barely detectable in adult tissues; in mouse it is expressed in a subset of extraembryonic tissues in a small window between 5.5 d postcoitum and 8.5 d postcoitum (9, 10). The status of ERRα and -β as true orphan nuclear receptors has been much debated. Vanacker et al. (11) have shown that the presence of an unknown serum component, which can be eliminated by charcoal treatment, is essential for the activities of ERRα and -β; however, other reports have shown that both ERRα and -β can bind coactivators in the absence of serum (12, 13). The status of ERRγ as a true orphan is more obvious. A recently reported structural analysis of ERRγ LBD complexed with steroid receptor coactivator 1 receptor-interacting domain has clearly shown that ERRγ LBD acquires the typical transcriptionally active conformation of agonist-bound nuclear receptor in the absence of any ligand (14). In addition, other functional evidence also suggests that ERRγ binds to coactivators constitutively, in the absence of any ligand (2–4, 6, 8).
Peroxisomal proliferator-activated receptor-γ (PPAR)γ coactivator 1 (PGC-1) is a tissue-specific coactivator that was first identified as a PPARγ-interacting protein (15); unlike the P160 and P300 family of coactivators, the expression of PGC-1 is limited to tissues such as heart, brain, skeletal muscle, kidney, and pancreas (16). PGC-1 plays a number of critical roles in regulating multiple aspects of energy metabolism including mitochondrial biogenesis, thermogenesis, gluconeogenesis, and fatty acid metabolism (15–20). In contrast, RIP140 is a unique corepressor that binds to the active conformation of nuclear receptors in the same fashion as coactivators and represses their activity by either 1) competition with coactivator binding to the activation function 2 interface of nuclear receptors (21–25), 2) by recruiting additional repressive molecules such as C-terminal binding protein (26), or 3) by recruitment of histone deacetylases (27, 28) or a combination of these mechanisms. Interestingly, receptor-interacting protein 140 (RIP140) has a similar tissue distribution to PGC-1, with high expression in brain, heart, stomach, and kidney (21). The fact that both these cofactors interact with nuclear receptors in their active state and that they have a similar tissue distribution indicates that these two proteins may act as a yin-yang control for nuclear receptor-mediated transcription depending on relative levels of their expression at a given time point.
In our previous report we have demonstrated that ERRγ is a constitutive activator of the small heterodimer partner (SHP) gene promoter, and this activation is achieved through its binding to one of the previously described steroidogenic factor 1 response element (SF-1RE) derivatives known as sft4 (8). Although ERRγ shows a strong binding to consensus SF-1RE/ERR response element (7), it only moderately activates a reporter construct containing such binding sites compared with sft4-driven reporter gene (7, 8). On the other hand, recent reports have demonstrated that ERRα and -β constitutively activate reporters driven by estrogen response element (ERE), palindromic thyroid hormone response element (TRE-pal), and SF-1RE (11–13). ERRγ was shown to be inactive on a TRE-pal and moderately active on a consensus SF-1RE (7, 8) although it strongly activates an ERE-luc reporter (6). Collectively, these results show that even though the DNA-binding domain of the ERR family members is highly conserved, with ERRβ and -γ sharing 98.9% sequence identity between them, and also that ERRγ LBD contains the strongest activation potential among the ERRs (7), ERRγ shows moderate or no activity on elements that are activated by ERRα and -β.
This study was designed to elucidate the mechanisms involved in the differential transcriptional activities of ERRγ. We demonstrate that, depending on the sequence of its target element, ERRγ exhibits differential affinity for PGC-1 and RIP140, which in turn may alter its transcriptional activity. Considering the remarkable similarity in the expression profiles of ERRγ, PGC-1, and RIP140, this report provides new insights into the regulation of the transcriptional activity of ERRγ by coactivators and corepressors in the absence of ligands.
RESULTS
Response Element-Specific Transactivation Potential of ERRγ
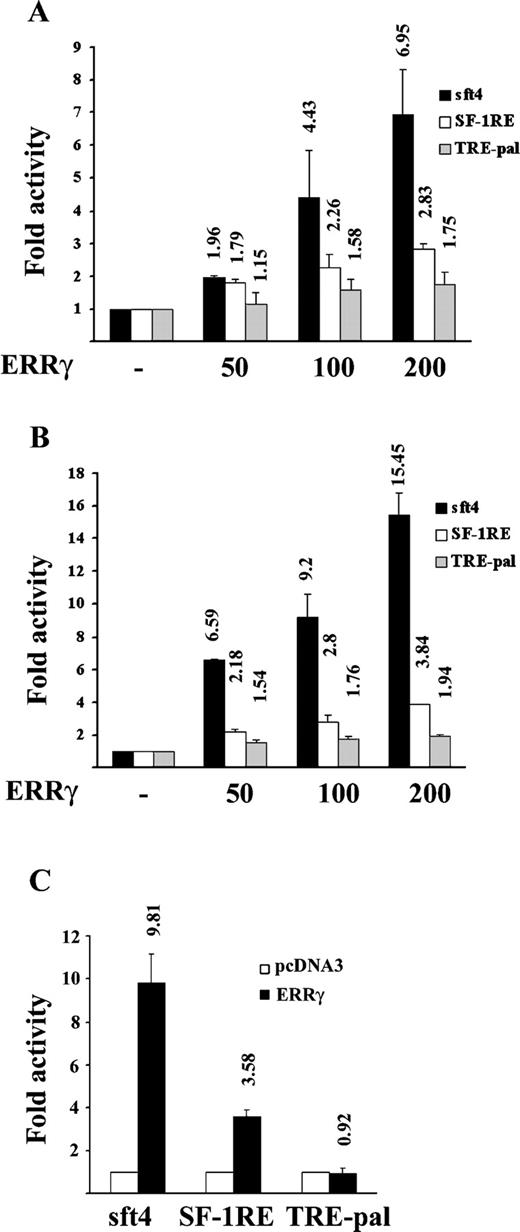
Transient transfection assays were performed to assess ERRγ transcriptional activity on luciferase reporters containing the previously described ERR response elements (6–8, 11, 12). Figure 1 shows a comparative account of ERRγ activity on different response elements including sft4, consensus SF-1RE, and TRE-pal. In CV-1 cells, ERRγ strongly activated a one copy (1×) sft4 reporter (6.95-fold) and moderately activated a 1× SF-1RE (2.83-fold). However, it caused a marginal activation of the 1× TRE-pal (1.75-fold) (Fig. 1A). A previous report demonstrated that the activity of ERRα depends on the copy number of the response elements, where ERRα did not activate a 1× SF-1RE element but strongly activated a 3× SF-1RE (11). We tried to determine whether the relative inactivity of ERRγ on a 1× TRE-pal could be overcome on a 3× response element. As demonstrated in Fig. 1B, on 3× TRE-pal ERRγ exhibited no substantial increase in activity in comparison to its activity on 1× TRE-pal (1.75-fold vs. 1.94-fold). However, a strong activation was observed on 3× sft4 (6.95-fold vs. 15.45-fold), and a moderate increase was also observed on 3× SF-1RE (2.83-fold vs. 3.84-fold). To determine whether this response element-specific activity of ERRγ was cell type specific, the same experiments were repeated in HeLa cells; as expected, ERRγ strongly activated 3× sft4 (9.81-fold) and moderately activated 3× SF-1RE (3.58-fold), but it did not affect the 3× TRE-pal activity (0.92-fold); however, the activity of ERRγ was lower than that in CV-1 cells (compare Fig. 1, B and C). Cos-7 cells also showed similar pattern of activities as in the case of CV-1, albeit with a slightly lower level of activity (data not shown).
