-
PDF
- Split View
-
Views
-
Cite
Cite
Dawn E. Hall, Philipp Zerbe, Sharon Jancsik, Alfonso Lara Quesada, Harpreet Dullat, Lina L. Madilao, Macaire Yuen, Jörg Bohlmann, Evolution of Conifer Diterpene Synthases: Diterpene Resin Acid Biosynthesis in Lodgepole Pine and Jack Pine Involves Monofunctional and Bifunctional Diterpene Synthases , Plant Physiology, Volume 161, Issue 2, February 2013, Pages 600–616, https://doi.org/10.1104/pp.112.208546
Close - Share Icon Share
Abstract
Diterpene resin acids (DRAs) are major components of pine (Pinus spp.) oleoresin. They play critical roles in conifer defense against insects and pathogens and as a renewable resource for industrial bioproducts. The core structures of DRAs are formed in secondary (i.e. specialized) metabolism via cycloisomerization of geranylgeranyl diphosphate (GGPP) by diterpene synthases (diTPSs). Previously described gymnosperm diTPSs of DRA biosynthesis are bifunctional enzymes that catalyze the initial bicyclization of GGPP followed by rearrangement of a (+)-copalyl diphosphate intermediate at two discrete class II and class I active sites. In contrast, similar diterpenes of gibberellin primary (i.e. general) metabolism are produced by the consecutive activity of two monofunctional class II and class I diTPSs. Using high-throughput transcriptome sequencing, we discovered 11 diTPS from jack pine (Pinus banksiana) and lodgepole pine (Pinus contorta). Three of these were orthologous to known conifer bifunctional levopimaradiene/abietadiene synthases. Surprisingly, two sets of orthologous PbdiTPSs and PcdiTPSs were monofunctional class I enzymes that lacked functional class II active sites and converted (+)-copalyl diphosphate, but not GGPP, into isopimaradiene and pimaradiene as major products. Diterpene profiles and transcriptome sequences of lodgepole pine and jack pine are consistent with roles for these diTPSs in DRA biosynthesis. The monofunctional class I diTPSs of DRA biosynthesis form a new clade within the gymnosperm-specific TPS-d3 subfamily that evolved from bifunctional diTPS rather than monofunctional enzymes (TPS-c and TPS-e) of gibberellin metabolism. Homology modeling suggested alterations in the class I active site that may have contributed to their functional specialization relative to other conifer diTPSs.
Conifer trees, including lodgepole pine (Pinus contorta) and jack pine (Pinus banksiana), produce complex mixtures of mono-, sesqui-, and diterpenoid specialized (i.e. secondary) metabolites, most prominently in the form of oleoresin, that can act as a physical and chemical defense against insect and pathogen attack (Phillips and Croteau, 1999; Keeling and Bohlmann, 2006a, 2006b; Zulak and Bohlmann, 2010; Boone et al., 2011). These oleoresin terpenoids also serve as a large-volume, renewable resource for industrial bioproducts, including solvents, flavors, fragrances, coatings, and resins (Langenheim, 2003; Bohlmann and Keeling, 2008; Bohlmann, 2011; Zerbe et al., 2012a). Oleoresin terpenoids are biosynthesized by a large family of catalytically diverse terpene synthases (TPSs) that contribute to a wide array of different compounds that can be produced by a single tree (Martin et al., 2004; Zulak et al., 2009; Keeling et al., 2011a). The many mono-, sesqui-, and diterpene synthases (diTPSs) of conifer specialized metabolism form the gymnosperm-specific TPS-d subfamily (Bohlmann et al., 1998a; Martin et al., 2004; Chen et al., 2011; Keeling et al., 2011a). The functional diversity of these conifer diTPSs appears to have evolved through events of repeated gene duplication and further sub- and neofunctionalization, leading to an expansion of the TPS-d multigene family (Martin et al., 2004; Keeling et al., 2008; Chen et al., 2011) that is the central player behind the chemical complexity of conifer specialized diterpenes. In addition, conifer diTPSs of general (i.e. primary) GA metabolism belong to the TPS-c and TPS-e/f subfamilies, which also include orthologous genes of angiosperms (Keeling et al., 2010; Chen et al., 2011).
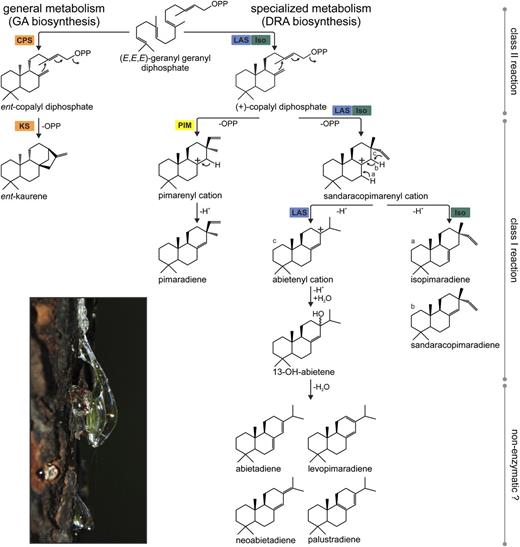
In gymnosperms and angiosperms, diterpene metabolites are formed through the multistep cycloisomerization of geranylgeranyl diphosphate (GGPP) catalyzed by diTPSs (Fig. 1). Despite their functional diversity, known diTPSs are structurally conserved with variations in three common α-helical domains, α, β, and γ, and associated active site motifs (Köksal et al., 2011a, 2011b; Zhou et al., 2012). diTPSs involved in general metabolism of GA biosynthesis in angiosperms and gymnosperms, as well as those involved in the biosynthesis of the large class of labdane-related specialized diterpenes in angiosperms, are monofunctional class II and class I enzymes (Keeling et al., 2010; Peters, 2010). The class II diTPSs contain a DxDD signature motif in the γβ-domain relevant for catalyzing the protonation-initiated cyclization of GGPP to a bicyclic diphosphate intermediate (Harris et al., 2005; Prisic and Peters, 2007; Xu et al., 2007a; Gao et al., 2009; Toyomasu et al., 2009; Falara et al., 2010; Keeling et al., 2010; Caniard et al., 2012; Sallaud et al., 2012). Class I diTPSs, which harbor DDxxD and NSE/DTE functional motifs in the α-domain, then catalyze the ionization of the diphosphate ester and subsequent rearrangement reactions (Xu et al., 2007a; Gao et al., 2009; Keeling et al., 2010; Caniard et al., 2012; Sallaud et al., 2012). In contrast, the archetypical diTPSs of nonvascular plants, such as Physcomitrella patens or Jungermannia subulata (Hayashi et al., 2006; Kawaide et al., 2011), the specialized diTPSs of the lycophyte Selaginella moellendorffii (Mafu et al., 2011; Sugai et al., 2011), and the gymnosperm diTPSs involved in diterpene resin acid (DRA) metabolism (Vogel et al., 1996; Peters et al., 2000; Schepmann et al., 2001; Martin et al., 2004; Ro and Bohlmann, 2006; Keeling et al., 2011a, 2011b; Zerbe et al., 2012a), are bifunctional class I/II enzymes that contain both functional active sites.
diTPS-catalyzed biosynthesis of general and specialized diterpenes in conifers. GA biosynthesis invariably derives from ent-CPP and ent-kaurene as intermediates. In contrast, the formation of DRAs proceeds via (+)-CPP and the rearrangement of various distinct carbocations to afford the mixture of known diterpenes in the oleoresin of coniferous trees (photo). Previously described or proposed diTPS catalytic functions involved in these individual pathways are highlighted. CPS, Copalyl diphosphate synthase; KS, kaurene synthase.
The formation of diterpenes in conifer DRA biosynthesis proceeds via the initial bicyclization of GGPP into (+)-copalyl diphosphate [(+)-CPP] at the class II active site of bifunctional diTPSs. The (+)-CPP intermediate then translocates to the class I active site and undergoes secondary cyclization and further rearrangements via intermediate carbocations (Peters et al., 2001). Well-characterized class I/II diTPSs of conifers include the isopimaradiene synthase-type (ISO) and levopimaradiene/abietadiene synthase-type (LAS) enzymes, which differ in the reactions catalyzed in the respective class I active sites (Martin et al., 2004; Keeling et al., 2008). ISO enzymes catalyze the direct deprotonation of the secondary sandaracopimarenyl cation to form isopimaradiene and minor quantities of sandaracopimaradiene (Fig. 1). The LAS-catalyzed reaction involves additional rearrangement reactions and water capture at C13, resulting in 13-hydroxy-8(14)-abietene as the initial product, which in vitro (and perhaps in vivo) readily dehydrates to afford abietadiene, levopimaradiene, palustradiene, and neoabietadiene (Keeling et al., 2011b). These different catalytic specificities of known gymnosperm diTPSs are predominantly determined by the conformation and amino acid composition of the class I active site. Targeted mutagenesis studies have shown that as little as a single residue substitution is sufficient to alter the product profile of these enzymes (Peters and Croteau, 2002a, 2002b; Wilderman and Peters, 2007; Keeling et al., 2008; Leonard et al., 2010; Zerbe et al., 2012b). To complete the biosynthesis of DRAs, stepwise oxidations at the C18 position of the products of LAS and ISO are catalyzed by cytochrome P450 enzymes of the CYP720B family, yielding the corresponding diterpene alcohols, aldehydes, and resin acids (Ro et al., 2005; Hamberger et al., 2011).
