Abstract
Background
Reports show miR-23b to be a cancer-related biomarker in various cancer types. Interestingly, it has a dual role of oncogenic and tumor-suppressive functions, depending on the cancer type. This study focused on the unknown association of miR-23b-3p with hepatocellular carcinoma (HCC).
Methods
Expression of miR-23b-3p was measured in nine HCC cell lines and 125 resected human HCC samples by TaqMan microRNA assays. To detect its downstream target, miR-23b-3p mimic and inhibitor constructs were transfected and analyzed.
Results
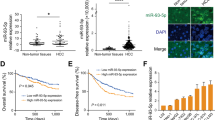
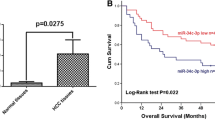
HepG2, a high miR-23b-3p-expressing cell line, was transfected with a miR-23b-3p inhibitor construct, whereas SK-Hep1, a low miR-23b-3p-expressing cell line, was transfected with a mimic construct. Proliferation of HCC cells was activated by miR-23b-3p overexpression and diminished by its knockdown. Then, 125 clinical HCC samples were examined to measure miR-23b-3p expression. Tumor expression of miR-23b-3p was upregulated in 48 cases (38%) and downregulated in 77 cases (62%). The upregulated cases were correlated with elderly patients (P = 0.015). These patients also showed significantly poor overall survival [hazard ratio (HR), 3.10; 95% conflidence interval (CI), 1.57–6.29; P = 0.001] in a multivariate analysis. Furthermore, mitochondrial metabolism-related genes (MICU3 and AUH) were detected as specific binding targets.
Conclusion
The study showed that miR-23b-3p functions as an oncogenic microRNA in HCC cell lines. Its overexpression in resected HCC tissues was a significant prognostic factor of overall survival. Both MICU3 and AUH may be candidate gene targets of miR-23b-3p.




Similar content being viewed by others
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Gan W, Huang JL, Zhang MX, et al. New nomogram predicts the recurrence of hepatocellular carcinoma in patients with negative preoperative serum AFP subjected to curative resection. J Surg Oncol. 2018; 117:1540–7.
Sugimoto H, Takeda S, Inoue S, Kaneko T, Watanabe K, Nakao A. Des-gamma-carboxy prothrombin (DCP) ratio, a novel parameter measured by monoclonal antibodies MU-3 and 19B7, as a new prognostic indicator for hepatocellular carcinoma. Liver Int. 2003;23:38–44.
Lu CY, Chen SY, Peng HL, Kan PY, Chang WC, Yen CJ. Cell-free methylation markers with diagnostic and prognostic potential in hepatocellular carcinoma. Oncotarget. 2017;8:6406–18.
Guo X, Lv X, Lv X, Ma Y, Chen L, Chen Y. Circulating miR-21 serves as a serum biomarker for hepatocellular carcinoma and correlated with distant metastasis. Oncotarget. 2017;8:44050–8.
Wu XM, Xi ZF, Liao P, et al. Diagnostic and prognostic potential of serum microRNA-4651 for patients with hepatocellular carcinoma related to aflatoxin B1. Oncotarget. 2017;8:81235–49.
Chen M, Hu W, Xiong CL, et al. miR-22 targets YWHAZ to inhibit metastasis of hepatocellular carcinoma and its down-regulation predicts a poor survival. Oncotarget. 2016;7:80751–64.
Begum S, Hayashi M, Ogawa T, et al. An integrated genome-wide approach to discover deregulated microRNAs in non-small cell lung cancer: clinical significance of miR-23b-3p deregulation. Sci Rep. 2015;5:13236.
Donadelli M, Dando I, Fiorini C, Palmieri M. Regulation of miR-23b expression and its dual role on ROS production and tumour development. Cancer Lett. 2014;349:107–113
Grossi I, Arici B, Portolani N, De Petro G, Salvi A. Clinical and biological significance of miR-23b and miR-193a in human hepatocellular carcinoma. Oncotarget. 2017;8:6955–69.
Zhuang L, Wang X, Wang Z, et al. MicroRNA-23b functions as an oncogene and activates AKT/GSK3beta/beta-catenin signaling by targeting ST7L in hepatocellular carcinoma. Cell Death Dis. 2017;8:e2804.