Differential Activation of sft4, SF-1RE, and TRE-pal-Containing Reporter Genes by ERRγ A, CV-1 cells were transfected with 200 ng of the indicated 1× luciferase reporter plasmids, 200 ng of RSV-β-galactosidase, and indicated doses of mammalian expression plasmids expressing ERRγ or empty vector, pcDNA3. B, CV-1 cells were transfected with 200 ng of the indicated 3× luciferase reporter plasmids, 200 ng of RSV-β-galactosidase, and indicated doses of ERRγ or pcDNA3 vector. C, HeLa cells were transfected with 200 ng of the indicated 3× reporter plasmids, 200 ng of RSV-β-galactosidase, and 200 ng of ERRγ or pcDNA3 vector. After transfections, cells were lysed, and the luciferase activity was measured, normalized against β-galactosidase activity, and plotted as fold activity, with 1-fold basal activity defined as the luciferase activity with pcDNA3. The results are expressed as mean ± se from three independent experiments performed in duplicate. The values on the top of each bar represent the respective mean fold activity.
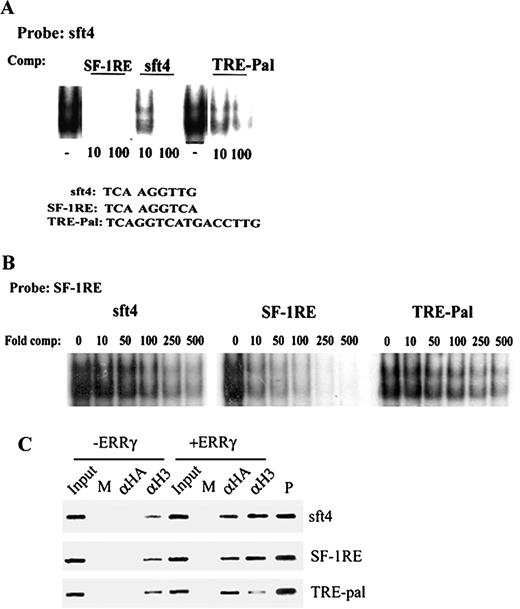
To investigate whether the relative inactivity of ERRγ on TRE-pal and lower activity on SF-1RE was due to a difference in its binding to these elements, a series of EMSAs were performed. Radiolabeled sft4-bound ERRγ was competed with 10- or 100-fold molar excess of unlabeled sft4, consensus SF-1RE, and TRE-pal. The strongest binding was observed on the consensus SF-1RE where 10-fold molar excess of the cold competitor was sufficient to successfully compete with the ERRγ-sft4 complex, whereas, ERRγ demonstrated approximately 10-fold weaker affinity for sft4 and TRE-pal. To further compare the affinities of ERRγ for sft4, SF-1RE, and TRE-Pal, an ERRγ-SF-1RE complex was competed with cold competitor oligonucleotides corresponding to sft4, SF-1RE, and TRE-pal. As demonstrated in Fig. 2B, and consistent with Fig. 2A, ERRγ showed the strongest affinity for SF-1RE, whereas it showed much weaker affinity for both sft4 and TRE-pal. Collectively, Figs. 1 and 2 suggest that DNA binding affinity is not a determinant for the transcriptional activity of ERRγ.
ERRγ Binds but Fails to Activate a TRE-pal A, Purified GST-ERRγ was incubated with end-labeled sft4 oligonucleotide, and the sft4-ERRγ complexes were competed with 10- or 100-fold molar excess of oligonucleotides corresponding to sft4, SF-1RE, or TRE-pal. B, Purified GST-ERRγ was incubated with end-labeled SF-1RE and competed with 0, 10-, 50-, 100-, 250-, or 500-fold molar excess of unlabeled sft4, SF-1RE, or TRE-pal. The resulting complexes were resolved by 5% nondenaturing PAGE and visualized by autoradiography. The competition assays were also repeated with in vitro transcribed and translated ERRγ with similar results. C, Cos-7 cells were transfected with sft4, SF-1RE, or TRE-pal with or without HA ERRγ as indicated in the figure. After cross-linking with 1% formaldehyde, soluble chromatin was prepared from the cells and incubated with preimmune serum (M), anti-HA (αHA), or antiacetylated H3 (αH3) antibodies. After immunoprecipitation the chromatin templates were purified and PCR amplified using primers flanking sft4, SF-1RE, or TRE-pal. Input indicates 5% of the total chromatin used for each immunoprecipitation, and plasmid control is indicated. Representative figures from three independent experiments are shown, and the data were highly reproducible.
Chromatin immunoprecipitation (ChIP) assays were performed to investigate whether ERRγ binds sft4, SF-1RE, and TRE-pal in vivo and differentially regulates the level of histone acetylation, which is a main determinant of the transcriptional status. Cos-7 cells were transfected with sft4, SF-1RE, and TRE-pal reporters with or without hemagglutinin (HA)-tagged ERRγ. After 48 h incubation, the cell extracts were immunoprecipitated with preimmune serum (mock), anti-HA, or antiacetylated histone histone 3 (H3) antibodies, and the presence of the target sequences in the chromatin immunoprecipitates was analyzed by PCR. As demonstrated in Fig. 2C, hemagglutinin (HA)-ERRγ was recruited to all the three ERR response elements (lane 7), whereas no chromatin was immunoprecipitated with anti-HA antibody in cells transfected with the reporters only (lane 3). A small level of acetylation was observed on chromatinated sft4, SF-1RE, and TRE-pal, which perhaps indicates basal transcription (lane 4). However, cotransfection of ERRγ substantially increased the acetylation of sft4 and SF-1RE, but not of a TRE-pal (compare lanes 4 and 8), indicating that although ERRγ is recruited to these elements in vivo, it only activates transcription of sft4, and SF-1RE-driven reporters, but not of a TRE-pal-driven one.
