Abstract
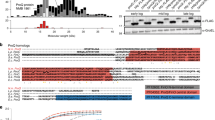
Currently, a number of structurally and functionally different thermosensitive elements, such as structurally and functionally different RNA thermometers, for controlling a variety of biological processes in bacteria, including virulence are known. These well-known RNA thermometers are structures, whether matched or mismatched, which are represented by either a single stretched hairpin structure or a few hairpins. Based on computer and thermodynamic analyses of 25 isolates of Salmonella enterica with complete genome, we have developed an algorithm and criteria to search for potential RNA thermometers, which will enable us to undertake a future search for potential riboswitches in the genomes of other socially significant pathogens. In addition to the well-known 4U RNA thermometer, another four hairpin-loop structures have been identified in S. enterica as new potential RNA thermometers and two of them are localized in 5′-UTR of virulence regulators gltB and yaeQ. They are highly conserved noncanonical structures and correspond to the necessary and sufficient conditions for forming RNA thermometers, since they are found in each of the 25 S. enterica genome isolates. We analyzed the thermosensitive motif in the pXO1 plasmid of Bacillus anthracis—an anthrax-causative pathogen—and visualized matched hairpins that form a cruciform structure in pUC8 supercoiled plasmid by atomic force microscopy.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Doudna, J. and Cech, T., The chemical repertoire of natural ribozymes, Nature, 2002, vol. 418, no. 6894, pp. 222–228.
Watson, P. and Fedor, M., The glmS riboswitch integrates signals from activating and inhibitory metabolites in vivo, Nat. Struct. Mol. Biol., 2011, vol. 18, no. 3, pp. 359–363.
Narberhaus, F. and Vogel, J., Regulatory RNAs in prokaryotes: here, there and everywhere, Mol. Microbiol., 2009, vol. 74, no. 2, pp. 261–269.
Klinkert, B. and Narberhaus, F., Microbial thermosensors, Cell Mol. Life Sci., 2009, vol. 66, no. 16, pp. 2661–2676.
Morita, M.T. Translational induction of heat shock transcription factor r32: evidence for a built-in RNA thermosensor, Genes Dev., 1999, vol. 13, no. 6, pp. 655–665.
Lybecker, M.C. and Samuels, D.S., Temperature-induced regulation of RpoS by a small RNA in Borrelia burgdorferi, Mol. Microbiol., 2007, vol. 64, no. 4, pp. 1075–1089.
Waldminghaus, T., Fippinger, A., Alfsmann, J., and Narberhaus, F., RNA thermometers are common in alpha- and gamma-proteobacteria, Biol. Chem., 2005, vol. 386, no. 12, pp. 1279–1286.
Waldminghaus, T., Heidrich, N., Brantl, S., and Narberhaus, F., FourU: a novel type of RNA thermometer in Salmonella, Mol. Microbiol., 2007, vol. 65, no. 2, pp. 413–424.
Hoe, N.P. and Goguen, J.D., Temperature sensing in Yersinia pestis: translation of the LcrF activator protein is thermally regulated, J. Bacteriol., 1993, vol. 175, no. 24, pp. 7901–7909.
Waldminghaus, T., Gaubig, L.C., and Narberhaus, F., Genome-wide bioinformatic prediction and experimental evaluation of potential RNA thermometers, Mol. Genet. Genom., 2007, vol. 278, pp. 555–564.
Neupert, J., Karcher, D., and Bock, R., Design of simple synthetic RNA thermometers for temperature-controlled gene expression in Escherichia coli, Nucleic Acids Res., 2008, vol. 36, no. 19, p. e124.
Wieland, M. and Hartig, J.S., RNA quadruplex-based modulation of gene expression, Chem. Biol., 2007, vol. 14, no. 7, pp. 757–763.
UNAFold: software for nucleic acid folding and hybridization. http://mfold.bioinfo.rpi.edu/cgi-bin/rna-form1.cgi
Brodskii, L.I., Drachev, A.L., Tatuzov, R.L., and Chumakov, K.M., The software package for biopolymer sequence analysis: GeneBee, Biopolym. Cell, 1991, vol. 7, pp. 10–14.
Limanskaya, O. and Limanskii, A., Imaging compaction of single supercoiled DNA molecules by atomic force microscopy, General Physiol. Biophys., 2008, vol. 27, no. 4, pp. 322–337.
Waldminghaus, T., Fippinger, A., Alfsmann, J., and Narberhaus, F., RNA thermometers are common in α- and γ-proteobacteria, Biol. Chem., 2005, no. 386, pp. 1279–1286.
Spirin, A.S., Molekulyarnaya biologiya. Struktura ribosomy i biosintez belka (Molecular Biology. The Structure of Ribosomes and Protein Synthesis), Moscow: Vyssh. shk., 1986.
