Background: Due to antibiotic abuse, the problem of bacterial
resistance is becoming increasingly serious, and rapid detection of bacterial
resistance has become an urgent issue. Because under the action of antibiotics,
different active bacteria have different metabolism of heavy water, antibiotic
resistance of bacteria can be identified according to the existence of a C-D peak
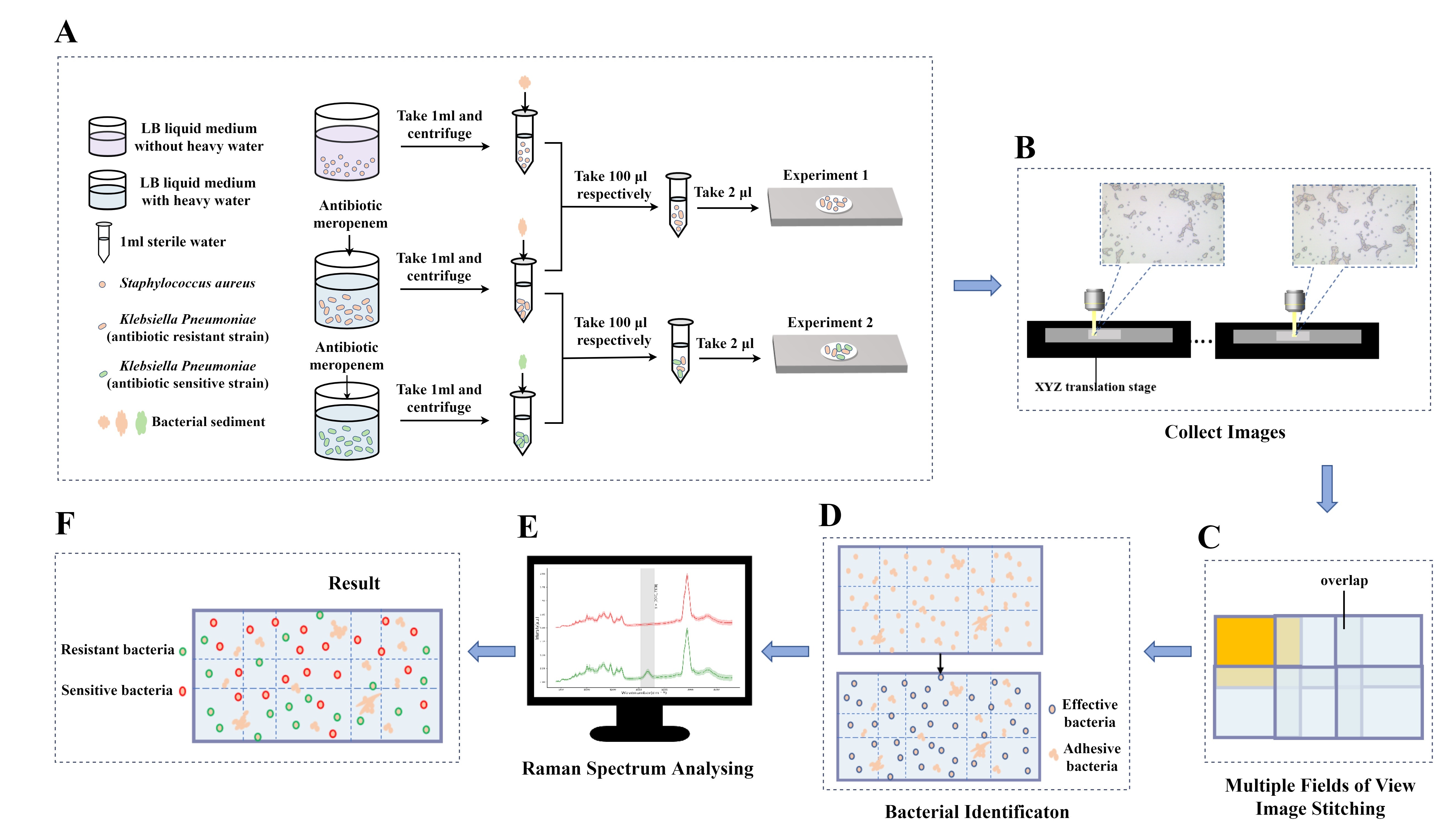
in the 2030–2400 cm range in the Raman spectrum. Methods: To
ensure data veracity, a large number of bacteria need to be detected, however,
due to the limitation of the field of view of the high magnification objective,
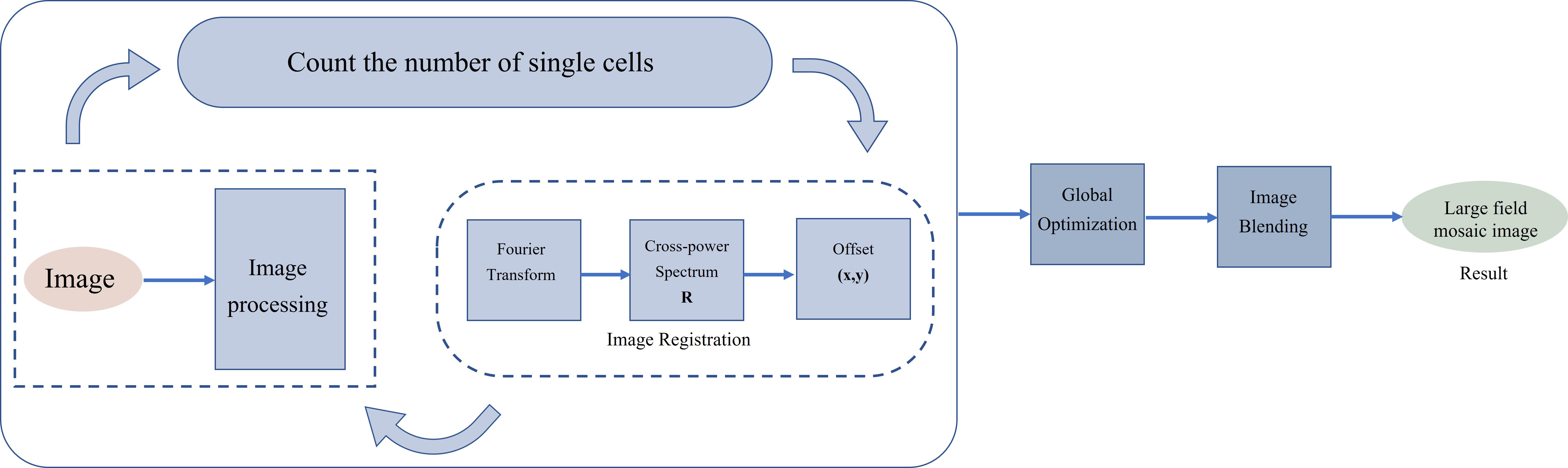
the number of single cells in a single field of view is very small. By combining
an image stitching algorithm, image recognition algorithm, and processing of
Raman spectrum and peak-seeking algorithm, can identify and locate single cells
in multiple fields of view at one time and can discriminate whether they are
Antimicrobial-resistant bacteria. Results: In experiments 1 and 2, 2706
bacteria in 9 11 fields of view and 2048 bacteria in 11 11
fields of view were detected. Results showed that in experiment 1, there are 1137
antibiotic-resistant bacteria, accounting for 42%, and 1569 sensitive bacteria,
accounting for 58%. In experiment 2, there are 1087 antibiotic-resistant
bacteria, accounting for 53%, and 961 sensitive bacteria, accounting for 47%.
It showed excellent performance in terms of speed and recognition accuracy as
compared to traditional manual detection approaches. And solves the problems of
low accuracy of data, a large number of manual experiments, and low efficiency
due to the small number of single cells in the high magnification field of view
and different peak-seeking parameters of different Raman spectra.
Conclusions: The detection and analysis method of bacterial Raman
spectra based on image stitching can be used for unattended, automatic, rapid and
accurate detection of single cells at high magnification with multiple fields of
view. With the characteristics of automatic, high-throughput, rapid, and accurate
identification, it can be used as an unattended, universal and non-invasive means
to measure antibiotic-resistant bacteria to screen for effective antibiotics,
which is of great importance for studying the persistence and spread of
antibiotics in bacterial pathogens.