DNA-Induced Differences in the Protease Sensitivity of ERRγ
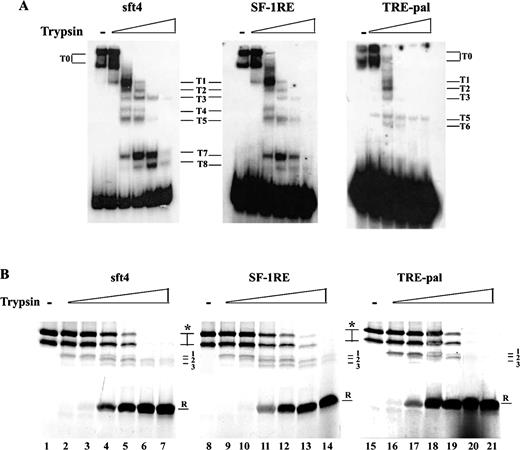
Because the DNA binding affinity of ERRγ is not the determinant for its activity, we assessed whether the DNA sequences could mediate structural alterations of the ERRγ protein itself and whether such structural changes, in turn, may be a cause for its differential activity. To study this, a limited proteolysis assay was performed, in which radiolabeled sft4, SF-1RE, and/or TRE pal-bound ERRγ was subjected to proteolysis by increasing doses of trypsin as described in the figure legend, and the resulting complexes were analyzed by native PAGE. As demonstrated in Fig. 3A, sft4-, SF-1RE-, or TRE-Pal-bound ERRγ digests, respectively, showed distinctively different patterns.
Protease Treatment of sft4-, SF-1RE-, and TRE-pal-Bound ERRγ Induces Different Patterns A, Purified GST-ERRγ was incubated with 5000 cpm of end-labeled oligonucleotides corresponding to sft4, SF-1RE, or TRE-pal. After incubation for 20 min, samples were treated with increasing doses of trypsin and incubated for 15 min; the samples were then resolved by 8% native PAGE and visualized by autoradiography. Digestion products are indicated by numbers; T0 indicates undigested products. B, In vitro transcribed and translated ERRγ was incubated with 20 pmol of oligonucleotides corresponding to sft4, SF-1RE, or TRE-pal. After incubation, increasing doses of trypsin were added and after further incubations, the reactions were resolved by 12% SDS-PAGE. Asterisks indicate undigested protein, arrows with numbers (1, 2, and 3) indicate intermediate products; R indicates trypsin-resistant products. Representative figures from three independent experiments are shown, and the data were highly reproducible.
To further confirm these differences in protease sensitivity, a different approach was employed in which radiolabeled in vitro transcribed and translated ERRγ was incubated with excess of oligonucleotides corresponding to sft4, SF-1RE, or TRE-pal, followed by digestion with the indicated doses of trypsin and analyzed by denaturing PAGE. As demonstrated in Fig. 3B, incubation with SF-1RE caused a right hand shift of the overall protease sensitivity of ERRγ in comparison to sft4 or TRE-pal. TRE-pal-bound ERRγ showed the highest trypsin sensitivity. An examination of the protease-resistant band (R) reveals that the band starts appearing at 0.5 ng in the case of TRE-pal (lane 17), whereas at the same dose the band does not appear in the case of sft4 or SF-1RE (lanes 3 and 10), and at the following dose (1 ng), although the band appears on SF-1RE, the intensity of the band is lower than both sft4 and TRE-pal (compare lanes 4, 11, and 18). Furthermore, examination of the intermediate bands (1, 2 and 3) revealed that the sensitivities of these bands were comparable between sft4 and SF-1RE; however, the stability of these bands had a further shift to left for TRE-pal (compare lanes 6, 7, 13, and 14 and 20 and 21). Taken together, Fig. 3, A and B, shows that on sft4, SF-1RE, or TRE-pal, ERRγ exhibits differential sensitivity to trypsin, which may be a result of distinct structural conformations of ERRγ while bound to these elements.
Identification of PGC-1 and RIP140 as Coregulators of ERRγ
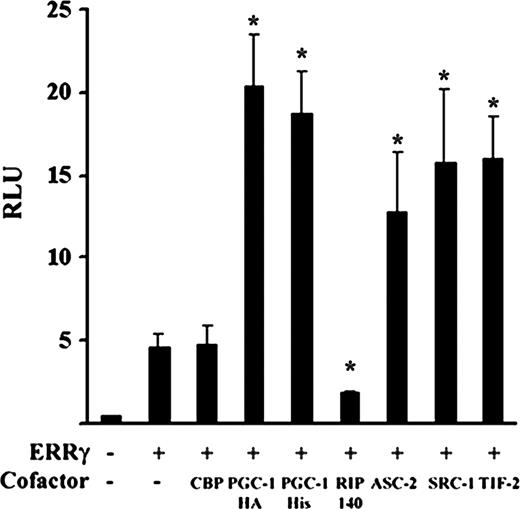
To explore the possibility that binding to different DNA sequences may alter the cofactor binding properties of ERRγ, we next attempted to identify the possible coactivator/corepressors of ERRγ. Consistent with previous findings, in transient transfection assays ERRγ activity was strongly augmented by the addition of P160 coactivators (transcription intermediary factor 2/ glucocorticoid receptor interaction protein 1, steroid receptor coactivator 1, amplified in breast cancer 1 (6, 12, 13), and RAP250/ activating signal cointegrator 2 (ASC2) (Fig. 4 and data not shown). In addition, we found that both human and mouse forms of the tissue-specific coactivator PGC-1 (HA tagged and His tagged, respectively) significantly increases ERRγ activity and that RIP140 strongly represses ERRγ activity while the cAMP response element binding protein-binding protein showed no effect (Fig. 4).
PGC-1 and RIP140 Act as Regulators of ERRγ Activity CV-1 cells were transfected with 200 ng of sft4 Luc, 200 ng of indicated cofactors, and 200 ng of RSV-β-galactosidase. Cells were lysed 48 h after transfection, and luciferase activity was measured, normalized against β-galactosidase activity, and plotted as relative light units (RLU). Bars represent mean ± se. Data represent three independent experiments performed in duplicate. Asterisks indicate the mean values are significantly different from the receptor control (P < 0.05) as determined by Student’s t test.