Chowdhury, S., Maris, C., Allain, F.H., and Narberhaus, F., Molecular basis for temperature sensing by an RNA thermometer, EMBO J., 2006, vol. 25, no. 11, pp. 2487–2497.
Lilley, D., Hairpin-loop formation by inverted repeats in supercoiled DNA molecules, Proc. Natl. Acad. Sci. U.S.A., 1980, vol. 77, no. 11, pp. 6468–6472.
Sinden, R. and Pettijohn, D., Cruciform transitions in DNA, J. Biol. Chem., 1984, vol. 259, no. 10, pp. 6593–6600.
Lyamichev, V., Panyutin, I., and Mirkin, S., The absence of cruciform structures from pAO3 plasmid DNA in vivo, J. Biomol. Struct. Dyn., 1984, vol. 2, no. 2, pp. 291–301.
Bevilacqua, P.C. and Blose, J.M., Structures, kinetics, thermodynamics and biological functions of RNA hairpins, Ann. Rev. Phys. Chem., 2008, vol. 58, pp. 79–103.
Cantor, Ch. and Schimmel, P., Biophysical Chemistry, Part III: The Behavior of Biological Macromolecules, New York: Freeman, 1980.
Antao, V.P. and Tinoco, I., Ghermodynamic parameters for loop formation in RNA and DNA hairpin tetraloops, Nucleic Acids Res., 1992, vol. 20, no. 4, pp. 819–824.
Panyutin, I., Lyamichev, V., and Lyubchenko, Y., A sharp structural transition in pAO3 plasmid DNA caused by increased superhelix density, FEBS Lett., 1982, vol. 148, no. 2, pp. 297–301.
Panyutin, I., Klishko, V., and Lyamichev, V., Kinetics of cruciform formation and stability of cruciform structure in superhelical DNA, J. Biomol. Struct. Dyn., 1984, vol. 1, no. 4, pp. 1311–1324.
Zarudnaya, M., Potyagailo, A., and Govorun, D., Conserved structural motifs in the 3′ untranslated region of genomic RNA of SARS-CoV virus, Biopolym. Cell, 2003, vol. 19, no. 3, pp. 298–303.
Limanskii, A.P., Visualization of cruciform structure in supercoiled DNA by atomic force microscopy, Biophysics, 2000, vol. 45, no. 6, pp. 1007–1010.
De Smit, M.H. and van Duin, J., Secondary structure of the ribosome binding site determines translational efficiency: a quantitative analysis, Proc. Natl. Acad. Sci. U.S.A., 1990, vol. 87, pp. 7668–7672.
Rychlik, W., Spencer, W.J., and Rhoads, R.E., Optimization of the annealing temperature for DNA amplification in vitro, Nucleic Acids Res., 1990, vol. 18, no. 21, pp. 6409–6417.
Limanskaya, O.Yu. and Limanskii, A.P., Imaging of T7 RNA polymerase elongation complexes by atomic force microscopy, Mol. Biol. (Moscow), 2008, vol. 42, no. 3, pp. 469–477.
Limanskaya, L.A. and Limanskii, A.P., S-DNA, over-supercoiled DNA with a 1.94- to 2.19-Å rise per base pair, Mol. Biol. (Moscow), 2006, vol. 40, no. 1, pp. 107–120.
Limanskaya, L.A. and Limanskii, A.P., Compaction of single supercoiled DNA molecules adsorbed onto amino mica, Russ. J. Bioorg. Chem., 2006, vol. 32, no. 5, pp. 444–459.
Vicari, D. and Artsimovitch, I., Virulence regulators RfaH and YaeQ do not operate in the same pathway, Mol. Genet. Genom., 2004, vol. 272, no. 5, pp. 489–496.
Johansson, J., Mandin, P., Renzini, A., et al., An RNA thermosensor controls expression of virulence genes in Listeria monocytogenes, Cell, 2002, vol. 10, pp. 551–561.
Lu, Z., Turner, D., and Mathews, D., A set of nearest neighbor parameters for predicting the enthalpy change of RNA secondary structure formation, Nucleic Acids Res., 2006, vol. 34, no. 17, pp. 4912–4924.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Ukrainian Text © O.Yu. Limanskaya, L.A. Murtazaeva, A.P. Limanskii, 2013, published in Tsitologiya i Genetika, 2013, Vol. 47, No. 5, pp. 12–21.
About this article
Cite this article
Limanskaya, O.Y., Murtazaeva, L.A. & Limanskii, A.P. Potential thermosensitive riboswitches in the genome of Salmonella. Cytol. Genet. 47, 268–275 (2013). https://doi.org/10.3103/S009545271305006X
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S009545271305006X