Abstract
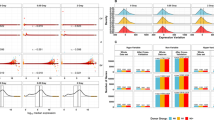
The level of spontaneous and radiation-induced DNA damage varies depending on genetic and environmental factors in human somatic cells. This variation may be associated with transcriptional changes in cells, allowing the use of gene expression levels as markers of individual sensitivity to mutagenic effects. This study aimed to identify and characterize differentially expressed genes (DEGs) in lymphocytes of individuals with various frequencies of endogenous γH2AX foci and radiation-induced micronuclei (n = 37). The low-focus group was characterized by 0.18 ± 0.02 endogenous γH2AX foci per cell and a 155.78 ± 47.19‰ radiation-induced micronucleus frequency. The high-focus group was characterized by 0.49 ± 0.07 foci/cell and a 78.44 ± 33.21‰ micronucleus frequency. Seven DEGs (ENST00000424415, CRNDE, ADAMTS1, ENST00000424084, EIF2A, PNPLA5, and FRG2C) (FDR < 0.2) were identified by gene expression analysis with microarrays. As the extracellular matrix metalloproteinase, ADAMTS1 is able to activate the latent form of TGFβ, and TGFβ is involved in radiation-induced cellular response; the effects of ADAMTS1 knockout and overexpression on the gene expression profile were further validated in adherent HeLa cells. Twenty-nine of 160 identified DEGs are involved in apoptosis, DNA DSB repair, G2/M cell cycle transition, and the TGFβ signaling pathway. Thus, ADAMTS1 may be useful as a potential target for antitumor therapy.


Similar content being viewed by others
REFERENCES
Sokolov, M. and Neumann, R., Global gene expression alterations as a crucial constituent of human cell response to low doses of ionizing radiation exposure, Int. J. Mol. Sci., 2015, vol. 17, no. 1, p. 55. https://doi.org/10.3390/ijms17010055
Foray, N., Bourguignon, M., and Hamada, N., Individual response to ionizing radiation, Mutat. Res., Rev. Mutat. Res., 2016, vol. 770, pp. 369—386. https://doi.org/10.1016/j.mrrev.2016.09.001
Rajaraman, P., Hauptmann, M., Bouffler, S., and Wojcik, A., Human individual radiation sensitivity and prospects for prediction, Ann. ICRP, 2018, vol. 47, nos. 3—4, pp. 126—141. https://doi.org/10.1177/0146645318764091
Andreassen, C.N., Schack, L.M.H., Laursen, L.V., and Alsner, J., Radiogenomics—current status, challenges and future directions, Cancer Lett., 2016, vol. 382, no. 1, pp. 127—136. https://doi.org/10.1016/j.canlet.2016.01.035
Wang, T.M., Shen, G.P., Chen, M.Y., et al., Genome-wide association study of susceptibility loci for radiation-induced brain injury, J. Natl. Cancer Inst., 2019, vol. 111, no. 6, pp. 620—628. https://doi.org/10.1093/jnci/djy150
Yang, D.W., Wang, T.M., Zhang, J.B., et al., Genome-wide association study identifies genetic susceptibility loci and pathways of radiation-induced acute oral mucositis, J. Transl. Med., 2020, vol. 18, no. 1, pp. 1—12. https://doi.org/10.1186/s12967-020-02390-0
Zheng, S. and Tao, W., Identification of novel transcriptome signature as a potential prognostic biomarker for anti-angiogenic therapy in glioblastoma multiforme, Cancers, 2021, vol. 13, no. 5, p. 1013. https://doi.org/10.3390/cancers13051013
Bhosle, S.M., Huilgol, N.G., and Mishra, K.P., Apoptotic index as predictive marker for radiosensitivity of cervical carcinoma: evaluation of membrane fluidity, biochemical parameters and apoptosis after the first dose of fractionated radiotherapy to patients, Cancer Detect. Prev., 2005, vol. 29, no. 4, pp. 369—375. https://doi.org/10.1016/j.cdp.2005.05.002
Azria, D., Riou, O., Castan, F., et al., Radiation-induced CD8 T-lymphocyte apoptosis as a predictor of breast fibrosis after radiotherapy: results of the prospective multicenter French Trial, EBioMedicine, 2015, vol. 2, no. 13, pp. 1965—1973. https://doi.org/10.1016/j.ebiom.2015.10.024
Anderson, R.M., Cytogenetic biomarkers of radiation exposure, Clin. Oncol., 2019, vol. 31, no. 5, pp. 311—318. https://doi.org/10.1016/j.clon.2019.02.009