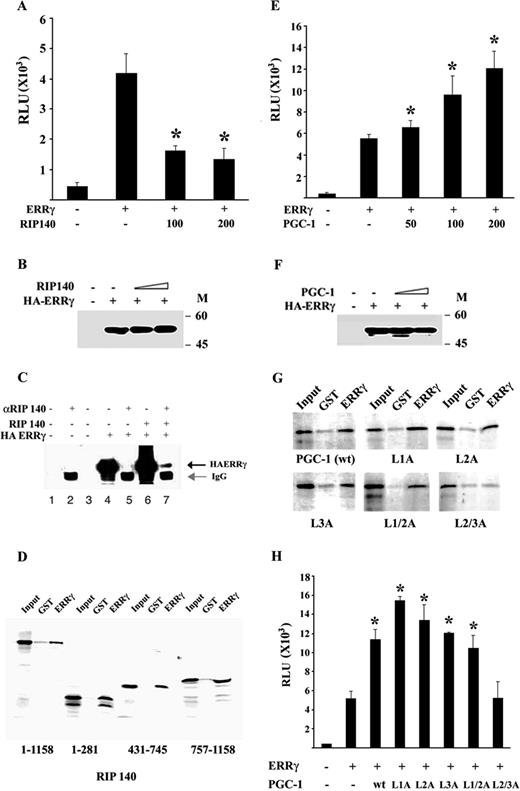
To further characterize the RIP140-mediated repression of ERRγ, transient transfection studies were performed. As demonstrated in Fig. 5A, RIP140 repressed the activity of ERRγ on sft4-Luc in a dose-dependent manner, and this repression was not due to any change in the level of ERRγ protein (Fig. 5B). To investigate whether the repressive action of RIP140 was dependent on a complex formation with ERRγ in vivo, a coimmunoprecipitation assay was performed. As demonstrated in Fig. 5C, HA-tagged ERRγ was coprecipitated with RIP140 indicating an in vivo association between ERRγ and RIP140. To further investigate whether ERRγ exhibits direct physical interaction with RIP140 and also to map the region within RIP140 responsible for such an interaction, a series of glutathione-S-transferase (GST) pull-down assays were performed. As expected, ERRγ interacted with RIP140 in solution, and the interaction interfaces included both the amino and carboxy termini of RIP140. Interestingly, the central part of RIP140 encoding amino acids 431–745, which overlaps with the interaction surface for testis-specific receptor 2 (TR2) (21), also formed strong complexes with ERRγ (Fig. 5D). Taken together, these results demonstrate that RIP140 may act as a corepressor for ERRγ.
RIP140 and PGC-1 Physically Interact with and Regulate ERRγ Transcriptional Activity A, CV-1 cells were transfected with 200 ng sft4 Luc, 100 or 200 ng of wild-type RIP140, and 200 ng of RSV-β-galactosidase. Cells were lysed 48 h after transfection, and luciferase activity was measured, normalized against β-galactosidase activity, and plotted as relative light units (RLU). Bars represent mean ± se. Data represent three independent experiments performed in duplicate. Asterisks indicate the mean values are significantly different from the receptor control (P < 0.05) as determined by Student’s t test. B, CV-1 cells were transfected with empty vector, PSG5HA ERRγ plus an empty vector, or increasing doses of RIP140 expression vector as indicated. Total protein isolated from the cell lysates was subjected to Western blot analysis 48 h after transfection with an anti-HA antibody. The data shown represent one of two independent experiments. C, Cos-7 cells were transfected with the indicated expression plasmids. cell lysates were immunoprecipitated 48 h after transfection with RIP140 antibody or preimmune serum as indicated in the figure. The complexes were resolved by 10% SDS-PAGE and detected by Western blotting with an anti-HA antibody. Lanes 4 and 6 represent 20% input. D, GST-ERRγ LBD was immobilized on glutathione Sepharose beads and incubated with 35S-labeled wild-type or indicated deletion mutants of RIP140. The result shown represents one of three independent experiments; the data were highly reproducible. E, CV-1 cells were transfected with 200 ng sft4 Luc, 200 ng RSV-β-galactosidase, and indicated doses of PGC-1. Cells were lysed 48 h after transfection, and luciferase activity was measured, normalized against β-galactosidase activity, and plotted as relative light units (RLU). Bars represent mean ± se. Data represent three independent experiments performed in duplicate. Asterisks indicate the mean values are significantly different from the receptor control (P < 0.05) as determined by Student’s t test. F, CV-1 cells were transfected with empty vector, PSG5HA ERRγ plus an empty vector, or increasing doses of PGC-1 expression vector as indicated. Total protein isolated from the cell lysates were subjected 48 h after transfection to Western blot analysis with an anti-HA antibody. The data shown represent one of two independent experiments. G, GST-ERRγ was immobilized on glutathione Sepharose beads and incubated with 35S-labeled wild-type or mutant PGC-1 as indicated in the figure. After extensive washing, samples were boiled in 1× sodium dodecyl sulfate sample buffer and resolved by 10% SDS-PAGE and visualized by autoradiography. GSTγ corresponds to GST ERRγ LBD. The result shown represents one of three independent experiments; the data were highly reproducible. H, CV-1 cells were transfected with 200 ng sft4 Luc, 200 ng RSV-β-galactosidase, and 200 ng of indicated PGC-1 constructs. Cells were lysed 48 h after transfection, and luciferase activity was measured, normalized against β-galactosidase activity, and plotted as relative light units (RLU). Bars represent mean ± se. Data represent three independent experiments performed in duplicate. Asterisks indicate the mean values are significantly different from the receptor control (P < 0.05) as determined by Student’s t test.
We also found that, in transient transfection assays, the tissue-specific coactivator PGC-1 strongly augments ERRγ activity in a dose-dependent manner (Fig. 5E), and this increase in ERRγ activity was not due to any change in the level of expression of ERRγ protein itself (Fig. 5F). The two proteins interacted physically in a GST-pull down assay (Fig. 5G). To determine the ERRγ interaction motif within PGC-1, a series of previously described PGC-1 mutants containing leucine to alanine substitutions in the receptor-interacting leucine-rich motifs (29) were tested for their interaction with ERRγ. Individual mutation of any of the three leucine-rich motifs (L1A, L2A, and L3A) did not significantly affect the ERRγ-PGC-1 interaction, but the double mutation of both L2 and L3 motifs (L2/3A) completely abolished this interaction, indicating that both these motifs are required for ERRγ-PGC-1 interaction (Fig. 5G). The GST pull-down assay results were supported by transient transfection data, which demonstrated that the double mutant L2/3A had no effect on the ERRγ activity, whereas the individual mutations of L1 and L2 and the double mutation of L1 and L2 (L1/2A) strongly augmented it (Fig. 5H).