Notably, the recently identified class I/II cis-abienol synthase from balsam fir (Abies balsamea; Zerbe et al., 2012a) as well as the class I taxadiene synthases from species of yew (Taxus spp.; Wildung and Croteau, 1996; Köksal et al., 2011a) employ distinct reaction mechanisms. cis-Abienol synthase catalyzes the formation of bicyclic diterpene alcohol cis-abienol via a labda-13-en-8-ol diphosphate intermediate, whereas taxadiene synthases facilitate the direct conversion of GGPP into the macrocyclic diterpene taxadiene.
Lodgepole pine is the main host species of the current large-scale outbreak of the mountain pine beetle (MPB; Dendroctonus ponderosae) and its associated pathogenic fungi, including the ascomycete Grosmannia clavigera (DiGuistini et al., 2011; Bohlmann, 2012; Keeling et al., 2012). In recent years, the MPB epidemic infested more than 15 million hectares of pine forest in western North America (Safranyik et al., 2010). A host range expansion of MPB into jack pine was recently documented as the epidemic expanded geographically eastwards across the Rocky Mountains barrier (Cullingham et al., 2011). While the diterpene profile of lodgepole pine saplings has been previously determined to contain the DRAs levopimaric acid, palustric acid, isopimaric acid, neoabietic acid, abietic acid, dehydroabietic acid, sandaracopimaric acid, and pimaric acid (Lewinsohn et al., 1993), little is known about the diterpenoid profile of jack pine. The effect of the jack pine and lodgepole pine oleoresin diterpenoids on MPB and its associated fungi is not known. However, in lodgepole pine, high levels of abietic acid and dehydroabietic acid were detected in the sapwood and heartwood following MPB attack, with pimaric acid, sandaracopimaric acid, isopimaric acid, and levopimaric acid/palustric acid also detected at low levels (Shrimpton, 1973). In a different bark beetle system, the DRAs abietic acid and isopimaric acid strongly inhibited spore germination, and abietic acid also strongly inhibited mycelial growth of Ophisotoma ips, a fungal symbiont of the bark beetle Ips pini (Kopper et al., 2005).
Despite the economic and ecological importance of pines and the importance of oleoresin diterpenes in pine defense and as bioproducts, no diTPSs have been identified in jack pine and lodgepole pine. In other species of pine, to the best of our knowledge, only a single bifunctional diTPS (PtLAS), which produces abietadiene, neoabietadiene, levopimaradiene, and palustradiene, has been characterized from loblolly pine (Pinus taeda; Ro and Bohlmann, 2006). Here, we report the identification of 11, and the functional characterization of seven, diTPSs from jack pine and lodgepole pine, including the discovery of four monofunctional diTPSs of DRA biosynthesis. The recombinant monofunctional diTPSs produced either pimaradiene or a combination of isopimaradiene and sandaracopimaradiene in vitro. Additionally, we characterized the bifunctional jack pine and lodgepole pine LAS-type diTPSs and demonstrate that these LAS enzymes produce a tertiary alcohol as the primary diterpene product. Assays in which the monofunctional pimaradiene and isopimaradiene synthases were coupled with the bifunctional jack pine and lodgepole pine LAS enzymes demonstrated that the class I active sites of monofunctional proteins can use the (+)-CPP intermediate produced by the class II active site of the bifunctional enzymes. Analysis of high-throughput sequence data indicated distinct expression levels of mono- and bifunctional diTPSs in the trees, and metabolite profiling was consistent with an in vivo role for these proteins in DRA biosynthesis.
RESULTS
Transcriptome Mining and Cloning of Jack Pine and Lodgepole Pine diTPSs
To facilitate the discovery of diTPSs in lodgepole pine and jack pine, we used transcriptome sequence resources developed by a combination of Sanger, 454, and Illumina sequencing of complementary DNA (cDNA) libraries made from sapling stem tissues (National Center for Biotechnology Information [NCBI] Short Read Archive accession number SRP009894). De novo transcriptome assemblies resulted in the identification of approximately 30,000 unigenes for each of the two pine species (D.E. Hall, M. Yuen, and J. Bohlmann, unpublished data). A TBLASTN search of these transcriptome assemblies against an in-house annotated database of 102 conifer TPSs (e.g. Keeling et al., 2011a) identified two full-length (FL) candidate diTPS sequences from each of jack pine (PbLAS1 and PbmISO1) and lodgepole pine (PcLAS1 and PcLAS2) as well as four partial diTPS-like sequences from jack pine and eight partial diTPS candidate genes from lodgepole pine. Based on the partial sequences, 5′ RACE was used to obtain two additional FL candidate diTPS sequences from jack pine (PbmPIM1 and PbmdiTPS1). Homology-based cloning led to the cloning of five additional FL diTPS candidates from lodgepole pine (PcmISO1, PcmPIM1, PcmdiTPS1, PcmdiTPS2, and PcmdiTPS3), for a total of four FL jack pine diTPS and seven FL lodgepole pine diTPS candidates (Supplemental Table S1). An overall length of 850 to 880 amino acids, along with a conserved γβα-tridomain architecture and presence of the characteristic KR(E/D)x6W motif downstream of the putative plastidial transit peptide, suggested that these 11 candidate genes encode for diTPSs, as opposed to hemi-, mono-, or sesqui-TPSs.
Sequence Analysis Suggests the Presence of Bifunctional Class I/II and Monofunctional Class I diTPSs in Jack Pine and Lodgepole Pine
Three of the FL diTPS-like sequences (PbLAS1, PcLAS1, and PcLAS2) showed 98% to 99% amino acid sequence identity to each other and to a previously characterized loblolly pine LAS (PtLAS; Ro and Bohlmann, 2006). Notably, PcLAS1 had a 21-bp insertion, which was not present in either PcLAS2 or PtLAS1. These sequences contained the class I and class II active site functional motifs (Fig. 2; Supplemental Fig. S1), suggesting that they were bifunctional class I/II diTPSs, resembling the known conifer diTPSs of DRA biosynthesis.
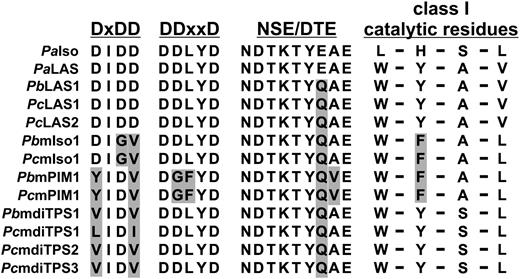
Protein sequence alignment of class II and class I diTPS signature motifs and select catalytic residues of conifer diTPSs. Protein sequence alignment of class II diTPS (DxDD) and class I diTPS (DDxxD, NSE/DTE) signature motifs, highlighting the loss of conserved Asp residues in the DxDD motif of all monofunctional diTPS candidates from jack pine and lodgepole pine. Variations of select catalytic residues that determine product specificity in P. abies LAS and ISO (Keeling et al., 2008) are also illustrated. Amino acids that differ from the corresponding residues present in P. abies LAS and ISO are shaded in gray. A complete alignment of the respective FL sequences is depicted in Supplemental Figure S1. PbmdiTPS1, P. banksiana monofunctional diTPS1; PcmdiTPS1, P. contorta monofunctional diTPS1; PcmdiTPS2, P. contorta monofunctional diTPS2; PcmdiTPS3, P. contorta monofunctional diTPS3; PbmPIM1, P. banksiana monofunctional pimaradiene synthase1; PcmPIM1, P. contorta monofunctional pimaradiene synthase1; PbmISO1, P. banksiana monofunctional isopimaradiene synthase1; PcmISO1, P. contorta monofunctional isopimaradiene synthase1; PcLAS2, P. contorta levopimaradiene/abietadiene synthase2; PcLAS1, P. contorta levopimaradiene/abietadiene synthase1; PbLAS2, P. banksiana levopimaradiene/abietadiene synthase1.
Unexpectedly, the eight remaining FL putative diTPS sequences contained only the class I signature motifs (NSE/DTE, DDxxD), but lacked either the conserved middle Asp residue (PcmISO1 and PbmISO1) or the first and last Asp residues (PcmdiTPS and PbmdiTPS) of the DxDD motif, which were previously shown to be critical for class II catalysis (Peters and Croteau, 2002b; Fig. 2; Supplemental Fig. S1). These eight sequences showed 66% to 73% amino acid identity to jack pine PbLAS1, lodgepole pine PcLAS1 and PcLAS2, and functionally characterized Norway spruce (Picea abies) PaISO and PaLAS of DRA specialized metabolism (Martin et al., 2004). Although representing putative monofunctional diTPSs, the eight sequences only showed 33% to 34% protein sequence identity to the monofunctional white spruce (Picea glauca) class II ent-copalyl diphosphate synthase (PgECPS) and class I ent-kaurene synthase (PgEKS) of GA metabolism (Keeling et al., 2010), suggesting roles in specialized as opposed to general metabolism. With 99% amino acid sequence identity to each other, PcmISO1 and PbmISO1, and likewise PcmPIM1 and PbmPIM1, presumably represent two pairs of orthologous genes from jack pine and lodgepole pine. The remaining class I diTPS candidates (PcmdiTPS1, PcmdiTPS2, PcmdiTPS3, and PbmdiTPS1), though highly similar among each other (97% to 98% protein sequence identity), showed a lower identity of 71% to 75% to the other pine diTPS candidates (Supplemental Table S1).