Bucher, M., Endesfelder, D., Roessler, U., et al., Analysis of chromosomal aberrations and γH2A.X foci to identify radiation-sensitive ataxia—telangiectasia patients, Mutat. Res., Genet. Toxicol. Environ. Mutagen., 2021, vol. 861, p. 503301. https://doi.org/10.1016/j.mrgentox.2020.503301
Guogytė, K., Plieskienė, A., Ladygienė, R., et al., Assessment of correlation between chromosomal radiosensitivity of peripheral blood lymphocytes after in vitro irradiation and normal tissue side effects for cancer patients undergoing radiotherapy, Genome Integr., 2017, vol. 8, p. 1. https://doi.org/10.4103/2041-9414.198907
Redon, C.E., Dickey, J.S., Bonner, W.M., and Sedelnikova, O.A., γ-H2AX as a biomarker of DNA damage induced by ionizing radiation in human peripheral blood lymphocytes and artificial skin, Adv. Space Res., 2009, vol. 43, no. 8, pp. 1171—1178. https://doi.org/10.1016/j.asr.2008.10.011
Markova, E., Vasilyev, S., and Belyaev, I., 53BP1 foci as a marker of tumor cell radiosensitivity, Neoplasma, 2015, vol. 62, no. 5, pp. 770—776. https://doi.org/10.4149/neo_2015_092
Belyaev, I.Y., Radiation-induced DNA repair foci: spatio-temporal aspects of formation, application for assessment of radio-sensitivity and biological dosimetry, Mutat. Res., Rev. Mutat. Res., 2010, vol. 704, nos. 1—3, pp. 132—141. https://doi.org/10.1016/j.mrrev.2010.01.011
Sedelnikova, O.A., Horikawa, I., Zimonjic, D.B., et al., Senescing human cells and ageing mice accumulate DNA lesions with unrepairable double-strand breaks, Nat. Cell Biol., 2004, vol. 6, no. 2, pp. 168—170. https://doi.org/10.1038/ncb1095
Han, J., Hendzel, M.J., Allalunis-Turner, J., Quantitative analysis reveals asynchronous and more than DSB-associated histone H2AX phosphorylation after exposure to ionizing radiation, Radiat. Res., 2006, vol. 165, no. 3, pp. 283—292. https://doi.org/10.1667/rr3516.1
Kato, T.A., Okayasu, R., Bedford, J.S., Comparison of the induction and disappearance of DNA double strand breaks and γ-H2AX foci after irradiation of chromosomes in G1-phase or in condensed metaphase cells, Mutat. Res. Fundam. Mol. Mech. Mutagen., 2008, vol. 639, nos. 1—2, pp. 108—112. https://doi.org/10.1016/j.mrfmmm.2007.11.006
Nakamura, A.J., Redon, C.E., Bonner, W.M., and Sedelnikova, O.A., Telomere-dependent and telomere-independent origins of endogenous DNA damage in tumor cells, Aging, 2009, vol. 1, pp. 212—218. https://doi.org/10.18632/aging.100019
Fumagalli, M., Rossiello, F., Clerici, M., et al., Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation, Nat. Cell Biol., 2012, vol. 14, no. 4, pp. 355—365. https://doi.org/10.1038/ncb2466
Bernadotte, A., Mikhelson, V.M., and Spivak, I.M., Markers of cellular senescence: telomere shortening as a marker of cellular senescence, Aging, 2016, vol. 8, no. 1, pp. 3—11. https://doi.org/10.18632/aging.100871
Georgoulis, A., Vorgias, C.E., Chrousos, G.P., and Rogakou, E.P., Genome instability and γH2AX, Int. J. Mol. Sci., 2017, no. 9, vol. 18, pp. 1979—1989. https://doi.org/10.3390/ijms18091979
Vasilev, S.A., Velichevskaya, A.I., Vishnevskaya, T.V., et al., Background level of γH2AX foci in human cells as a factor of individual radiosensitivity, Radiats. Biol., Radioekol., 2015, vol. 55, no. 4, pp. 402—410. https://doi.org/10.7868/S0869803115040128
Martin, O.A., Ivashkevich, A., Choo, S., et al., Statistical analysis of kinetics, distribution and co-localisation of DNA repair foci in irradiated cells: cell cycle effect and implications for prediction of radiosensitivity, DNA Repair., 2013, vol. 12, no. 10, pp. 844—855. https://doi.org/10.1016/j.dnarep.2013.07.002
Muller, S., Neusser, M., Kohler, D., and Cremer, M., Preparation of complex DNA probe sets for 3D FISH with up to six different fluorochromes, Cold Spring Harb. Protoc., 2007, vol. 2007, no. 5. https://doi.org/10.1101/pdb.prot4730
Melnikov, A.A., Vasilyev, S.A., Musabaeva, L.I., et al., Indication of cytogenetic abnormalities in peripheral blood lymphocytes of patients with malignant neoplasms under fast neutron therapy, Tyumen. Med. Zh., 2012, no. 4, pp. 76—78.