DNA-Dependent Cofactor Recruitment by ERRγ
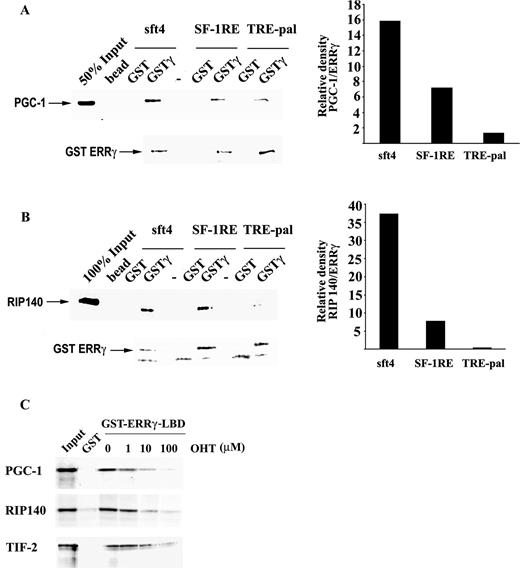
To investigate whether the response element-specific conformational alteration and the resulting activity of ERRγ were due to differential cofactor recruitment, a DNA-dependent pull-down assay was performed. GST-purified ERRγ was bound to immobilized sft4, SF-1RE, and TRE-pal and, after washing out of excess ERRγ, the complexes were incubated with radiolabeled RIP140 and/or PGC-1. As demonstrated in Fig. 6A, sft4-bound ERRγ showed the strongest affinity for PGC-1 whereas SF-1RE-bound ERRγ had a lower affinity. Interestingly, TRE-pal-bound ERRγ exhibited by far the lowest affinity for PGC-1 (compare with the lower panel, which shows Coomassie blue staining of the same gel and is used as a loading control; the position of ERRγ is indicated), suggesting that the lack of ERRγ activity on TRE-pal and the lower activity on SF-1RE may be a direct result of reduced coactivator binding. Surprisingly, however, TRE-pal-bound ERRγ showed no interaction with RIP140, whereas it strongly interacted with both sft4- and SF-1RE-bound ERRγ (Fig. 6B). Although TRE-pal-bound ERRγ was expected to bind a corepressor more efficiently than sft4-bound ERRγ, given the property of RIP140 to interact with nuclear receptors in their active conformations as in the case of liganded receptors, our results demonstrate that TRE-pal-bound ERRγ is in an inactive state, and that SF-1RE-bound ERRγ is in a partially active state, where its activity may be regulated by the cellular levels of RIP140. In agreement with these results and previous reports (2, 4) OHT, an inverse agonist of ERRγ, significantly reduced the interactions between ERRγ and RIP140 (Fig. 6C) and also the interactions between ERRγ and PGC-1 and transcription intermediary factor 2 (Fig. 6C).
Recruitment of RIP140 and PGC-1 to ERRγ Is Regulated by the Sequence of the Response Element A and B, ABCD assays were performed with immobilized sft4, SF-1RE, or TRE-pal oligonucleotides. Purified GST or full-length GST-ERRγ was allowed to interact with DNA. Unbound proteins were removed by extensive washing followed by incubation with 35S-labeled PGC-1 (A) or RIP140 (B). After extensive washing the complexes were resolved by SDS-PAGE. The lower panels indicate the Coomassie blue-stained gels that serve as loading controls, and the position of GST-ERRγ in the gels is indicated. The bands corresponding to PGC-1 or RIP140 in the autoradiographs were quantitated and normalized against the respective GST-ERRγ bands in the Coomassie blue-stained gels using Imagegauge software (Fuji Photo Film Co., Ltd., Tokyo, Japan) and plotted. Similar results were obtained in two independent experiments. C, Immobilized GST ERRγ LBD was preincubated with 1, 10, or 100 μm OHT, followed by incubation with the indicated coregulators. After extensive washing the complexes were resolved by SDS-PAGE and visualized by autoradiography. Similar results were obtained in three independent experiments.
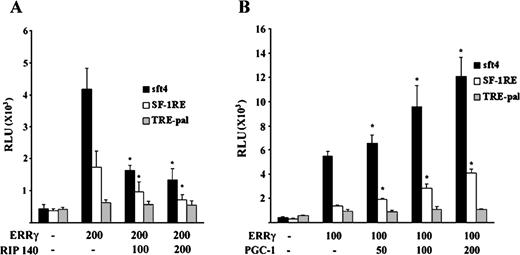
To further investigate whether the DNA-dependent differential cofactor recruitment by ERRγ can actually occur in vivo, a series of transient transfection assays were performed. As demonstrated in Fig. 7A, RIP140 did not inhibit ERRγ activity on TRE-pal, although it efficiently inhibited ERRγ activity on sft4 and SF-1RE. Similarly, PGC-1 robustly augmented ERRγ activity on sft4 and modestly augmented ERRγ activity on SF-1RE but failed to do so on a TRE-pal-driven reporter. Taken together, these results indicate that binding to different DNA elements alters the coregulator binding properties of ERRγ.
Overexpression of RIP140 or PGC-1 Selectively Regulates the Transcriptional Activity of ERRγ Depending on the Response Element A, CV-1 cells were transfected with 200 ng of indicated reporters, 200 ng of ERRγ or empty vector, indicated doses of RIP140, and 200 ng of RSV-β-galactosidase. B, CV-1 cells were transfected with 200 ng of indicated reporters, 200 ng of ERRγ or empty vector, indicated doses of PGC-1 and 200 ng of RSV-β-galactosidase. Cells were lysed 48 h after transfection, and luciferase activity was measured, normalized against β-galactosidase activity, and plotted as relative light units (RLU). Bars represent mean ± se. Data represent three independent experiments performed in duplicate. Asterisks indicate the mean values are significantly different from the receptor control (P < 0.05) as determined by Student’s t test.