Characterization of Bifunctional PcLAS1, PcLAS2, and PbLAS1
In vitro enzyme assays using affinity-purified recombinant PcLAS1, PcLAS2, and PbLAS1 and C10, C15, or C20 prenyl diphosphate substrates were performed to substantiate functional annotations. Using liquid chromatography-mass spectrometry (LC-MS), the major diterpene products of PcLAS1, PcLAS2, and PbLAS1 in assays with GGPP (C20) were identified as epimers of the diterpene tertiary alcohol 13-hydroxy-8(14)-abietene, by comparison with retention times and mass spectra of the products of P. abies PaLAS (Keeling et al., 2011b). Gas chromatography-mass spectrometry (GC-MS) analysis, which causes dehydration of 13-hydroxy-8(14)-abietene (Keeling et al., 2011b), identified the diterpene olefins abietadiene, levopimaradiene, and neoabietadiene as the three major products in direct comparison to the authentic compounds (Supplemental Table S2), consistent with the three principal products of the previously characterized PtLAS, which had been analyzed exclusively by GC-MS (Ro and Bohlmann, 2006). Assays of PcLAS1, PcLAS2, and PbLAS1 with farnesyl diphosphate (FPP; C15) did not yield any products. PcLAS1 produced trace amounts of the acyclic monoterpenes myrcene, linalool, and geraniol with geranyl diphosphate (GPP; C10) or neryl diphosphate (C10); and PbLAS1 produced trace amounts of myrcene with GPP. Enzyme assays with the remaining eight jack pine and lodgepole pine diTPS candidates did not yield any products when incubated with GPP, FPP, or GGPP, except that both PcmISO1 and PbmISO1 produced trace amounts of myrcene, ocimene, linalool, and geraniol with GPP as substrate. These results established PcLAS1, PcLAS2, and PbLAS1 as bifunctional class I/II diTPSs, which converted GGPP into the diterpene tertiary alcohol 13-hydroxy-8(14)-abietene as the primary product. Conversely, the remaining eight candidate diTPSs were found to be nonfunctional in the conversion of GGPP, consistent with the lack of a catalytic DxDD motif in the class II active site.
Coupled Enzyme Assays Identify Monofunctional Conifer diTPSs Involved in Specialized Metabolism
The lack of an intact DxDD motif in the class II active site of PcmPIM1, PbmPIM1, PcmISO1, PcmISO1, PbmdiTPS1, PcmdiTPS1, PcmdiTPS2, and PcmdiTPS3 suggested that these proteins may require copalyl diphosphate (CPP), but not GGPP, as a substrate. As CPP is not commercially available, we employed coupled assays with different CPP synthases to test if any of these eight enzymes were active as monofunctional class I diTPS. Specifically, we used the maize (Zea mays) AN2 protein (Harris et al., 2005) to produce ent-CPP and a protein variant of PaLAS (PaLAS:D611A; Zerbe et al., 2012a) to produce (+)-CPP with GGPP as substrate. Following quantitative conversion of GGPP to CPP with these enzymes, conversion of CPP was then tested by addition of the candidate class I pine diTPS proteins.
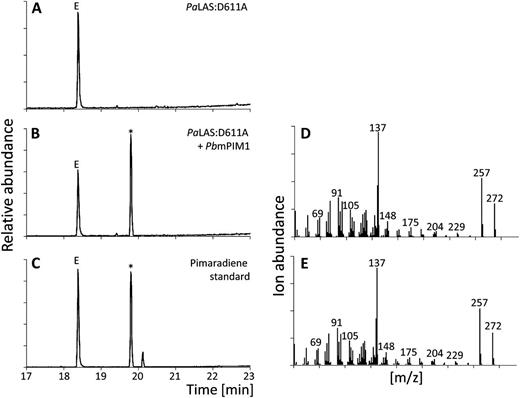
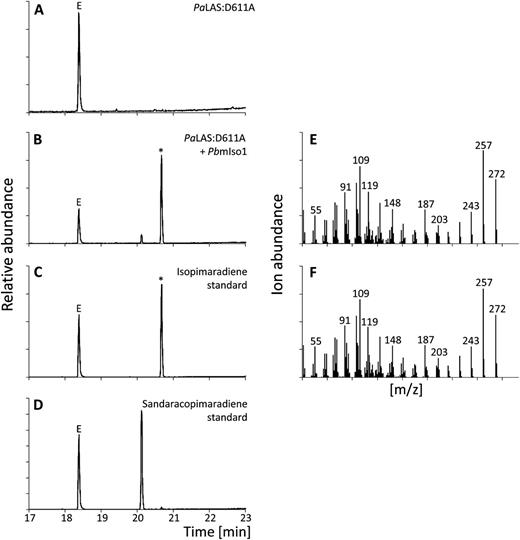
When PcmPIM1 and PbmPIM1 were incubated with PaLAS:D611A and GGPP [i.e. providing (+)-CPP as a substrate], we identified diterpene product profiles of 99% and 95% pimaradiene, respectively (Fig. 3, Supplemental Fig. S2; Supplemental Table S2), establishing PcmPIM1 and PbmPIM1 as monofunctional, class I pimaradiene synthases (PIM) and adding a new function to the portfolio of conifer diTPSs. Coupled assays of PcmISO1 with PaLAS:D611A yielded a mixture of at least 90% isopimaradiene and up to 10% sandaracopimaradiene. Similarly, when PbmISO1 was coupled with PaLAS:D611A, 88% isopimaradiene and 12% sandaracopimaradiene were produced (Fig. 4; Supplemental Fig. S3; Supplemental Table S2). Further LC-MS analysis verified that the reaction products of PcmPIM1 and PcmISO1 were diterpene olefins, as opposed to a C13 alcohol.
Diterpene product profiles of PbmPIM1. GC-MS analysis of reaction products of PaLAS:D611A (A) and PaLAS:D611A combined with PbmPIM1 (B) in comparison to a pimaradiene standard (C). D and E, Mass spectra of reaction products marked with an asterisk in B and C. E, Internal standard 1-eicosene. m/z, Mass-to-charge ratio.
Diterpene product profiles of PbmISO1. GC-MS analysis of reaction products of PaLAS:D611A (A) and PaLAS:D611A combined with PbmISO1 (B) compared with retention times of authentic isopimaradiene (C) and sandaracopimaradiene (D) standards. E and F, Mass spectra of reaction products marked with an asterisk in B and C. E, Internal standard 1-eicosene. m/z, Mass-to-charge ratio.
Coupled assays with PcmdiTPS1, PcmdiTPS2, PcmdiTPS3, and PbmdiTPS1 did not produce any obvious diterpene compounds using GC-MS and LC-MS analyses. Coupled assays with ZmAN2 did not result in significant product formation for any of the eight candidate pine diTPSs. Taken together, the combinatorial approach of coupled enzyme assays identified two pairs of orthologous monofunctional class I diTPSs, with PcmPIM1 and PbmPIM1 representing new PIM-type diTPSs and PcmISO1 and PbmISO1 representing new monofunctional ISO-type diTPSs. The class I activity of these enzymes appeared to be specific for (+)-CPP and directly produced diterpene olefins as opposed to unstable diterpene alcohols.
Bifunctional Class I/II PcLAS1 and PbLAS1 Can Supply (+)-CPP for the Monofunctional Class I PcmPIM1, PbmPIM1, PcmISO1, and PbmISO1
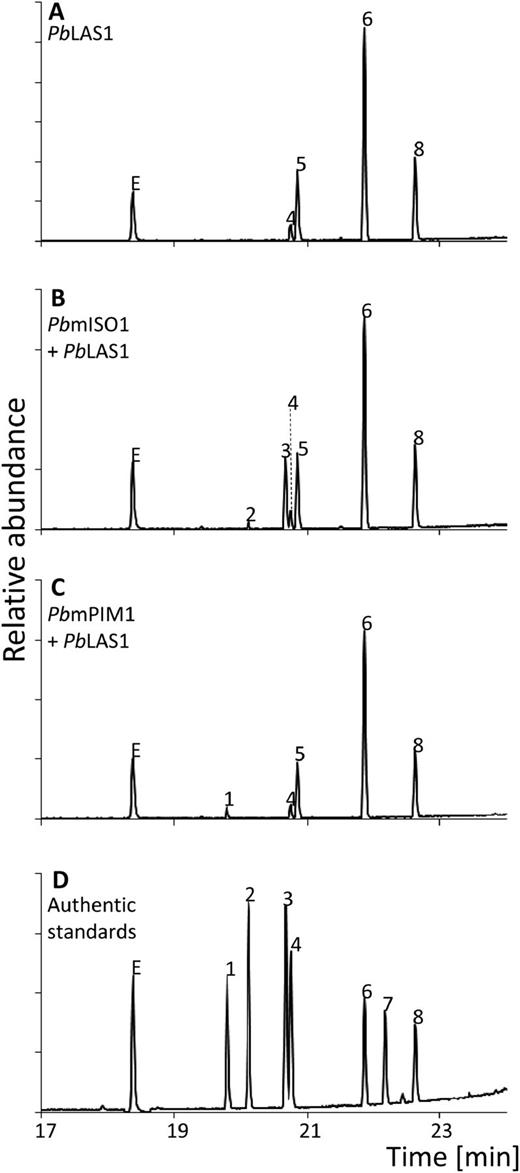
Previous studies demonstrated that (+)-CPP is released from the class II active site of the bifunctional Abies grandis abietadiene synthase (AgAS), prior to catalysis of the class I reaction (Peters et al., 2001). To determine in vitro if (+)-CPP released from an intact bifunctional class I/II diTPS can act as a substrate for the monofunctional pine diTPSs, coupled assays with PbLAS and either PbmPIM1 or PbmISO1 and GGPP as a substrate were conducted and compared with the activity of the LAS enzyme alone. Combining PbLAS1 and PbmISO1 led to the accumulation of additional product peaks of sandaracopimaradiene and isopimaradiene (peaks 2 and 3 in Fig. 5B) in addition to abietadiene, levopimaradiene, palustradiene, and neoabietadiene, which were produced by PbLAS1 alone (Fig. 5A). Isopimaradiene accounted for 8% of the total product profile in these assays (Supplemental Table S2). Coupled assays of PbLAS1 and PbmPIM1 resulted in the accumulation of 1.6% pimaradiene as a new product peak (peak 1 in Fig. 5C; Supplemental Table S2).