Savchenko, R.R., Murashkina, A.A., Fishman, V.S., et al., Effect of ADAMTS1 differential expression on the radiation-induced response of HeLa cell line, Russ. J. Genet., 2021, vol. 57, no. 7, pp. 856—862. https://doi.org/10.1134/S1022795421070127
Edie, S., Zaghloul, N.A., Leitch, C.C., et al., Survey of human chromosome 21 gene expression effects on early development in Danio rerio, G3 (Bethesda, MD), 2018, vol. 8, no. 7, pp. 2215—2223. https://doi.org/10.1534/g3.118.200144
GEO accession viewer. https://www.ncbi.nlm.nih.gov/ geo/query/acc.cgi?acc=GSE97000. Accessed May 19, 2021.
Tilton, S.C., Markillie, L.M., Hays, S., et al., Identification of differential gene expression patterns after acute exposure to high and low doses of low-LET ionizing radiation in a reconstituted human skin tissue, Radiat. Res., 2016, vol. 186, no. 5, pp. 531—538. https://doi.org/10.1667/rr14471.1
Cilensek, Z.M., Yehiely, F., Kular, R.K., and Deiss, L.P., A member of the GAGE family of tumor antigens is an anti-apoptotic gene that confers resistance to Fas/CD95/APO-1, interferon-gamma, taxol and gamma-irradiation, Cancer Biol. Ther., 2002, vol. 1, no. 4, pp. 379—386. https://doi.org/10.4161/cbt.1.4.11
Herbert, K., Binet, R., Lambert, J.P., et al., BRN2 suppresses apoptosis, reprograms DNA damage repair, and is associated with a high somatic mutation burden in melanoma, Genes Dev., 2019, vol. 33, nos. 5—6, pp. 310—332. https://doi.org/10.1101/gad.314633.118
You, Y., Wen, R., Pathak, R., et al., Latexin sensitizes leukemogenic cells to gamma-irradiation-induced cell-cycle arrest and cell death through Rps3 pathway, Cell Death Dis., 2014, vol. 5, no. 10. e1493. https://doi.org/10.1038/cddis.2014.443
Tanaka, Y., Imamura, J., Kanai, F., et al., Runx3 interacts with DNA repair protein Ku70, Exp. Cell Res., 2007, vol. 313, no. 15, pp. 3251—3260. https://doi.org/10.1016/j.yexcr.2007.06.012
Jiang, J., Han, P., Qian, J., et al., Knockdown of ALPK2 blocks development and progression of renal cell carcinoma, Exp. Cell Res., 2020, vol. 392, no. 2, p. 112029. https://doi.org/10.1016/j.yexcr.2020.112029
Eckers, J.C., Kalen, A.L., Xiao, W., et al., Selenoprotein P inhibits radiation-induced late reactive oxygen species accumulation and normal cell injury, Int. J. Radiat. Oncol. Biol. Phys., 2013, vol. 87, no. 3, pp. 619—625. https://doi.org/10.1016/j.ijrobp.2013.06.2063
Graham, L.D., Pedersen, S.K., Brown, G.S., et al., Colorectal neoplasia differentially expressed (CRNDE), a novel gene with elevated expression in colorectal adenomas and adenocarcinomas, Genes Cancer, 2011, vol. 2, no. 8, pp. 829—840. https://doi.org/10.1177/1947601911431081
Ellis, B.C., Molloy, P.L., and Graham, L.D., CRNDE: a long non-coding RNA involved in CanceR, neurobiology, and development, Front. Genet., 2012, vol. 3, p. 270. https://doi.org/10.3389/fgene.2012.00270
Zhang, X., Sun, S., Pu, J.K.S., et al., Long non-coding RNA expression profiles predict clinical phenotypes in glioma, Neurobiol. Dis., 2012, vol. 48, no. 1, pp. 1—8. https://doi.org/10.1016/j.nbd.2012.06.004