DISCUSSION
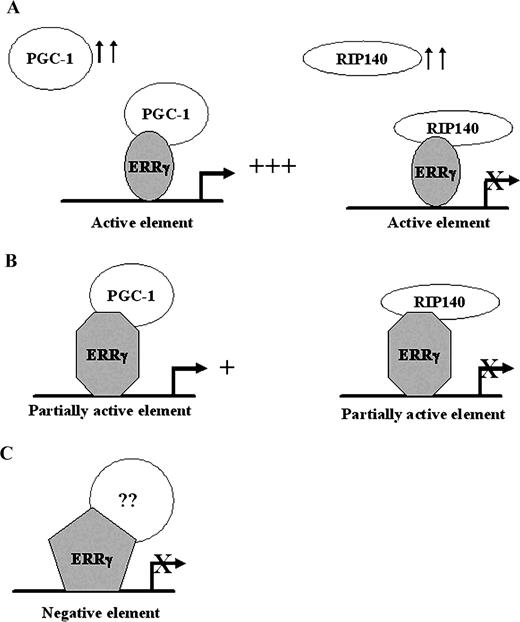
In this report we have demonstrated that the transcriptional activity of the orphan nuclear receptor ERRγ is regulated by the sequence of its response elements. Our studies provide evidence that individual response elements may alter the conformation of ERRγ, which in turn may alter its coregulator binding properties. We have also identified two such coregulators, namely PGC-1 and RIP140, which exhibit a similar tissue distribution as ERRγ. Several lines of evidence provided in this study, including in vitro and in vivo interaction of these proteins with ERRγ and response element-specific regulation of ERRγ-mediated transcription by these proteins, indicate that the sequence of ERRγ response elements as well as the relative levels of these two cofactors may regulate ERRγ activity and expression of ERRγ target genes in vivo. Figure 8 represents a model for the DNA-dependent regulation of ERRγ activity. Induction of PGC-1 or RIP140 level strongly augments or represses ERRγ activity, respectively, on active elements such as sft4, on which the ERRγ becomes highly permissive for binding these coregulators. On a partially active element, such as an SF-1RE, the ERRγ becomes less permissive for PGC-1 binding, indicating that induction of PGC-1 may lead to moderate activation of such response elements, and an induction of RIP140 may strongly repress it. However, RIP140 fails to recognize the altered structure of ERRγ on a negative element (TRE-pal), and PGC-1 shows a weak interaction. It is possible that ERRγ may interact with some as yet unidentified corepressor while it is bound to a TRE-pal.
Model for DNA- and Coregulator-Dependent Transcription by ERRγ Induction of PGC-1 or RIP140 differentially regulates ERRγ activity on active, partially active, or inactive ERRγ-response elements owing to the DNA-dependent structural alterations of the ERRγ. See Discussion for further explanations.
Although ERRα and -β were the first orphan nuclear receptors described and their physiological roles have been addressed in detail (1, 3, 5, 9, 13, 20, 30–36), relatively little understanding has so far been achieved regarding the mechanism of action of the members of this nuclear receptor subfamily, ERRγ being its newest member. Structural analyses and several functional studies have identified ERRγ as a true orphan nuclear receptor that can bind cofactors and function as an activator of transcription in the absence of ligand (6, 8, 14). However, the detailed mechanism of ERRγ activity and the regulation of its activity remain obscure. Although ERRγ can bind a wide range of DNA elements ranging from extended half-sites derived from sft4, a functional and natural ERRγ response element (8), consensus SF-1RE, and double half-sites including inverted palindromes IR3 (ERE), IR0 (TRE-pal), and direct repeats (DR0) (7, 12, 13, 37–40), it exhibits highly selective transcriptional activity on these elements. Whereas it robustly activates an sft4-driven reporter, it shows moderate activity on SF-1RE and no significant activity on a TRE-pal, indicating that, in addition to being a transcriptional activator, it may also act as efficient repressor of some of its target genes where it may suppress SF-1- and TR-mediated transcription by competition for their binding sites. ChIP assays also support this hypothesis because ERRγ bound to all three response elements but only caused hyperacetylation of histone H3 when associated with sft4 and SF-1RE. A detailed investigation of this response element-specific transcriptional property of ERRγ revealed that, upon binding different DNA elements, ERRγ exhibits differential sensitivity to trypsin, which most probably reflects a DNA-induced structural alteration. Such DNA-dependent conformational changes have also been described for a number of other nuclear receptors including ER, thyroid hormone receptor (TR) (41–44), and transcription factors such as the POU domain-containing transcription factor Pit-1 (45). Pit-1 activates transcription when bound to its recognition site on the prolactin (PRL) gene whereas, upon binding its recognition element on the GH gene, Pit-1 represses transcription. Recent crystallographic studies of Pit-1 bound to the PRL or GH gene recognition sequences have shown dramatic differences in the domain spacing and resulting binding of nuclear receptor corepressor to Pit-1 when bound to its recognition sequence on PRL or GH genes (45). However, it also is formally possible that ERRγ may bind differently to each of the ERR response elements described here in ways that affect the availability of the cofactor interaction interfaces and result in these differential transcriptional activities without changing the structural conformation, and further studies are necessary to accurately elucidate the mechanism for this DNA-dependent cofactor interaction of ERRγ.
In this report we also describe the identification of two coregulators of ERRγ, namely PGC-1 and RIP140. Whereas PGC-1 acts as a strong coactivator for ERRγ, RIP140 acts as a corepressor. A number of nuclear receptors, including PPAR, ER, glucocorticoid receptor, TR, and mineralocorticoid receptor, have been identified as interacting partners for PGC-1. However, the uniqueness of the PGC-1 and ERRγ interaction lies in the fact that, unlike with other receptors, PGC-1 interaction with ERRγ requires both the second and third leucine-rich motifs, while for other receptors only the second leucine-rich motif is responsible for the interaction (16, 20, 46, 47). While this study was in progress, a number of reports have also demonstrated that PGC-1 acts as a coactivator for both ERRα and ERRγ (37–40, 48), thus confirming our results. However, despite the fact that ERRα and ERRγ share a highly similar LBD sequence, ERRα-PGC-1 interaction is dependent mainly on the third leucine-rich motif of PGC-1, which resides in the repressor domain of this coactivator (37, 38). The RIP140-ERRγ interaction is also unique when compared with other nuclear receptors, which interact with either the amino terminus, or carboxy terminus, or both the receptor-interacting domains of RIP140 (22–27), because the ERRγ interaction with RIP140 is also mediated by the central part of this molecule. A similar interaction has been reported in the case of TR2, which can also interact with the central part of RIP140; unlike ERRγ, however, it fails to recognize the carboxy terminus receptor-interacting domain of RIP140 (21).