Diterpene product profiles of coupled assays with monofunctional PbmISO1 or PbmPIM1 and bifunctional PbLAS1. GC-MS analysis of reaction products from coupled enzyme assays of monofunctional PbmISO1 and PbmPIM1 with bifunctional PbLAS1, indicating the conversion of (+)-CPP, released from the LAS class II active site, to the different pimaradienes through the class I activity of PbmPIM1 and PbmISO1. Product profiles, with GGPP as substrate, of PbLAS1 (A), PbLAS1 combined with PbmISO1 (B), and PbLAS1 combined with PbmPIM1 (C). D, Authentic standards: 1, pimaradiene; 2, sandaracopimaradiene; 3, isopimaradiene; 4, palustradiene; 5, levopimaradiene; 6, abietadiene; 7, dehydroabietadiene; 8, neoabietadiene. E, Internal standard 1-eicosene.
Similarly, incubation of PcLAS1 with either PcmISO1 or PcmPIM1 led to the production of sandaracopimaradiene and isopimaradiene or pimaradiene, respectively, in addition to diterpene olefins produced by PcLAS1 (Supplemental Fig. S4; Supplemental Table S2). The pimaradiene and isopimaradiene peaks accounted for 3.7% and 10.5% of the total diterpene olefins produced in these assays (Supplemental Table S2).
DRA Profiling of Jack Pine and Lodgepole Pine Tissue Samples Confirms the Presence of Pimaradiene-, Sandaracopimaradiene-, and Isopimaradiene-Derived DRAs
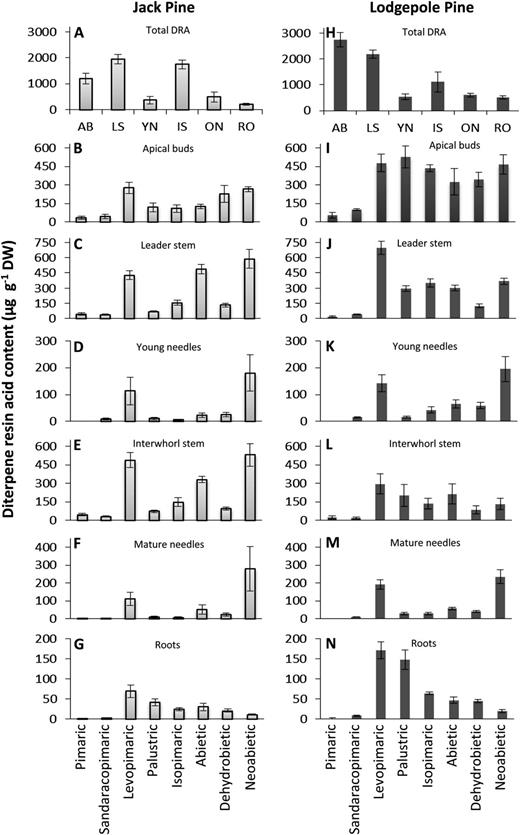
To test if the in vitro products of the newly discovered monofunctional class I diTPSs are relevant for the in vivo diterpene metabolite profiles of jack pine and lodgepole pine, we produced diterpenoid metabolite profiles of extracts from jack pine and lodgepole pine saplings. Extracts were prepared from six tissue types of each species, including apical buds, leader stem tissues, young needles, interwhorl stem tissue, mature needles, and roots (Fig. 6). GC-MS analysis of organic solvent extracts identified the DRAs as the most abundant diterpenoids across all tested tissue types in both jack pine and lodgepole pine. Lesser amounts of the corresponding aldehydes, alcohols, and olefins were also present (Supplemental Figs. S5 and S6), which is consistent with recent diterpenoid profiling of Sitka spruce in which DRAs accounted for more than 90% of the total diterpenoids in stem tissues (Hamberger et al., 2011). Detailed analysis of the total DRA content in the jack pine and lodgepole pine tissues revealed that the abundance of these metabolites was highest in apical buds, leader stem, and interwhorl stem tissues for both species (Fig. 6, A and H). The total amount of DRAs was similar across both species for each tissue type, with the exception of apical buds, where lodgepole pine had higher levels of DRAs.
DRA contents in 3-year-old jack pine and lodgepole pine saplings. Total DRA content (A and H), apical buds (AB; B and I), leader stem (LS; C and J), young needles (YN; D and K), top interwhorl stem (IS; E and L), top interwhorl mature needles (ON; F and M), and roots (RO; G and N). ses are based on five biological replicates with at least two technical replicates per sample. DW, Dry weight.
Pimaric acid, sandaracopimaric acid, and isopimaric acid, which correspond to the diterpene olefins produced by PcmPIM1, PbmPIM1, PcmISO1, and PbmISO1, were present in all (sandaracopimaric acid and isopimaric acid) or several (pimaric acid) of the analyzed tissues of jack pine and lodgepole pine. Pimaric acid and sandaracopimaric acid were detected as the least abundant DRAs, while higher levels of isopimaric acid were detected in most tissue types (Fig. 6). In extracts from lodgepole pine, levopimaric acid was the most abundant DRA in the leader stem, and a mix of similar levels of levopimaric acid, palustric acid, isopimaric acid, abietic acid, dehydroabietic acid, and neoabietic acid was detected in extracts from interwhorl stem tissue. In extracts from jack pine leader stem and the upper interwhorl stem tissues, levopimaric acid, abietic acid, and neoabietic acid were the most abundant DRAs. In both young and mature needles, levopimaric acid and neoabietic acid were the most abundant DRAs in extracts from lodgepole pine and jack pine. In roots, similar DRA profiles were observed from both pine species; however, the macrocyclic diterpenoid thunbergol was also detected in four of the five biological replicates from lodgepole pine.
The results from metabolite profiling of DRAs confirm that the products of PcmPIM1, PbmPIM1, PcmISO1, and PbmISO1 are relevant for the oleoresin composition of jack pine and lodgepole pine, although the corresponding DRAs are of relatively low abundance.
Mapping of the 454 and Illumina Sequence Data Identifies PcLAS1/2 and PbLAS1 as the Most Highly Expressed Genes
Mapping of the FL diTPS sequences against the jack pine and lodgepole pine 454 and Illumina transcriptome data obtained from sapling stem tissues confirmed the presence of the monofunctional class II and the bifunctional class I/II diTPSs in both data sets, with the exception of PcmdiTPS3, which was present only in the larger Illumina dataset with higher coverage (Table I). Due to the high identity (>99%) of PcLAS1 and PcLAS2 on the nucleotide level, it was not possible to map these genes independently; however, targeted analysis of the 21-bp segment that is present in PcLAS1, but not PcLAS2, suggested that PcLAS1 represents 86% of the Illumina reads and 79% of the 454 reads that are attributed to transcripts of these two bifunctional diTPSs.
Abundance of candidate diTPSs in the jack pine and lodgepole pine transcriptome data
| . | No. of Reads . | . | |
|---|---|---|---|
| Gene Name and Species . | 454 . | Illumina . | Relative Expressiona . |
| Lodgepole pine | |||
| PcLAS1, PcLAS2 | 614 | 16,560 | 100.00 |
| PcmISO1 | 1 | 111 | 0.67 |
| PcmPIM1 | 17 | 456 | 2.80 |
| PcmdiTPS1 | 7 | 132 | 0.80 |
| PcmdiTPS2 | 12 | 279 | 1.70 |
| PcmdiTPS3 | 0 | 51 | 0.31 |
| Jack pine | |||
| PbLAS1 | 516 | 69,608 | 100.00 |
| PbmISO1 | 131 | 12,611 | 18.10 |
| PbmPIM1 | 2 | 570 | 0.82 |
| PbmdiTPS1 | 12 | 928 | 1.30 |
| . | No. of Reads . | . | |
|---|---|---|---|
| Gene Name and Species . | 454 . | Illumina . | Relative Expressiona . |
| Lodgepole pine | |||
| PcLAS1, PcLAS2 | 614 | 16,560 | 100.00 |
| PcmISO1 | 1 | 111 | 0.67 |
| PcmPIM1 | 17 | 456 | 2.80 |
| PcmdiTPS1 | 7 | 132 | 0.80 |
| PcmdiTPS2 | 12 | 279 | 1.70 |
| PcmdiTPS3 | 0 | 51 | 0.31 |
| Jack pine | |||
| PbLAS1 | 516 | 69,608 | 100.00 |
| PbmISO1 | 131 | 12,611 | 18.10 |
| PbmPIM1 | 2 | 570 | 0.82 |
| PbmdiTPS1 | 12 | 928 | 1.30 |
Relative gene expression was calculated based on the number of reads in the Illumina data set when PcLAS1 and PcLAS2 and PbLAS1 are set at 100% for lodgepole pine and jack pine, respectively.