Szafron, L.M., Balcerak, A., Grzybowska. E.A., et al., The novel gene CRNDE encodes a nuclear peptide (CRNDEP) which is overexpressed in highly proliferating tissues, PLoS One, 2015, vol. 10, no. 5. e0127475. https://doi.org/10.1371/journal.pone.0127475
Han, P., Li, J.W., Zhang, B.M., et al., The lncRNA CRNDE promotes colorectal cancer cell proliferation and chemoresistance via miR-181a-5p-mediated regulation of Wnt/β-catenin signaling, Mol. Cancer, 2017, vol. 16, no. 1, pp. 1—13. https://doi.org/10.1186/s12943-017-0583-1
Liu, C., Hou, J., Shan, F., et al., Long non-coding RNA CRNDE promotes colorectal carcinoma cell progression and paclitaxel resistance by regulating miR-126-5p/ATAD2 axis, OncoTargets Ther., 2020, vol. 13, pp. 4931—4942. https://doi.org/10.2147/OTT.S237580
Ellis, B.C., Graham, L.D., and Molloy, P.L., CRNDE, a long non-coding RNA responsive to insulin/IGF signaling, regulates genes involved in central metabolism, Biochim. Biophys. Acta, Mol. Cell Res., 2014, vol. 1843, no. 2, pp. 372—386. https://doi.org/10.1016/j.bbamcr.2013.10.016
Wu, D., Han, B., Guo, L., and Fan, Z., Molecular mechanisms associated with breast cancer based on integrated gene expression profiling by bioinformatics analysis, J. Obstet. Gynaecol., 2016, vol. 36, no. 5, pp. 615—621. https://doi.org/10.3109/01443615.2015.1127902
Ewan, K.B., Henshall-Powell, R.L., Ravani, S.A., et al., Transforming growth factor-B1 mediates cellular response to DNA damage in situ, Cancer Res. 2002, vol. 62, no. 20, pp. 5627—5631.
Kirshner, J., Jobling, M.F., Pajares, M.J., et al., Inhibition of transforming growth factor-β1 signaling attenuates ataxia telangiectasia mutated activity in response to genotoxic stress, Cancer Res., 2006, vol. 66, no. 22, pp. 10861—10869. https://doi.org/10.1158/0008-5472.can-06-2565
Wiegman, E.M., Blaese, M.A., Loeffler, H., et al., TGFβ-1 dependent fast stimulation of ATM and p53 phosphorylation following exposure to ionizing radiation does not involve TGFβ-receptor I signaling, Radiother. Oncol., 2007, vol. 83, no. 3, pp. 289—295. https://doi.org/10.1016/j.radonc.2007.05.013
Funding
The present study was supported by the Russian Foundation for Basic Research, projects no. 14-04-31867 (experiments conducted with PBMCs of healthy individuals) and no. 19-34-90143 (experiments conducted with cell lines).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The authors declare no conflict of interest.
Statement of compliance with standards of research involving humans as subjects. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants involved in the study.
Additional information
Translated by A. Kazantseva
Rights and permissions
About this article
Cite this article
Vasilyev, S.A., Savchenko, R.R., Belenko, A.A. et al. ADAMTS1 Is Differentially Expressed in Human Lymphocytes with Various Frequencies of Endogenous γH2AX Foci and Radiation-Induced Micronuclei. Russ J Genet 58, 1235–1244 (2022). https://doi.org/10.1134/S102279542210012X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S102279542210012X