Because RIP140 interacted with ERRγ in solution we thought that the stronger interaction of ERRγ with RIP140 on a TRE-pal might be the reason for the relative inactivity of ERRγ on a TRE-pal, but ABCD assays revealed that TRE-pal-bound ERRγ fails to interact with RIP140. Because RIP140 interacts with liganded nuclear receptors in their active state (22–28), it seems plausible that ERRγ may adopt an inactive state on TRE-pal, which is not recognized by RIP140. This hypothesis is further strengthened by the fact that ERRγ bound to TRE-pal showed little or no interaction with PGC-1. Consistent with the results from the ABCD assay, transient transfection assays revealed that, unlike on sft4 and SF-1RE, the transcriptional activity of ERRγ on TRE-pal was not affected by overexpression of RIP140 or PGC-1. It may also be reasonable to assume that TRE-pal-bound ERRγ may bind some as yet unidentified corepressor. Because both RIP140 and PGC-1 share a very similar tissue distribution with ERRγ (8, 16, 21), it is highly plausible that these two coregulators may provide a response element-dependent yin-yang control of ERRγ-mediated transcription in vivo.
Considering that the total number of nuclear receptors in vertebrates discovered to date is around 50, and with an extremely small chance of discovery of new nuclear receptors, it remains intriguing how this relatively limited number of nuclear receptors critically regulates nearly every aspect of vertebrate development, adult physiology, and even pathophysiology. Recent studies indicate that the mechanisms involved in diversification of the function of a nuclear receptor may include: 1) structural modifications after alternative splicing or alternative promoter usage, giving rise to different A/B regions where different N termini can give rise to receptor isoforms with distinct transcriptional activities; 2) posttranslational modifications of nuclear receptors from acetylation to phosphorylation, leading to differential activation potency of the nuclear receptors and even altering their subcellular localization; 3) spatial and temporal expression, relative levels, and relative specificities of coregulator proteins, and 4) response element-driven alterations in the conformation of the receptor. These conformational changes may also influence the structural and functional properties of the coregulator proteins as evidenced by the homeodomain transcription factor HNF-1α (49), where a loss of function mutation actually increases the interaction of the transcription factor with coactivator/cointegrator proteins cAMP response element binding protein-binding protein and pCAF; such interaction, however, obliterates the histone acetylase property of these proteins. Similarly, when PPARα is bound to its target element on acyl-coenzyme A (CoA) oxidase (AOx) gene, the receptor activity is inhibited by overexpression of SHP, but, conversely, when PPARα binds to its target element on enoyl-CoA hydratase/3-hydroxyacyl-CoA dehydrogenase gene promoter, its activity is augmented by overexpression of SHP (50). These data, together with our report, clearly demonstrate the importance of cis elements in regulation of transcription.
In summary, our report demonstrates that ERRγ activity depends on the response elements that it binds, leading to differential interaction with coregulator proteins in the presence of DNA. Furthermore, physical interactions of ERRγ with two different coregulator proteins, namely PGC-1 and RIP140, which have a similar tissue distribution with ERRγ indicate that the relative levels of these two proteins may regulate ERRγ transcriptional activity in vivo.
MATERIALS AND METHODS
Plasmids
PSG5HA ERR3/ERRγ was a kind gift from Dr. Michael R. Stallcup; PGL2–3× SF-1RE luc was a kind gift from Dr. Jean Marc Vanacker; and 3× TRE-pal TATA luc was kindly donated by Dr. Philippe Lefebvre. The HA-tagged expression plasmids for wild-type and mutant mouse PGC-1 were kindly provided by Dr. Anastasia Kralli; the His-tagged human PGC-1 was a kind gift from Dr. Bruce Spiegelman; and the expression plasmids for wild-type RIP140 and the deletion mutants were kind gifts from Dr. Johanna Zilliacus. PGL2–3X sft4, GST ERRγ full length, and GST ERRγ LBD plasmids are described elsewhere (8). PGL3–3× sft4 and SF-1RE were constructed by restriction digestion of PGL2–3× sft4 or SF-1RE with KpnI and BglII and cloning the fragments into PGL3 promoter vector (Promega Corp., Madison, WI). PGL3–3× TRE-pal were constructed by PCR amplification of the element from 3× TRE-pal TATA luc, and recloning the PCR product into SmaI and BglII sites of the PGL3 promoter vector. PGL3–1× sft4, SF-1RE, and TRE-pal were constructed by annealing oligonucleotides corresponding to the respective elements and cloning them into 5′-KpnI and 3′-BglII sites of PGL3 promoter vector. All the constructs were checked by sequencing.
Cell Culture, Transient Transfection Assays, and Western Blotting
CV-1, HeLa, and Cos-7 cells were maintained in DMEM (Invitrogen, San Diego, CA) in the presence of 10% fetal bovine serum (Invitrogen). For luciferase assays, CV-1 or HeLa cells were plated on 24-well plates and after 24 h the cells were transfected by Lipofectamine Plus reagent (Invitrogen) according to the manufacturer’s instructions. For luciferase assays, cells were transfected with 200 ng of reporter gene, 200 ng of the internal control, Rous sarcoma virus (RSV)-β-galactosidase, and varying doses of mammalian expression plasmids. Total DNA in each transfection was adjusted to 1 μg by adding the appropriate amount of pcDNA3 vector. Approximately 48 h after transfection, cells were harvested, and the luciferase activity was measured as described previously (8) and normalized against β-galactosidase activity. The β-galactosidase activity was not affected by cotransfection of either ERRγ or the cofactors used in this study. Western blottings were performed as described elsewhere (8). Briefly, CV-1 cells were plated on 10-cm dishes and transfected with pcDNA3 alone (Mock) or PSG5HA ERRγ with or without increasing doses of cofactor plasmids. Cells were lysed 48 h after transfection, total protein was harvested, and 20 μg of total protein, as determined by Bradford assay (Sigma Chemical Co., St. Louis, MO), was resolved by 12% SDS-PAGE and immobilized on nitrocellulose membranes (Hybond-C, AP Biotech, Uppsala, Sweden), and HA ERRγ was detected using a monoclonal antiserum against HA (MMS101R-500, BAbCO, Richmond, CA).