| . | No. of Reads . | . | |
|---|---|---|---|
| Gene Name and Species . | 454 . | Illumina . | Relative Expressiona . |
| Lodgepole pine | |||
| PcLAS1, PcLAS2 | 614 | 16,560 | 100.00 |
| PcmISO1 | 1 | 111 | 0.67 |
| PcmPIM1 | 17 | 456 | 2.80 |
| PcmdiTPS1 | 7 | 132 | 0.80 |
| PcmdiTPS2 | 12 | 279 | 1.70 |
| PcmdiTPS3 | 0 | 51 | 0.31 |
| Jack pine | |||
| PbLAS1 | 516 | 69,608 | 100.00 |
| PbmISO1 | 131 | 12,611 | 18.10 |
| PbmPIM1 | 2 | 570 | 0.82 |
| PbmdiTPS1 | 12 | 928 | 1.30 |
| . | No. of Reads . | . | |
|---|---|---|---|
| Gene Name and Species . | 454 . | Illumina . | Relative Expressiona . |
| Lodgepole pine | |||
| PcLAS1, PcLAS2 | 614 | 16,560 | 100.00 |
| PcmISO1 | 1 | 111 | 0.67 |
| PcmPIM1 | 17 | 456 | 2.80 |
| PcmdiTPS1 | 7 | 132 | 0.80 |
| PcmdiTPS2 | 12 | 279 | 1.70 |
| PcmdiTPS3 | 0 | 51 | 0.31 |
| Jack pine | |||
| PbLAS1 | 516 | 69,608 | 100.00 |
| PbmISO1 | 131 | 12,611 | 18.10 |
| PbmPIM1 | 2 | 570 | 0.82 |
| PbmdiTPS1 | 12 | 928 | 1.30 |
Relative gene expression was calculated based on the number of reads in the Illumina data set when PcLAS1 and PcLAS2 and PbLAS1 are set at 100% for lodgepole pine and jack pine, respectively.
Based on the number of reads present for each diTPS in the Illumina data set, we calculated the transcript abundance of the individual diTPSs relative to the abundance of PcLAS1/2 for lodgepole pine and relative to PbLAS1 for jack pine (Table I). In jack pine, transcript abundance of PbmISO1 was 5.5-fold lower than that of PbLAS1, whereas transcript abundance of PbmPIM1 and PbmdiTPS1 was 77- and 122-fold lower, respectively. In lodgepole pine, transcript abundance of PcmPIM1 and PcmdiTPS2 was 36- and 59-fold lower than that of PcLAS1/2, respectively, and transcripts of PcmISO1, PcmdiTPS1, and PcmdiTPS3 were at least 100 times less abundant than PcLAS1/2 (Table I). For logistic reasons, transcriptome sequencing was done with 2-year-old saplings, while DRA analysis was done with 3-year-old saplings. The relatively low transcript abundance of the monofunctional diTPSs PcmPIM1, PbmPIM1, PcmISO1, and PbmISO1, relative to the bifunctional PcLAS1/2 and PbLAS1, is in agreement with the relative abundance of the corresponding DRAs described above.
Evolution of Monofunctional Class I diTPSs of Specialized Metabolism
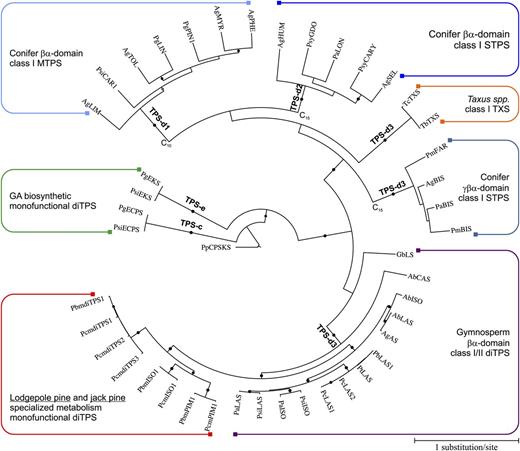
Based on their monofunctional activity and role in DRA biosynthesis, the newly discovered class I diTPSs of jack pine and lodgepole pine introduce a new functionality to the catalytic landscape of specialized conifer diTPSs. To gain a deeper understanding of their evolution, we performed a phylogenetic analysis, including mono- and bifunctional diTPSs of DRA biosynthesis, monofunctional class I and class II diTPSs of GA biosynthesis, and select conifer monofunctional class I mono-, sesqui-, and diTPSs. It was previously shown that bifunctional class I/II diTPSs of conifer DRA biosynthesis belong to a gymnosperm-specific TPS-d3 subfamily, while the monofunctional gymnosperm diTPSs of GA biosynthesis cluster together with angiosperm diTPSs of GA biosynthesis in the TPS-c and TPS-e/f subfamilies (Keeling et al., 2010; Chen et al., 2011). A maximum likelihood-based phylogenetic analysis places the jack pine and lodgepole pine mono- and bifunctional diTPSs in the gymnosperm-specific TPS-d3 clade, containing primarily diTPSs and few γβα-domain sesqui-TPSs, clearly distant from GA biosynthetic ECPS and EKS (Fig. 7). The bifunctional class I/II PbLAS1, PcLAS1, and PcLAS2 are closely related to previously characterized bifunctional LAS and ISO enzymes from grand fir (Vogel et al., 1996), Norway spruce (Martin et al., 2004), loblolly pine (Ro and Bohlmann, 2006), Sitka spruce (Keeling et al., 2011a), and balsam fir (Zerbe et al., 2012a). The eight monofunctional diTPSs (PcmPIM1, PbmPIM1, PcmISO1, PcmISO1, PbmdiTPS1, PcmdiTPS1, PcmdiTPS2, and PcmdiTPS3) form a distinct branch within the TPS-d3 family adjacent to the bifunctional class I/II diTPSs of DRA biosynthesis, but situated distantly from other monofunctional gymnosperm TPS, such as Taxus spp. taxadiene synthase (Wildung and Croteau, 1996), γβα-domain gymnosperm sesqui-TPS (Bohlmann et al., 1998b, Martin et al., 2004; Huber et al., 2005, McAndrew et al., 2011), and other βα-domain conifer mono-TPSs. Furthermore, the orthologous pairs of PcmPIM1/PbmPIM1 and PcmISO1/PcmISO1, for which we showed biochemical functions, are separated from the four remaining diTPS candidates (PbmdiTPS1, PcmdiTPS1, PcmdiTPS2, and PcmdiTPS3) for which no activity was observed.
Phylogenetic tree of FL diTPS from jack pine and lodgepole pine with previously characterized members of the gymnosperm-specific TPS-d family. P. patens ent-kaurene/kaurenol synthase (PpCPS/KS; accession no. BAF61135) was used to root the tree. Branches indicated with dots represent bootstrap support more than 80% (100 repetitions). Alterations in the substrate specificity from GGPP (C20) to FPP (C15) and GPP (C10) are indicated. Abbreviations and NCBI accession numbers are as follows: PcmISO1 (JQ240314), PbmISO1 (JQ240313), PcmPIM1 (JQ240315), PbmPIM1 (JQ240316), PcmdiTPS1 (JQ240318), PbmdiTPS1 (JQ240317), PcmdiTPS2 (JQ240319), PcmdiTPS3 (JQ240320), PtLAS (Q50EK2), PcLAS2 (JQ240311), PcLAS1 (JQ240310), PbLAS1 (JQ240312), PsiLAS (ADZ45517), PaLAS (Q675L4), PsiISO (ADZ45512), PaISO (Q675L5), AbLAS (JN254805), AgAS (Q38710), AbISO (JN254806), AbCAS (JN254808), GbLS (Q947C4), TcTXS (ABC25488), TbTXS (Q41594), AgBIS (O81086), PaBIS (AAS47689), PmBIS (Q4QSN4), PmFAR (ADX42737), PsiEKS (ADB55710), PgEKS (ADB55708), ΨECPS (ADB55709), PgECPS (ADB55707), AgHUM (O64405), AgSEL (AAC05727), AgLIM (AAB70907), AgTOL (AAF61454), AgPHE (Q9M7D1), AgMYR (AAB71084), PaLON (AAS47695), PgPIN1 (ADZ45507), PgLIN (ADZ45500), PsiCAR1 (ADZ45511), PsyCARY (ABV44452), and PsyGDO (ABV44453).
These results show that monofunctional gymnosperm diTPSs evolved independently on at least three separate occasions, one being the evolution of monofunctional diTPSs of GA biosynthesis in gymnosperms and angiosperms, the second being the evolution of a monofunctional taxadiene synthase, and the third being the emergence of monofunctional diTPSs of pine DRA biosynthesis described here. The latter type of monofunctional class I diTPS appears to have evolved by loss of functionality of the class II active site that remained intact in the similar class I/II LAS and ISO enzymes.
DISCUSSION
Monofunctional diTPSs Are New Modules in the Biosynthesis of Multiproduct DRAs
Conifers produce a diverse and dynamic mixture of large volumes of terpenoids, which contribute to the plasticity of a broad defense shield against many generalist herbivores and protect against specialist insect pests and pathogens (Keeling and Bohlmann, 2006a; Zulak and Bohlmann, 2010). DRAs are a major component of the chemical and physical defense of conifers (Phillips and Croteau, 1999) and have been implicated, for example, in pine defense to the bark beetle associated fungus O. ips (Kopper et al., 2005) and in Sitka spruce resistance to the white pine weevil (Robert et al., 2010). The committed reactions in DRA biosynthesis are catalyzed by diTPSs (Keeling and Bohlmann, 2006a, 2006b) and cytochrome P450 enzymes (Ro et al., 2005; Hamberger et al., 2011). Previously, it has been shown that conifer diTPSs involved in specialized metabolism of DRAs are bifunctional class I/II enzymes that catalyze the conversion of GGPP, through the stable intermediate (+)-CPP, to produce diterpene olefins or alcohols (Vogel et al., 1996; Peters et al., 2000; Schepmann et al., 2001; Martin et al., 2004; Ro and Bohlmann, 2006; Keeling et al., 2011b; Zerbe et al., 2012a). Conversely, the general GA metabolism in gymnosperms and angiosperms, and likewise the specialized metabolism of labdane-related diterpenoids in angiosperms, involve separate consecutively acting monofunctional class I and class II enzymes (Richman et al., 1999; Xu et al., 2004; Harris et al., 2005; Prisic and Peters, 2007; Xu et al., 2007a; Gao et al., 2009; Toyomasu et al., 2009; Keeling et al., 2010; Falara et al., 2010; Caniard et al., 2012).
