Abstract
Background
Hepatitis C virus (HCV) infection produces chronic hepatitis, cirrhosis, and, ultimately, hepatocellular carcinoma (HCC). A molecular analysis of the damaged liver tissues infected with HCV may identify specific gene-expression profiles associated with a risk for liver carcinogenesis.
Methods
Forty patients with HCV-positive HCC were classified into two groups: single nodular HCC group (n = 28) and multicentric HCC group (n = 12). Using a complementary DNA microarray, we compared the gene-expression patterns of the noncancerous liver tissue specimens between the two groups. We also identified the differentially expressed genes related to multicentric recurrence in the liver remnant. We then evaluated whether a specific gene-expression profile can accurately estimate the risk for multicentric hepatocarcinogenesis.
Results
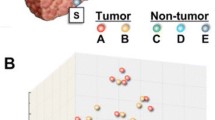
We selected the 230 differentially expressed genes in the multicentric HCC group. A hierarchical clustering analysis identified a cluster that might be closely associated with the multicentric occurrence of HCC. On the basis of the gene-expression profiling of the 36 genes commonly associated with both multicentric HCC and multicentric recurrence, we created a scoring system to estimate the risk for multicentric hepatocarcinogenesis. The prediction score of patients in the multicentric HCC group with multicentric recurrence (19.9 ± 9.2) was significantly higher (P < .05) than that in the single nodular HCC group without multicentric recurrence (−1.8 ± 12.7).
Conclusions
Specific gene-expression signatures in noncancerous liver tissue may help to accurately predict the risk for developing HCC.



Similar content being viewed by others
References
El Serag HB, Mason AC. Rising incidence of hepatocellular carcinoma in the United States. N Engl J Med 1999;340:745–50
Tsukuma H, Hiyama T, Tanaka S, et al. Risk factors for hepatocellular carcinoma among patients with chronic liver disease. N Engl J Med 1993;328:1797–801
Sherlock S. Viruses and hepatocellular carcinoma. Gut 1994;35:828–32
Tarao K, Rino Y, Ohkawa S, et al. Association between high serum alanine aminotransferase levels and more rapid development and higher rate of incidence of hepatocellular carcinoma in patients with hepatitis C virus-associated cirrhosis. Cancer 1999;86:589–95
Utsunomiya T, Matsumata T, Adachi E, et al. Limitations of current preoperative liver imaging techniques for intrahepatic metastatic nodules of hepatocellular carcinoma. Hepatology 1992;16:694–701
Shimada M, Takenaka K, Gion T, et al. Prognosis of recurrent hepatocellular carcinoma: a 10-year surgical experience in Japan. Gastroenterology 1996;111:720–6
Poon RT, Fan ST, Ng IO, et al. Different risk factors and prognosis for early and late intrahepatic recurrence after resection of hepatocellular carcinoma. Cancer 2000;89:500–7
Takenaka K, Adachi E, Nishizaki T, et al. Possible multicentric occurrence of hepatocellular carcinoma: a clinicopathological study. Hepatology 1994;19:889–94
Kumada T, Nakano S, Takeda I, et al. Patterns of recurrence after initial treatment in patients with small hepatocellular carcinoma. Hepatology 1997;25:87–92
Iizuka N, Oka M, Yamada Okabe H, et al. Oligonucleotide microarray for prediction of early intrahepatic recurrence of hepatocellular carcinoma after curative resection. Lancet 2003;361:923–9
Kurokawa Y, Matoba R, Takemasa I, et al. Molecular-based prediction of early recurrence in hepatocellular carcinoma. J Hepatol 2004;41:284–91
Kim JW, Ye Q, Forgues M, et al. Cancer-associated molecular signature in the tissue samples of patients with cirrhosis. Hepatology 2004;39:518–27
Liver Cancer Study Group of Japan. 2000. The General Rules for the Clinical and Pathological Study of Primary Liver Cancer. 4th ed. Tokyo: Kanahara Shuppan, p 32–3
Ueno S, Aoki D, Maeda T, et al. Preoperative assessment of multicentric occurrence in synchronous small and multiple hepatocellular carcinoma based on image-patterns and histological grading of non-cancerous region. Hepatol Res 2004;29:24–30
Utsunomiya T, Shimada M, Taguchi KI, et al. Clinicopathologic features and postoperative prognosis of multicentric small hepatocellular carcinoma. J Am Coll Surg 2000;190:331–5
Nishida K, Mine S, Utsunomiya T, et al. Global analysis of altered gene expressions during the process of esophageal squamous cell carcinogenesis in the rat: a study combined with a laser microdissection and a cDNA microarray. Cancer Res 2005;65:401–9
Utsunomiya T, Okamoto M, Hashimoto M, et al. A gene-expression signature can quantify the degree of hepatic fibrosis in the rat. J Hepatol 2004;41:399–406
Eisen MB, Spellman PT, Brown PO, et al. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 1998;95:14863–8
Brazma A, Hingamp P, Quackenbush J, et al. Minimum information about a microarray experiment (MIAME)—toward standards for microarray data. Nat Genet 2001;29:365–71
Colombat M, Paradis V, Bieche I, et al. Quantitative RT-PCR in cirrhotic nodules reveals gene expression changes associated with liver carcinogenesis. J Pathol 2003;201:260–7
Anders RA, Yerian LM, Tretiakova M, et al. cDNA microarray analysis of macroregenerative and dysplastic nodules in end-stage hepatitis C virus-induced cirrhosis. Am J Pathol 2003;162:991–1000
Greer P, Haigh J, Mbamalu G, et al. The Fps/Fes protein-tyrosine kinase promotes angiogenesis in transgenic mice. Mol Cell Biol 1994;14:6755–63
Orlovsky K, Theodor L, Malovani H, et al. Gamma interferon down-regulates Fer and induces its association with inactive Stat3 in colon carcinoma cells. Oncogene 2002;21:4997–5001
Larsson N, Segerman B, Howell B, et al. Op18/stathmin mediates multiple region-specific tubulin and microtubule-regulating activities. J Cell Biol 1999;146:1289–302
Neben K, Korshunov A, Benner A, et al. Microarray-based screening for molecular markers in medulloblastoma revealed STK15 as independent predictor for survival. Cancer Res 2004;64:3103–11
Vauthey JN, Walsh GL, Vlastos G, et al. Importance of field cancerisation in clinical oncology. Lancet Oncol 2000;1:15–6
Acknowledgments
Supported by Grants-in-Aid for Scientific Research (17109013, 17591411, 17591413, 17015032, and 16390381), the Japan Society for the Promotion of Science, and a Health and Labor Sciences Research Grant on Hepatitis and BSE (14230801; the Ministry of Health, Labor and Welfare of Japan). Also supported by CREST, JST, Uehara Memorial Foundation, Yasuda Medical Research Foundation, Japanese Foundation for Multidisciplinary Treatment of Cancer, and Princess Takamatsu Cancer Research Fund.
Author information
Authors and Affiliations
Corresponding author
Additional information
M. Okamoto and T. Utsunomiya contributed equally to this study.
Rights and permissions
About this article
Cite this article
Okamoto, M., Utsunomiya, T., Wakiyama, S. et al. Specific Gene-Expression Profiles of Noncancerous Liver Tissue Predict the Risk for Multicentric Occurrence of Hepatocellular Carcinoma in Hepatitis C Virus–Positive Patients. Ann Surg Oncol 13, 947–954 (2006). https://doi.org/10.1245/ASO.2006.07.018
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1245/ASO.2006.07.018