Supporting data for "Ximmer: A System for Improving Accuracy and Consistency of CNV Calling from Exome Data"
Dataset type: Virtual-Machine, Software, Workflow
Data released on August 23, 2018
Sadedin SP; Ellis JA; Masters SL; Oshlack A (2018): Supporting data for "Ximmer: A System for Improving Accuracy and Consistency of CNV Calling from Exome Data" GigaScience Database. https://doi.org/10.5524/100495
While exome and targeted next generation DNA sequencing are primarily used for detecting single nucleotide changes and small indels, detection of copy number variants (CNVs) can provide highly valuable additional information from the data. Although there are dozens of exome CNV detection methods available, these are often difficult to use and accuracy varies unpredictably between and within data sets.
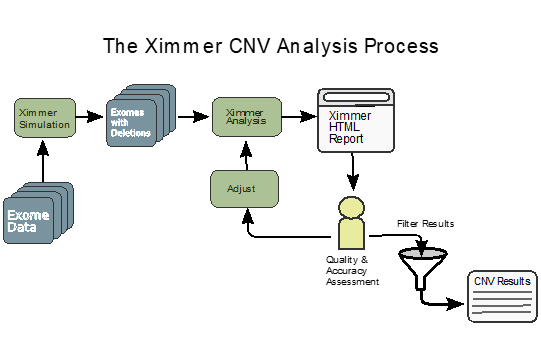
We present Ximmer, a tool which supports an end to end process for evaluating, tuning and running analysis methods for detection of CNVs in germline samples. Ximmer includes a simulation framework, implementations of several commonly used CNV detection methods, and a visualisation and curation tool which together enable interactive exploration and quality control of CNV results. Using Ximmer, we comprehensively evaluate CNV detection on four data sets using five different detection methods. We show that application of Ximmer can improve accuracy and aid in quality control of CNV detection results. In addition, Ximmer can be used to run analyses and explore CNV results in exome data.
Ximmer offers a comprehensive tool and method for applying and improving accuracy of CNV detection methods for exome data.
Additional details
Read the peer-reviewed publication(s):
- Sadedin, S. P., Ellis, J. A., Masters, S. L., & Oshlack, A. (2018). Ximmer: a system for improving accuracy and consistency of CNV calling from exome data. GigaScience, 7(10). https://doi.org/10.1093/gigascience/giy112 (PubMed:30192941)
Additional information:
https://github.com/ssadedin/ximmer
Accessions (data not in GigaDB):
Click on a table column to sort the results.
Table Settings| File Name | Description | Sample ID | Data Type | File Format | Size | Release Date | File Attributes | Download |
|---|---|---|---|---|---|---|---|---|
| Description of the deposited files. | Readme | TEXT | 2.85 kB | 2018-08-15 | MD5 checksum: cf542268ed6e87eeb014ccd3cb7656d1 |
|||
| Archival copy of the github repository https://github.com/ssadedin/ximmer downloaded 14 Aug 2018. Ximmer is a tool designed to help users of exome and targeted genomic sequencing data accurately detect and interpret copy number variants. Please visit the GitHub repository for the most recent version. | GitHub archive | archive | 226.06 MB | 2018-08-15 | license: Lesser General Public License MD5 checksum: fc40dcd7522ac1539c946f2f602d9060 |
|||
| Zip archive containing the Ximmer report for the SureSelect samples analysed in the manuscript. | Structural variation | zip | 1.37 MB | 2018-08-15 | MD5 checksum: 40c23febdb58d67551343bfcb0709559 |
|||
| Zip archive containing the HTML Ximmer report for the Nimblegen samples analysed in the manuscript. | Structural variation | zip | 925.10 kB | 2018-08-15 | MD5 checksum: a7933bd8af5d2e8dd637cb23ca3b30b8 |
|||
| Ximmer report for the Nextera samples analysed in the manuscript are not available to the public due to ethical restrictions on the dataset used. Readers are encouraged to contact the authors directly if they wish to gain access to these data. | Other | TEXT | 528 B | 2018-08-15 | MD5 checksum: d240419fea857209d680749105a9d809 |
|||
| Ximmer report for the TruSeq samples analysed in the manuscript are not available to the public due to ethical restrictions on the dataset used. Readers are encouraged to contact the authors directly if they wish to gain access to these data. | Other | TEXT | 528 B | 2018-08-15 | MD5 checksum: d240419fea857209d680749105a9d809 |
| Date | Action |
|---|---|
| August 23, 2018 | Dataset publish |
| August 28, 2018 | Manuscript Link added : 10.1093/gigascience/giy112 |
| August 30, 2018 | Manuscript Link removed : 10.1093/gigascience/giy112 |
| November 3, 2018 | Manuscript Link added : 10.1093/gigascience/giy112 |
| November 29, 2018 | File ximmer-report-phs001272.txt updated |
| November 11, 2022 | Manuscript Link updated : 10.1093/gigascience/giy112 |