To investigate DRA biosynthesis in jack pine and lodgepole pine, we generated deep transcriptome sequence inventories and de novo assemblies from which we identified, cloned, and characterized 11 FL diTPSs. Functional characterization of the three jack pine and lodgepole pine LAS enzymes (PbLAS1, PcLAS1, and PcLAS2), which produce the unstable diterpene tertiary alcohol 13-hydroxy-8(14)-abietene, suggested that water capture by a carbocation at the class I active site is a conserved and common reaction mechanism of LAS enzymes across several conifer species. This mechanism was recently proposed when 13-hydroxy-8(14)-abietene, which dehydrates to form abietadiene, palustradiene, levopimaradiene, and neoabietadiene, was first discovered as the primary product of the LAS from Norway spruce and balsam fir (Keeling et al., 2011b; Zerbe et al., 2012a).
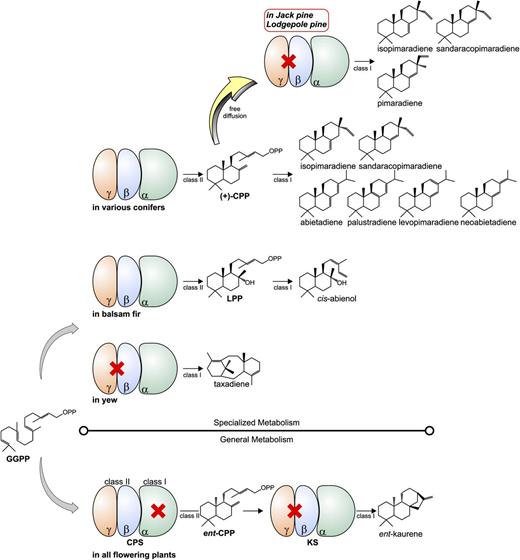
Unexpectedly, absence of the class II active site motif DxDD suggested that eight of the 11 candidate sequences were not typical bifunctional conifer diTPSs. Functional characterization identified the orthologous pair of PbmPIM1 and PcmPIM1 as single-product pimaradiene synthases, while the orthologous pair of PbmISO1 and PcmISO1 are isopimaradiene synthases, which also produced small amounts of sandaracopimaradiene. Monofunctional class I diTPSs of specialized DRA metabolism have not been previously reported. The discovery of the orthologous genes in two closely related pine species makes it unlikely that these monofunctional diTPSs represent cloning or sequencing artifacts. The only other known example of a monofunctional class I diTPS of specialized metabolism in a gymnosperm is taxadiene synthase (Wildung and Croteau, 1996), which converts GGPP directly into the macrocyclic taxadiene backbone without a bicyclic diphosphate intermediate. The monofunctional pimaradiene synthases described here are, to the best of our knowledge, the first reported gymnosperm enzymes that produce predominantly pimaradiene and extend the scope of known conifer diTPS functions involved in DRA formation beyond the previously known ISO and LAS enzymes (Fig. 8).
Schematic of diterpene biosynthesis in gymnosperms, including conifers and Taxus spp. A portfolio of diTPS with distinct modular architecture drives the formation of the wide array of naturally occurring diterpene metabolites in gymnosperms. As in angiosperm general and specialized diterpene metabolism, two monofunctional class II and class I diTPSs function in GA biosynthesis in conifer general metabolism. In contrast, bifunctional class I/II diTPSs are responsible for the formation of specialized abietane- and pimarane-related compounds of conifer oleoresin. Monofunctional diTPSs of gymnosperm specialized metabolism are the class I taxadiene synthases in species of yew and the newly discovered monofunctional ISO and PIM enzymes of jack pine and lodgepole pine. The latter enzymes can utilize the (+)-CPP pool of bifunctional LAS and ISO diTPS to form pimaradiene, sandaracopimaradiene, and isopimaradiene, which are present as the corresponding DRAs in lodgepole and jack pine. LPP, Labda-13-en-8-ol diphosphate; CPS, CPP synthase; KS, kaurene synthase.
Considering the modularity of plant specialized (i.e. secondary) metabolism, our discovery of monofunctional class I diTPSs of DRA biosynthesis highlights that in-depth mining and functional analysis of plant transcriptomes can reveal new elements of biosynthetic pathways, even when all essential enzymes of the pathway were thought to be known. Matrix-type pathway systems of specialized metabolism may allow for more than one route to arrive at particular metabolites. The DRA biosynthesis of conifers is emerging as a model for such a dynamic metabolic network that is built from multiple diTPSs and cytochrome P450s of variable functions and activities to produce a diverse array of similar diterpene compounds (Keeling et al., 2008, 2010; Hamberger et al., 2011).
A Role for the Lodgepole Pine and Jack Pine Monofunctional diTPSs in DRA Biosynthesis
The diterpenoid profiles in the different tissues of jack pine and lodgepole pine confirmed that the bifunctional diTPSs, which produced the abietane-type diterpene olefins (abietadiene, neoabietadiene, palustradiene, and levopimaradiene), and the monofunctional diTPSs that formed pimarane-type olefins (isopimaradiene, pimaradiene, and sandaracopimaradiene) supplied seven of the eight predominant diterpene structures detected, with only the formation of the dehydroabietane backbone not accounted for. Biosynthesis of dehydroabietadiene has not yet been resolved in any plant species. Our analysis found that, in all tissues tested, abietane-type DRAs were more abundant than pimarane-type DRAs. These profiles are consistent with the metabolite profiles of Sitka spruce stem tissue where DRAs accounted for 92% of the total diterpenes, the majority of which were of the abietane type (Hamberger et al., 2011) and with previous DRA analysis of stem sections from 1-year-old lodgepole pine saplings where the abietane-type DRAs, as well as isopimaric acid, were the most abundant diterpenoids (Lewinsohn et al., 1993). We conclude that the newly identified monofunctional PIM (PbmPIM1 and PcmPIM1) and ISO enzymes (PbmISO1 and PcmISO1) can account for the three pimarane-type DRAs of lodgepole pine, which accumulate in relatively low amounts in the tissues tested. The low abundance of the pimarane-type DRAs may be attributed to the low level of transcript abundance of the monofunctional diTPSs and to their requirement for a (+)-CPP substrate released by a separate, presumably bifunctional, diTPS.
Mapping of transcriptome sequence data to the functionally characterized diTPSs allowed us to assess the relative transcript abundance of these genes in lodgepole pine and jack pine leader and interwhorl stem tissue. PcLAS1, PcLAS2, and PbLAS1 were the most highly abundant transcripts, consistent with the high levels of abietane-type DRAs. In jack pine, PbmISO1 and PbmPIM1 transcripts were detected at 5- and 122-fold lower levels than PbLAS1, consistent with the lower levels of pimarane-type DRAs, as well as with the greater abundance of isopimaric acid than pimaric acid in the leader and interwhorl stem tissues. In lodgepole pine, PcLAS1/2 transcripts were at 149- and 36-fold higher levels than PcmISO1 and PcmPIM1, respectively. The lower expression of PcmPIM1 is consistent with the lower quantities of pimaric acid in these tissues; however, isopimaric acid is more abundant than pimaric acid in all tissue extracts of lodgepole pine, which suggests that other TPS may be involved in the production of isopimaric acid. Bifunctional ISO enzymes have been previously identified from Norway spruce (Martin et al., 2004), Sitka spruce (Keeling et al., 2011a), and balsam fir (Zerbe et al., 2012a); thus, it is possible that an orthologous bifunctional ISO exists in pine, which may account for the discrepancy between the transcript abundance and metabolite levels in these tissues. However, despite the deep transcriptome sequences, no obvious ISO candidate was found in lodgepole pine or jack pine.
The monofunctional class I diTPSs identified in this study were active with (+)-CPP, but not with ent-CPP or GGPP, as substrate. While ent-CPP is produced in conifers by monofunctional class II diTPSs dedicated to GA biosynthesis (Keeling et al., 2010), a monofunctional class II diTPS that produces (+)-CPP has not yet been described for a gymnosperm and has so far only been reported from the angiosperm Salvia miltiorrhiza (Gao et al., 2009). The transcriptome resources available for pine and other conifers (e.g. Sitka spruce and white spruce; Ralph et al., 2008; Rigault et al., 2011) also did not reveal any significant candidates for (+)-CPP-producing class II diTPSs. However, it has been previously demonstrated that the (+)-CPP formed by the bifunctional class I/II AgAS can freely diffuse from the class II active site (Peters et al., 2001). In agreement with these earlier findings, we showed that the bifunctional class I/II diTPSs PcLAS1 and PbLAS1 could supply (+)-CPP for the monofunctional class I diTPSs in vitro, which confirms that (+)-CPP can diffuse from the class II active site of the pine LAS proteins. Assuming that bifunctional and monofunctional diTPSs are present in the same cells and subcellular compartments, presumably in the epithelial cells of resin ducts (Abbott et al., 2010; Zulak et al., 2010), the more abundant bifunctional LAS may be a source of (+)-CPP for monofunctional diTPSs to yield the low quantities of pimarane-type DRAs as detected in the lodgepole pine and jack pine metabolite profiles (Fig. 8).