Protein Interaction Tests
GST pull-down assays were performed as described elsewhere (8). Briefly, GST-ERRγ or GST only were expressed in Escherichia coli BL21 (DE3) pLys bacterial culture and recovered on glutathione-Sepharose 4B beads (Amersham Pharmacia Biotech, Arlington Heights, IL), The GST fusion proteins were incubated with 35S-labeled coregulators that were expressed by coupled in vitro transcription and translation (TNT, Promega Corp.) in the presence or absence of OHT as indicated in the figure legends. After incubation, specifically bound proteins were eluted, resolved by SDS-PAGE, and analyzed by autoradiography. The coimmunoprecipitation assay was conducted as described previously (51). Briefly, Cos-7 cells were plated on 15-cm dishes, and after 24 h the cells were transfected with PSG5 HA ERRγ plus an empty vector or RIP140 expression vector. A green fluorescent protein-expressing plasmid, EGFPC1 (CLONTECH, Palo Alto, CA) was also transfected as an internal control, and the efficiency of transfection as determined by visual inspection was about 50%. Cell extracts were incubated 48 h post transfection with anti-RIP140 antibody (sc-8997, Santa Cruz Biotechnology, Inc., Santa Cruz, CA) followed by Western blot analysis of the immunocomplexes with a monoclonal anti-HA antibody (BAbCO).
EMSA
Bacterially expressed GST-fused ERRγ was purified using GST Sepharose beads (Amersham Pharmacia Biotech), as described elsewhere (8). The purified GST ERRγ was incubated with 10,000 cpm of end-labeled oligonucleotide probes in the presence or absence of indicated fold molar excesses of competitor oligonucleotides. The samples were resolved by 5% nondenaturing PAGE and visualized by autoradiography. The sft4 sequence corresponds to GATCCACATGACTTCTGGAGTCAAGGTTGTTTGGA, where the core ERRγ binding sequence is underlined. The oligonucleotide designated as SF-1RE had the same flanking sequences as sft4 with mutations only in the core sequences as represented in the figure.
Limited Proteolysis Assay
Limited proteolysis assay by EMSA was performed as described elsewhere (8). Briefly, bacterially expressed and purified GST-ERRγ was incubated with 5000 cpm of end-labeled oligonucleotides corresponding to sft4, SF-1RE, and TRE-pal in EMSA binding buffer in a reaction volume of 10 μl. Each reaction contained 10 mm Tris, pH 8.0; 40 mm KCl, 6% glycerol, 0.05% Nonidet P40, 1 mm dithiothreitol, 4 mm spermidine, 0.05 mm ZnCl2, 2.5μg BSA, 500 ng poly(dI-dC), and 200 ng of sheared and denatured salmon sperm DNA. After 10 min incubation on ice, 0, 1, 3, 5, 10, or 20 ng of trypsin (Promega Corp.) was added and the incubation continued for another 15 min at room temperature. After incubation the reactions were stopped by mixing 10 μl of 1× EMSA binding buffer and putting the samples on ice. The samples were then immediately subjected to 8% polyacrylamide gel electrophoresis and analyzed by autoradiography. Limited proteolysis assay by SDS-PAGE was performed by incubating 6 μl of radiolabeled in vitro transcribed and translated ERRγ with 20 pmol of sft4, SF-1RE, or TRE-pal in the presence of EMSA buffer in a reaction volume of 10 μl. After a 15-min incubation on ice, 0, 0.5, 1, 3, 5, 10, or 20 ng of trypsin were added and the incubation continued for another 15 min at room temperature. The reactions were stopped by adding an equal volume of 2× sodium dodecyl sulfate loading buffer heated at 95 C for 5 min and were resolved by 12% SDS-PAGE. The gels were then dried and analyzed by autoradiography.
DNA-Dependent Protein-Protein Interaction (ABCD) Assay
ABCD assays were performed as described elsewhere (52) with minor modifications. Briefly, the oligonucleotides corresponding to sft4, SF-1RE, or TRE-pal were prepared by annealing a 5′-biotinylated forward strand to the reverse strand. Annealed oligos (10 pmol) were immobilized on 100 μg of streptavidin paramagnetic beads (Promega Corp.). The immobilized oligos were then incubated with 1 μg of purified GST-ERRγ, and after extensive washing the immobilized DNA-bound ERRγ was incubated with in vitro transcribed and translated 35S-labeled PGC-1 or RIP140. After further washing the specifically bound complexes were resolved on SDS-PAGE and analyzed by autoradiography.
ChIP Assay
ChIP assays were performed as described elsewhere (53) with minor modifications. Briefly, Cos-7 cells were plated in 15-cm dishes and 24 h after plating the cells were transfected with sft4, SF-1RE, or TRE-pal luciferase reporter plasmids in the presence or absence of HA ERRγ and/or indicated cofactors by standard DEAE dextran method. Soluble chromatin from these cells was prepared 48 h after transfection and immunoprecipitated with monoclonal antibodies against HA (BAbCO), acetylated histone3 (H3) (06–599; Upstate Biotechnology, Inc., Lake Placid, NY), 6× histidine (His) (8916–1; CLONTECH) or polyclonal antibody against RIP140 (Santa Cruz Biotechnology, Inc.). The final DNA extractions were amplified by PCR using primers specific to vector sequences flanking the sft4, SF-1RE, or TRE-pal sites and analyzed on agarose gels.
Acknowledgments
We thank Drs. Michael R. Stallcup, Jean Marc Vanacker, Philippe Lefebvre, Bruce Spiegelman, Anastasia Kralli, and Johanna Zilliacus for kindly supplying plasmid DNAs, and members of the Unit for Receptor Biology at the Department of Biosciences at Novum for sharing materials and for fruitful discussions.
This work was supported by Swedish Cancer Society and Karo Bio AB. H.S.C. was supported by Korean Research Foundation Grant KRF-2000-015-DS0037.
Abbreviations:
- ChIP,
Chromatin immunoprecipitation;
- CoA,
coenzyme A;
- ER,
estrogen receptor;
- ERE,
estrogen response element;
- ERR,
ER-related receptor;
- GST,
glutathione-S-transferase;
- HA,
hemagglutinin;
- H3,
histone 3;
- LBD,
ligand-binding domain;
- OHT,
4-hydroxytamoxifen;
- PGC-1,
peroxisome proliferator-activated receptor γ coactivator 1;
- PPARγ,
peroxisome proliferator-activated receptor γ;
- RIP140,
receptor-interacting protein 140;
- RSV,
Rous sarcoma virus;
- SF-1RE,
steroidogenic factor 1 response element;
- SHP,
small heterodimer partner;
- TR,
thyroid hormone receptor;
- TRE-pal,
palindromic thyroid hormone response element.