Tovar V, Alsinet C, Villanueva A, et al. IGF activation in a molecular subclass of hepatocellular carcinoma and pre-clinical efficacy of IGF-1R blockage. J Hepatol. 2010;52:550–9.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods San Diego Calif. 2001;25:402–8.
Vultur A, Gibhardt CS, Stanisz H, Bogeski I. The role of the mitochondrial calcium uniporter (MCU) complex in cancer. Pflugers Archiv Eur J Physiol. 2018;470:1149–63.
Brennan LE, Nakagawa J, Egger D, Bienz K, Moroni C. Characterisation and mitochondrial localisation of AUH, an AU-specific RNA-binding enoyl-CoA hydratase. Gene. 1999;228:85–91.
Wallace DC. Mitochondria and cancer. Nat Rev Cancer. 2012;12:685–98.
Zong WX, Rabinowitz JD, White E. Mitochondria and cancer. Mol Cell. 2016;61:667–76.
Fujii T, Fuchs BC, Yamada S, et al. Mouse model of carbon tetrachloride induced liver fibrosis: histopathological changes and expression of CD133 and epidermal growth factor. BMC Gastroenterol. 2010;10:10–79.
Yamada S, Nomoto S, Fujii T, et al. Correlation between copy number of mitochondrial DNA and clinico-pathologic parameters of hepatocellular carcinoma. Eur J Surg Oncol. 2006;32:303–7.
Ohsawa S, Sato Y, Enomoto M, Nakamura M, Betsumiya A, Igaki T. Mitochondrial defect drives non-autonomous tumour progression through Hippo signalling in Drosophila. Nature. 2012;490:547–51.
Barajas JM, Reyes R, Guerrero MJ, Jacob ST, Motiwala T, Ghoshal K. The role of miR-122 in the dysregulation of glucose-6-phosphate dehydrogenase (G6PD) expression in hepatocellular cancer. Sci Rep. 2018;8:9105.
Lee YS, Dutta A. MicroRNAs in cancer. Annu Rev Pathol. 2009;4:199–227.
Mohamed AA, Ali-Eldin ZA, Elbedewy TA, El-Serafy M, Ali-Eldin FA, AbdelAziz H. MicroRNAs and clinical implications in hepatocellular carcinoma. World J Hepatol. 2017;9:1001–7.
Zaman MS, Thamminana S, Shahryari V, et al. Inhibition of PTEN gene expression by oncogenic miR-23b-3p in renal cancer. PloS ONE. 2012;7:e50203.
Ji Q, Hao X, Zhang M, et al. MicroRNA miR-34 inhibits human pancreatic cancer tumor-initiating cells. PloS ONE. 2009;4:e6816.
Gao P, Tchernyshyov I, Chang TC, et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature. 2009;458:762–5.
Giampazolias E, Tait SW. Mitochondria and the hallmarks of cancer. FEBS J. 2016;283:803–14.
Campos-Viguri GE, Jiménez-Wences H, Peralta-Zaragoza O, et al. miR-23b as a potential tumor suppressor and its regulation by DNA methylation in cervical cancer. Infect Agents Cancer. 2015;10:42.
Jin L, Wessely O, Marcusson EG, Ivan C, Calin GA, Alahari SK. Prooncogenic factors miR-23b and miR-27b are regulated by Her2/Neu, EGF, and TNF-alpha in breast cancer. Cancer Res. 2013;73:2884–96.
Zhou R, Hu G, Gong AY, Chen XM. Binding of NF-kappaB p65 subunit to the promoter elements is involved in LPS-induced transactivation of miRNA genes in human biliary epithelial cells. Nucleic Acids Res. 2010;38:3222–32.
Bisio A, De Sanctis V, Del Vescovo V, et al. Identification of new p53 target microRNAs by bioinformatics and functional analysis. BMC Cancer. 2013;13:552.
Acknowledgements
We thank H. Nikki March, PhD, from the Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript. We also thank Dr. Mikako Ito in Nagoya University for use of the Extracellular flux analyzer system (Agilent, Santa Clara, CA, USA).
Funding
This study was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI Grant-in-Aid for Scientific Research (C) Nos. JP15H06281 and JP19K18143.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hayashi, M., Yamada, S., Kurimoto, K. et al. miR-23b-3p Plays an Oncogenic Role in Hepatocellular Carcinoma. Ann Surg Oncol 28, 3416–3426 (2021). https://doi.org/10.1245/s10434-020-09283-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-020-09283-y