Monofunctional diTPSs of DRA Biosynthesis Originated from Bifunctional diTPSs
The bifunctional ent-kaurene/kaurenol synthases of the nonvascular plants P. patens and J. subulata are thought to represent ancestral plant diTPSs (Hayashi et al., 2006; Kawaide et al., 2011). Lineage-specific gene duplications followed by sub- and neofunctionalization of such archetypical class I/II diTPSs are likely events along the path that led to the families of diTPSs of general and specialized metabolism found in species of conifers (gymnosperms) and angiosperms (Chen et al., 2011). A phylogeny of the jack pine and lodgepole pine diTPSs with previously characterized conifer TPS of the TPS-c, TPS-e/f, and TPS-d subfamilies (Martin et al., 2004; Chen et al., 2011) illustrates these events leading to a variety of functionally diverse diTPS of specialized metabolism (Fig. 7). The topology of the phylogenetic tree suggests that the monofunctional class I diTPSs of specialized metabolism in lodgepole pine and jack pine have evolved relatively recently through gene duplication of a bifunctional diTPS followed by loss of the class II activity and additional functional diversification. While the bifunctional LAS enzymes of lodgepole pine and jack pine have orthologs in other conifers within and outside of the pine genus, e.g. in loblolly pine, spruce, and true firs, monofunctional class I diTPSs of specialized metabolism have not been found in other conifers. It is possible that they represent a lineage-specific clade of the TPS-d3 group that originated in a common ancestor of the closely related species of lodgepole pine and jack pine, perhaps after the separation from loblolly pine and after the separation of the pine, spruce, and fir genera.
Alternatively, a direct origin of monofunctional class I diTPSs of specialized metabolism from monofunctional class I EKS could be considered, since mutagenesis studies with angiosperm and conifer enzymes demonstrated that functional evolution of EKS to a pimarane-type diTPS requires exchange of only one amino acid in the class I active site (Xu et al., 2007b; Zerbe et al., 2012b). However, the evolutionary distance of the pine monofunctional diTPSs to members of the GA biosynthetic TPS-c and TPS-e families inferred by the phylogenetic tree (Fig. 7) along with the low sequence similarity between both enzyme groups do not support this hypothesis.
Overall, the phylogeny indicates that events of monofunctionalization (i.e. a form of subfunctionalization from a duplicated bifunctional ancestor) have occurred multiple times independently in the evolution of gymnosperm diTPSs, leading to class II and class I enzymes of general GA metabolism, the Taxus spp. taxadiene synthases, and the newly described monofunctional pine diTPS (Fig. 8). Beyond the diTPSs, monofunctionalization ultimately also led to the large family of bidomain conifer mono- and sesqui-TPSs of the TPS-d1 and TPS-d2 groups.
Unique features in the active-site composition of the monofunctional diTPSs of lodgepole pine and jack pine underpin the proposed relatively recent origin from a class I/II diTPS ancestor. Homology modeling of all eight PcmPIM1 and PbmPIM1, mISO1, and mdiTPS based on the crystal structure of Taxus brevifolia taxadiene synthase (Köksal et al., 2011a; Protein Data Bank IDs 3P5P and 3P5R) illustrated that the monofunctional pine diTPSs exhibited the conserved γβα-domain folding pattern (Supplemental Fig. S7). Within the functional α-domain (downstream of the conserved RRWW segment; Supplemental Fig. S1) of PcmPIM1 and PbmPIM1 and PcmdiTPS and PbmdiTPS, 27 and 28 residues, respectively, were found to be unique compared with characterized bifunctional LAS and ISO enzymes and the monofunctional PcmISO1 and PbmISO1 enzymes. Among these, Ala-718 and Ser-721 of PcmPIM1 and PbmPIM1 are located adjacent to the hinge segment between helix G1 and G2, the composition of which is critical for the product specificity of LAS and ISO (Peters et al., 2003; Wildermann and Peters, 2007; Keeling et al., 2008) and may have contributed to the neofunctionalization toward PIM functionality (Supplemental Fig. S7). Notably, key residues in the class I active site that determine the product specificity of ISO- and LAS-type diTPSs (Keeling et al., 2008) are not conserved in the monofunctional ISO from pine, whereas these amino acids are strictly conserved in the pine bifunctional LAS enzymes, consistent with the product profiles of these enzymes. Similar differences in these catalytic residues have also been shown for the balsam fir ISO (Zerbe et al., 2012a) and suggest that formation of the tertiary abietenyl+ carbocation en route to abietane-type diterpenes, rather than the direct deprotonation to isopimaradiene depends on the strict conservation in these positions. In the four PcmdiTPSs and PbmdiTPSs, for which no obvious function was found, it was noticeable that the side chains of Phe-592, located upstream of the DDxxD motif, and likewise Phe-814 and His-817 on helix J protrude into the active site cavity and may cause a steric hindrance, possibly impeding catalytic activity (Supplemental Fig. S7). It can be speculated that these enzymes may have evolved from functional diTPSs into a trough of no function, from where they may evolve toward new diTPS activities or simply represent dead-end mutations of functional diTPSs.
MATERIALS AND METHODS
Plant Materials
One-year-old jack pine (Pinus banksiana clone ID PSB 410 1 + 0) and lodgepole pine (Pinus contorta clone ID PLI 144) saplings were obtained from the University of Alberta and from the British Columbia Ministry of Forests, Lands, and Natural Resource Operations, respectively. These trees were maintained for 2 or 3 years outdoors at the University of British Columbia as described previously for other conifers used in our research (Hall et al., 2011). For RNA extractions, four 2-year-old saplings of each species were moved into the greenhouse and were maintained at 24°C and 16 h light per day for 2 weeks prior to induction of defense gene expression. For induction, the stem of each tree was wounded with a razor blade and the aerial portion of the tree was sprayed with 50 mL of 0.1% methyl jasmonate. One-quarter of the stem (combined bark and xylem) of each tree was harvested 2, 6, 24, and 48 h following treatment, and the four time points from each individual were pooled prior to RNA extraction. For metabolite profiling, five 3-year-old saplings of each species were moved into the greenhouse for 2 weeks prior to harvest. Six tissue types were harvested from each tree and were flash frozen and stored at −80°C until processing with the exception that combined bark and xylem tissue was cut into 1-cm sections prior to freezing.
Transcriptome Mining, RNA and cDNA Work, and RACE
Details of the transcriptome resources developed by Sanger, 454, and Illumina sequencing will be described elsewhere. In brief, total RNA was extracted from the bark tissue of a single jack pine and a single lodgepole pine individual and was subjected to cDNA library construction and Sanger sequencing. Also, total RNA was extracted from the combined bark and xylem tissues of a second lodgepole pine individual and a second jack pine individual and was submitted to the McGill University and Génome Québec Innovation Centre for cDNA library construction, Roche-GS-FLX Titanium 454 sequencing, and Illumina sequencing. Raw sequences obtained from each platform were trimmed and filtered prior to the production of four unique assemblies containing (1) Sanger sequences only (assembled with CAP3), (2) 454 sequences only (assembled with Newbler), (3) 454 and Sanger sequences combined (assembled with Newbler), and (4) Illumina sequences only (assembled with Trinity).
A TBLASTN search with 102 previously characterized conifer TPSs (Keeling and Bohlmann, 2006a; Keeling et al., 2011a) as query sequences was used to identify diTPS-like sequences in the lodgepole pine and jack pine transcriptomic resources. Individual candidate sequences identified in the automated TBLASTN search of the transcriptome assembly were reassembled using Phrap (De la Bastide and McCombie, 2007) and manually validated to obtain a finalized set of diTPS candidates. This sequence information was used to design target-specific primers (Supplemental Table S3) for 5′ RACE or for cloning of FL cDNA. For the cloning of FL diTPS, RNA was extracted from a single jack pine sapling and a single lodgepole pine sapling and converted to cDNA for RACE or FL cDNA cloning as described previously (Hall et al., 2011). Amounts of 50 ng of jack pine RNA and 90 ng of lodgepole pine RNA were converted to cDNA using the Superscript III first-strand synthesis system (Invitrogen), and the cDNA was used as a template with gene specific primers (Supplemental Table S3) to amplify FL cDNA clones. For RACE, 1 μg of total RNA was processed using the SMARTer RACE cDNA amplification kit (Clontech), and this cDNA was used as a template with gene specific primers (Supplemental Table S3) and the universal primer mix as per the manufacturer’s protocol. Modifications of FL cDNAs were made with truncations upstream of the KRx8W motif to produce pseudomature proteins for protein expression without plastid transit peptides (Martin et al., 2004).
Protein Expression and diTPS Enzyme Assays
N-terminally truncated cDNAs of each candidate diTPS were cloned into pET28b(+) in frame with an N-terminal 6xHis tag using the In-Fusion PCR cloning system (Clontech) with primers listed in Supplemental Table S3. Recombinant proteins were expressed in Escherichia coli, Ni affinity purified, and characterized using single vial assays as described previously (O’Maille et al., 2004; Keeling et al., 2008; Hall et al., 2011). Each enzyme (35 µL) was initially assayed separately with 20 μ m GPP (Echelon), 70 μ m E,E-FPP (Echelon), and 40 μ m GGPP (Sigma-Aldrich) in the appropriate buffers (Keeling et al., 2011b), and the products were analyzed by GC-MS. diTPS candidates, which did not yield products with GPP, FPP, or GGPP, were then targeted for coupled assays with either (1) the maize (Zea mays) ent-CPP synthase, which produces ent-CPP (ZmAN2; Harris et al., 2005), or (2) a protein variant of Picea abies LAS (PaLAS:D611A; Zerbe et al., 2012a) in which the class I active site has been inactivated, resulting in the production of (+)-CPP. Coupled assays were performed as described previously (Keeling et al., 2010; Zerbe et al., 2012b) except that each assay contained 40 μ m GGPP and equal amounts (50 μg) of each recombinant protein. Assays containing ZmAN2 and PaLAS:D611A were preincubated for 2 h at 30°C to allow the accumulation of CPP, at which time 50 μg of the candidate diTPS protein was added, overlaid with pentane containing 1.6 μ m eicosene (Sigma-Aldrich) as an internal standard, and incubated for a further 1 h at 30°C. Production of CPP was confirmed by incubation of ZmAN2 or PaLAS:D611A with GGPP for 2 h at 30°C, at which point 10 units of calf alkaline phosphatase (Invitrogen) was added to the aqueous layer and incubated overnight at 37°C. Coupled assays in which PcLAS1 or PbLAS1 were used to supply (+)-CPP were immediately overlaid with pentane containing eicosene and incubated for 1 h total at 30°C. All proteins were characterized based on at least three independent duplicate assays.
Diterpene Extraction
Samples from six tissue types, (1) apical buds, (2) combined bark and xylem from the leader, (3) young needles from the leader, (4) combined bark and xylem from the first interwhorl, (5) needles from the first interwhorl, and (6) roots of jack pine and lodgepole pine saplings, were extracted in 1.5 mL of tert-butyl methyl ether (Sigma-Aldrich) containing 417 μ m 12,14-dichlorodehydroabietic acid (Orchid Cellmark) as described previously (Robert et al., 2010). A volume of 200 μL of ether extract was transferred to a fresh vial and derivatized with 2.0 μ m trimethylsilyldiazomethane in diethyl ether (Sigma-Aldrich) as described previously (Robert et al., 2010). Extractions were repeated with five biological replicates and three technical replicates per tissue type.
GC-MS and LC-MS Analysis of diTPS Assay Products and Metabolite Extractions
All GC-MS was completed on an Agilent 6890A Series GC system with an Agilent 7683 autosampler, coupled to an Agilent 5975 Inert XL mass spectrometric detector at 70 eV (Agilent Technologies). For enzyme assays, 0.2 to 1 μL of the pentane overlay was injected and compounds were separated on a DB-WAX capillary column (diameter = 250 µm, length = 30 m, and film thickness = 0.25 µm) with a helium flow of 1 mL/min and an initial temperature of 40°C for 5 min, increasing by 10°C per min to a final temperature of 240°C for 10 min. Two microliters of the dephosphorylated reaction products were separated on an SGE SolGel Wax column (diameter = 250 μm, length = 30 m, and film thickness = 0.25 μm) with an initial helium gas flow of 1.2 mL/min and an initial temperature of 40°C held for 2 min, increasing by 25°C per min to a final hold temperature of 250°C for 5 min. Diterpene content in extracts from jack pine and lodgepole pine tissues were separated on an AT-1000 column (diameter = 250 μm, length = 30 m, and film thickness = 0.25 μm) with a helium flow of 1 mL/min and an initial temperature of 150°C increasing by 1.5°C per min to 220°C, then by 20°C per min to a final hold of 240°C for 20 min. LC-MS was completed using methods described previously (Keeling et al., 2011b). Response factors were used for quantification and were calculated based on comparison with a known concentration of 12,14-dichlorodehydroabietic acid (metabolite extractions) or eicosene (recombinant enzyme assays).
Mapping of Illumina and 454 Sequences to Candidate diTPS Genes
Adapter-trimmed Illumina reads were mapped back to the FL diTPS sequences using BWA version 0.5.9-r16. Reads were allowed one mismatch at the alignment step and mapped as paired with maximum insert size of 350 bp. Single-end 454 Titanium sequences were mapped using BWA-SW with a stringent mismatch penalty score of 9.
Phylogenetic Analysis
Protein sequence alignments were generated using Dialign 2.2.1 (Morgenstern, 2004) and further manual inspection. Phylogenetic analyses were performed on the basis of a maximum likelihood-based approach using the PhyML-aBayes version 3.0.1 beta (Anisimova et al., 2011) with four rate substitution categories, LG substitution model, BIONJ starting tree, and 100 bootstrap repetitions. Phylogenetic trees were visualized using the Interactive Tree of Life v2 (iTOL) Web server (Letunic and Bork, 2011).
Computational Structure Analysis
Homology models of PcmPIM1, PbmPIM1, PcmISO1, PcmISO1, PbmdiTPS1, PcmdiTPS1, PcmdiTPS2, and PcmdiTPS3 were built on the crystal structure of taxadiene synthase from Taxus brevifolia (Protein Data Bank IDs 3P5R and 3P5P) using the same approach as reported previously (Keeling et al., 2011a). All models were of high quality with more than 91% of residues assigned to most favored regions in Ramachandran plot statistics, and pairwise structural comparisons showed a high similarity with the taxadiene synthase structure with root mean square deviations of less than 1 Å.
Sequence data from this article can be found in the NCBI GenBank database under accession numbers SRP009894 for the transcriptome resources of lodgepole pine and jack pine and under the accession numbers JQ240310 to JQ240320 for the FL cDNAs PcLAS1 (JQ240310), PcLAS2 (JQ240311), PbLAS1 (JQ240312), PbmISO1 (JQ240313), PcmISO1 (JQ240314), PcmPIM1 (JQ240315), PbmPIM1 (JQ240316), PbmdiTPS1 (JQ240317), PcmdiTPS1 (JQ240318), PcmdiTPS2 (JQ240319), and PcmdiTPS3 (JQ240320).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. Complete amino acid alignment of the FL diTPS from jack pine and lodgepole pine with select previously characterized conifer diTPS.
Supplemental Figure S2. Diterpene product profiles of PcmPIM1.
Supplemental Figure S3. Diterpene product profiles of PcmISO1.
Supplemental Figure S4. Diterpene product profiles of coupled assays with monofunctional PcmISO1 or PcmPIM1 and bifunctional PcLAS1.
Supplemental Figure S5. Representative GC-MS traces of the diterpene olefin, alcohol, aldehyde, and resin acid content of extracts from six tissue types of jack pine.
Supplemental Figure S6. Representative GC-MS traces of the diterpene olefin, alcohol, aldehyde, and resin acid content of extracts from six tissue types of lodgepole pine.
Supplemental Figure S7. Homology modeling of pine monofunctional diterpene synthases.
Supplemental Table S1. Amino acid sequence identity matrix of Pinus banksiana (Pb) and Pinus contorta (Pc) diterpene synthase candidate genes with previously characterized diTPS from Pinus taeda (PtLAS), Picea abies (PaLAS and PaISO), and Picea glauca ECPS and EKS.
Supplemental Table S2. Relative amount of individual diterpenes produced by the recombinant diTPS characterized in this study.
Supplemental Table S3. Gene-specific primers used for 5′ RACE or to amplify FL genes from lodgepole pine and jack pine cDNA.
Note Added in Proof
In addition to the previous report of a monofunctional (+)-CPP synthase from Salvia miltiorhizza (Gao et al., 2009), while our paper was under review, Wu et al. (Wu Y, Zhou K, Toyomasu T, Sugawara C, Oku M, Abe S, Usui M, Chandler PM, Peters RJ [2012] Functional characterization of wheat copalyl diphosphate synthases sheds light on the early evolution of labdane-related diterpenoid metabolism in the cereals. Phytochemistry 84: 40–46) published an article on wheat (Triticum aestivum) diTPS showing a (+)-CPP synthase functionality.
ACKNOWLEDGMENTS
We thank Ms. Karen Reid (University of British Columbia) for outstanding laboratory and project management support, Dr. Christopher I. Keeling (University of British Columbia) for helpful discussions and preparation of the PaLAS:D611A construct, Dr. Reuben Peters (Iowa State University) for the ZmAN2 construct, Dr. Janice Cooke (University of Alberta) for the jack pine material, and Dr. Nicholas Ukrainetz (British Columbia Ministry of Forests, Lands, and Natural Resources) for the lodgepole pine material.
Glossary
- TPS
terpene synthase
- diTPS
diterpene synthase
- GGPP
geranylgeranyl diphosphate
- DRA
diterpene resin acid
- (+)-CPP
(+)-copalyl diphosphate
- ISO
isopimaradiene synthase
- LAS
levopimaradiene/abietadiene synthase
- MPB
mountain pine beetle
- NCBI
National Center for Biotechnology Information
- FL
full-length
- ECPS
ent-copalyl diphosphate synthase
- EKS
ent-kaurene synthase
- LC-MS
liquid chromatography-mass spectrometry
- GC-MS
gas chromatography-mass spectrometry
- FPP
farnesyl diphosphate
- GPP
geranyl diphosphate
- CPP
copalyl diphosphate
- PIM
pimaradiene synthase
- cDNA
complementary DNA
LITERATURE CITED
Author notes
This work was supported by funding from the Natural Sciences and Engineering Research Council of Canada (Discovery Grant to J.B. and Postdoctoral Fellowship to D.E.H.) and by Genome British Columbia, Genome Alberta, and Genome Canada in support of the Tria Project (funds to J.B.). J.B. is a University of British Columbia Distinguished University Scholar.
These authors contributed equally to the article.
Corresponding author; e-mail bohlmann@msl.ubc.ca.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Jörg Bohlmann (bohlmann@msl.ubc.ca).
The online version of this article contains Web-only data.
Open Access articles can be viewed online without a subscription.